Abstract
Recent studies have revealed that newly emerging transformed cells are often apically extruded from epithelial tissues. During this process, normal epithelial cells can recognize and actively eliminate transformed cells, a process called epithelial defence against cancer (EDAC). Here, we show that mitochondrial membrane potential is diminished in RasV12-transformed cells when they are surrounded by normal cells. In addition, glucose uptake is elevated, leading to higher lactate production. The mitochondrial dysfunction is driven by upregulation of pyruvate dehydrogenase kinase 4 (PDK4), which positively regulates elimination of RasV12-transformed cells. Furthermore, EDAC from the surrounding normal cells, involving filamin, drives the Warburg-effect-like metabolic alteration. Moreover, using a cell-competition mouse model, we demonstrate that PDK-mediated metabolic changes promote the elimination of RasV12-transformed cells from intestinal epithelia. These data indicate that non-cell-autonomous metabolic modulation is a crucial regulator for cell competition, shedding light on the unexplored events at the initial stage of carcinogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Fialkow, P. J. Clonal origin of human tumors. Biochim. Biophys. Acta 458, 283–321 (1976).
Nowell, P. C. The clonal evolution of tumor cell populations. Science 194, 23–28 (1976).
Morata, G. & Ripoll, P. Minutes: mutants of Drosophila autonomously affecting cell division rate. Dev. Biol. 42, 211–221 (1975).
de la Cova, C., Abril, M., Bellosta, P., Gallant, P. & Johnston, L. A. Drosophila myc regulates organ size by inducing cell competition. Cell 117, 107–116 (2004).
Moreno, E. & Basler, K. dMyc transforms cells into super-competitors. Cell 117, 117–129 (2004).
Tamori, Y. et al. Involvement of Lgl and Mahjong/VprBP in cell competition. PLoS Biol. 8, e1000422 (2010).
Karim, F. D. & Rubin, G. M. Ectopic expression of activated Ras1 induces hyperplastic growth and increased cell death in Drosophila imaginal tissues. Development 125, 1–9 (1998).
Brumby, A. M. & Richardson, H. E. scribble mutants cooperate with oncogenic Ras or Notch to cause neoplastic overgrowth in Drosophila. EMBO J. 22, 5769–5779 (2003).
Hogan, C. et al. Characterization of the interface between normal and transformed epithelial cells. Nat. Cell Biol. 11, 460–467 (2009).
Kajita, M. et al. Interaction with surrounding normal epithelial cells influences signalling pathways and behaviour of Src-transformed cells. J. Cell Sci. 123, 171–180 (2010).
Wu, S. K. et al. Cortical F-actin stabilization generates apical-lateral patterns of junctional contractility that integrate cells into epithelia. Nat. Cell Biol. 16, 167–178 (2014).
Leung, C. T. & Brugge, J. S. Outgrowth of single oncogene-expressing cells from suppressive epithelial environments. Nature 482, 410–413 (2012).
Norman, M. et al. Loss of Scribble causes cell competition in mammalian cells. J. Cell Sci. 125, 59–66 (2012).
Kajita, M. et al. Filamin acts as a key regulator in epithelial defence against transformed cells. Nat. Commun. 5, 4428 (2014).
Ohoka, A. et al. EPLIN is a crucial regulator for extrusion of RasV12-transformed cells. J. Cell Sci. 128, 781–789 (2015).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Sciacovelli, M., Gaude, E., Hilvo, M. & Frezza, C. The metabolic alterations of cancer cells. Methods Enzymol. 542, 1–23 (2014).
Koppenol, W. H., Bounds, P. L. & Dang, C. V. Otto Warburg’s contributions to current concepts of cancer metabolism. Nat. Rev. Cancer 11, 325–337 (2011).
Cairns, R. A., Harris, I. S. & Mak, T. W. Regulation of cancer cell metabolism. Nat. Rev. Cancer 11, 85–95 (2011).
Warburg, O. On the origin of cancer cells. Science 123, 309–314 (1956).
Levayer, R., Hauert, B. & Moreno, E. Cell mixing induced by myc is required for competitive tissue invasion and destruction. Nature 524, 476–480 (2015).
Kato, M., Li, J., Chuang, J. L. & Chuang, D. T. Distinct structural mechanisms for inhibition of pyruvate dehydrogenase kinase isoforms by AZD7545, dichloroacetate, and radicicol. Structure 15, 992–1004 (2007).
Wynn, R. M. et al. Pyruvate dehydrogenase kinase-4 structures reveal a metastable open conformation fostering robust core-free basal activity. J. Biol. Chem. 283, 25305–25315 (2008).
Sutendra, G. et al. Mitochondrial activation by inhibition of PDKII suppresses HIF1a signaling and angiogenesis in cancer. Oncogene 32, 1638–1650 (2013).
Koukourakis, M. I. et al. Pyruvate dehydrogenase and pyruvate dehydrogenase kinase expression in non small cell lung cancer and tumor-associated stroma. Neoplasia 7, 1–6 (2005).
Hur, H. et al. Expression of pyruvate dehydrogenase kinase-1 in gastric cancer as a potential therapeutic target. Int. J. Oncol. 42, 44–54 (2013).
Lu, C. W. et al. Overexpression of pyruvate dehydrogenase kinase 3 increases drug resistance and early recurrence in colon cancer. Am. J. Pathol. 179, 1405–1414 (2011).
Chu, Q. S. et al. A phase I open-labeled, single-arm, dose-escalation, study of dichloroacetate (DCA) in patients with advanced solid tumors. Invest. New Drugs 33, 603–610 (2015).
Dunbar, E. M. et al. Phase 1 trial of dichloroacetate (DCA) in adults with recurrent malignant brain tumors. Invest. New Drugs 32, 452–464 (2014).
Strum, S. B. et al. Case report: sodium dichloroacetate (DCA) inhibition of the “Warburg Effect” in a human cancer patient: complete response in non-Hodgkin’s lymphoma after disease progression with rituximab-CHOP. J. Bioenerg. Biomembr. 45, 307–315 (2013).
Garon, E. B. et al. Dichloroacetate should be considered with platinum-based chemotherapy in hypoxic tumors rather than as a single agent in advanced non-small cell lung cancer. J. Cancer Res. Clin. Oncol. 140, 443–452 (2014).
Kami, K. et al. Metabolomic profiling of lung and prostate tumor tissues by capillary electrophoresis time-of-flight mass spectrometry. Metabolomics 9, 444–453 (2013).
Shestov, A. A. et al. Quantitative determinants of aerobic glycolysis identify flux through the enzyme GAPDH as a limiting step. eLife 3, e03342 (2014).
Tang, X. et al. A joint analysis of metabolomics and genetics of breast cancer. Breast Cancer Res. 16, 415 (2014).
Pfeiffer, T., Schuster, S. & Bonhoeffer, S. Cooperation and competition in the evolution of ATP-producing pathways. Science 292, 504–507 (2001).
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009).
Imajo, M., Ebisuya, M. & Nishida, E. Dual role of YAP and TAZ in renewal of the intestinal epithelium. Nat. Cell Biol. 17, 7–19 (2015).
Semenza, G. L., Roth, P. H., Fang, H. M. & Wang, G. L. Transcriptional regulation of genes encoding glycolytic enzymes by hypoxia-inducible factor 1. J. Biol. Chem. 269, 23757–23763 (1994).
Gordan, J. D., Thompson, C. B. & Simon, M. C. HIF and c-Myc: sibling rivals for control of cancer cell metabolism and proliferation. Cancer Cell 12, 108–113 (2007).
Kim, J. W., Tchernyshyov, I., Semenza, G. L. & Dang, C. V. HIF-1-mediated expression of pyruvate dehydrogenase kinase: a metabolic switch required for cellular adaptation to hypoxia. Cell Metab. 3, 177–185 (2006).
Papandreou, I., Cairns, R. A., Fontana, L., Lim, A. L. & Denko, N. C. HIF-1 mediates adaptation to hypoxia by actively downregulating mitochondrial oxygen consumption. Cell Metab. 3, 187–197 (2006).
Kluza, J. et al. Inactivation of the HIF-1alpha/PDK3 signaling axis drives melanoma toward mitochondrial oxidative metabolism and potentiates the therapeutic activity of pro-oxidants. Cancer Res. 72, 5035–5047 (2012).
Atsumi, T. et al. High expression of inducible 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase (iPFK-2; PFKFB3) in human cancers. Cancer Res. 62, 5881–5887 (2002).
Christofk, H. R. et al. The M2 splice isoform of pyruvate kinase is important for cancer metabolism and tumour growth. Nature 452, 230–233 (2008).
Wu, W. & Zhao, S. Metabolic changes in cancer: beyond the Warburg effect. Acta Biochim. Biophys. Sin. 45, 18–26 (2013).
Semenza, G. L. HIF-1: upstream and downstream of cancer metabolism. Curr. Opin. Genet. Dev. 20, 51–56 (2010).
de la Cova, C. et al. Supercompetitor status of Drosophila Myc cells requires p53 as a fitness sensor to reprogram metabolism and promote viability. Cell Metab. 19, 470–483 (2014).
Hou, B. H. et al. Optical sensors for monitoring dynamic changes of intracellular metabolite levels in mammalian cells. Nat. Protoc. 6, 1818–1833 (2011).
Hsu, P. D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Maruyama, T. et al. Corrigendum: increasing the efficiency of precise genome editing with CRISPR-Cas9 by inhibition of nonhomologous end joining. Nat. Biotechnol. 34, 210 (2016).
el Marjou, F. et al. Tissue-specific and inducible Cre-mediated recombination in the gut epithelium. Genesis 39, 186–193 (2004).
Kawamoto, S. et al. A novel reporter mouse strain that expresses enhanced green fluorescent protein upon Cre-mediated recombination. FEBS Lett. 470, 263–268 (2000).
Perry, S. W., Norman, J. P., Barbieri, J., Brown, E. B. & Gelbard, H. A. Mitochondrial membrane potential probes and the proton gradient: a practical usage guide. BioTechniques 50, 98–115 (2011).
Hogan, C. et al. Rap1 regulates the formation of E-cadherin-based cell-cell contacts. Mol. Cell Biol. 24, 6690–6700 (2004).
Yamamoto, S. et al. A role of the sphingosine-1-phosphate (S1P)-S1P receptor 2 pathway in epithelial defense against cancer (EDAC). Mol. Biol. Cell 27, 491–499 (2016).
Imamura, H. et al. Visualization of ATP levels inside single living cells with fluorescence resonance energy transfer-based genetically encoded indicators. Proc. Natl Acad. Sci. USA 106, 15651–15656 (2009).
Acknowledgements
We thank K. Rajewsky for the establishment of knocked-in ES cells harbouring RasV12-eGFP. We also thank H. Harada for the HIF1 reporter-expressing vector, C. Kuo for the R-spondin-producing cell line and T. Yoshimori for useful advice on EM analyses. Y.Fujita is supported by Japan Society for the Promotion of Science (JSPS) Grant-in-Aid for Scientific Research on Innovative Areas 26114001, Grant-in-Aid for Scientific Research (A) 26250026 and the AMED Strategic Japanese–Swiss Cooperative Program. Y.Fujita is also supported by the Takeda Science Foundation. S.Kon is supported by Japan Society for the Promotion of Science (JSPS) Grant-in-Aid for Scientific Research on Innovative Areas 26112701, the Kato Memorial Bioscience Foundation and The YASUDA Medical Foundation.
Author information
Authors and Affiliations
Contributions
S.Kon designed experiments and generated most of the data. K.I. carried out qrtPCR experiments and metabolic analyses. H.K. and S.Kitamoto designed and analysed the cell-competition mouse model. S.I., H.Y., Y.Y., T.Matsumoto, H.W., R.Egami, A.N., I.K., S.T., R.N. and T.Maruyama assisted in experiments. M.K., A.S., R.Enoki, S.H., S.Y., J.K. and M.Ishii carried out time-lapse experiments. T.K. carried out electron microscopic analyses. J.-M.N., Y.Onodera, Y.Fujioka and Y.Ohba carried out FRET analysis. H.I., T.Soga and M.R.D. assisted in metabolic analyses. Y.S., M.O., J.-i.M. and T.Sato assisted in ex vivo and in vivo experiments. T.Shirai, T.Morita, M.Imajo and E.N. assisted in iGT experiments. Y.Fujita conceived and designed the study. The manuscript was written by S.Kon and Y.Fujita with assistance from the other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 The mitochondrial membrane potential is decreased in Src-transformed cells but not in Scribble-knockdown cells when they are surrounded by normal cells.
(a) TMRM incorporation in Src-transformed cells. MDCK-pTR cSrcY527F-GFP cells were mixed with normal MDCK cells or cultured alone, and loaded with 50 nM TMRM (red). Arrows indicate Src-transformed cells showing diminished fluorescence intensity of TMRM. (b) TMRM incorporation in Scribble-knockdown cells. MDCK-pTR Scribble shRNA cells were mixed with normal MDCK cells or cultured alone, and incubated with tetracycline for 48 h and loaded with 50 nM TMRM (red). Arrows indicate Scribble-knockdown cells showing the comparable fluorescence intensity of TMRM to that in the surrounding normal cells. (c) Establishment of doxycycline-inducible myc-RasV12 MDCK cell lines. Doxycline-induced expression of myc-RasV12 protein is determined by western blotting. (d) Immunofluorescence images of xz sections of myc-RasV12 MDCK cells surrounded by normal MDCK cells. (e) Quantification of the apical extrusion of myc-RasV12 cells. Data are mean ± s.e.m. n = 2 independent experiments. These data demonstrate that myc-RasV12 cells are apically extruded when surrounded by normal cells, similarly for GFP-RasV12 cells. (f) TMRM incorporation in myc-RasV12 cells. MDCK-pTRE3G myc-RasV12 cells were fluorescently labelled with CMFDA dye (green), and co-cultured with normal MDCK cells or cultured alone, and loaded with 50 nM TMRM (red). This result shows that non-cell-autonomous reduction of TMRM incorporation also occurs in myc-RasV12 cells. (g) Immunofluorescence and ratiometric images of MitoTracker Green (MTG) and TMRM. MDCK-pTRE3G myc-RasV12 cells were fluorescently labelled with CMAC dye (blue), and co-cultured with normal MDCK cells or cultured alone. Cells were incubated with 200 nM MTG for 2 h, washed briefly and subsequently loaded with 50 nM TMRM for 30 min. Scale bars, 10 μm (a,b,d,f,g). (h) Quantification of the ratio of TMRM to MTG. Data are mean ± s.e.m. ∗P < 0.001, unpaired two-tailed t-test; n = 50 and 36 cells pooled from three independent experiments. For Supplementary Fig. 1c, unprocessed original scans of immunoblotting are also shown in Supplementary Fig. 9. Statistics source data for e,h are provided in Supplementary Table 2.
Supplementary Figure 2 Mitophagosome-like structures are frequently observed in RasV12 cells that are surrounded by normal epithelial cells.
Electron microscopic images of MDCK-pTR GFP-RasV12 cells cultured alone or surrounded by normal cells. The areas in the white boxes are shown below at higher magnification, demonstrating mitophagosome-like structures. In RasV12 cells surrounded by normal cells, on average 0.65 mitophagosome-like structures in a single RasV12 cell per slice (51 slices); in RasV12 cells cultured alone, on average 0.01 mitophagosome-like structures in a single RasV12 cell per slice (84 slices). Scale bars, 2 μm (upper panels) and 0.5 μm (lower panels).
Supplementary Figure 3 Knockdown/knockout of PDK4 inhibits PDH phosphorylation and LDHA accumulation in RasV12-transformed cells surrounded by normal cells.
(a) The illustration for the mode of action of PDK and LDHA. (b) A targeting scheme and DNA sequences of the wild type and PDK4-null MDCK-pTRE3G GFP-RasV12 cell lines. PAM motifs are underlined. The red spacing indicates a deleted nucleotide. (c–n) Effect of PDK4-knockdown or-knockout on p-PDH (c–e), TMRM incorporation (f,g), LDHA (h–j) or apical extrusion (k–n). MDCK-pTR GFP-RasV12 PDK4 shRNA cells (c,h), MDCK-pTR GFP-RasV12 Luciferase shRNA cells (d,f,i) or MDCK-pTRE3G GFP-RasV12 PDK4-sgRNA cells (e,g,j) were mixed with normal MDCK cells or MDCK-pTR Luciferase shRNA cells. Cells were loaded with 50 nM TMRM (f,g) or stained with Hoechst 33342 (blue) and anti-p-PDH (c–e) or anti-LDHA (h–j) antibody (red). (g) Note that PDK4-knockout rather promoted TMRM incorporation in RasV12 cells surrounded by normal cells. (k) Immunofluorescence images of xz sections of MDCK-pTR GFP-RasV12 Luciferase-shRNA cells surrounded by normal MDCK cells or of MDCK-pTR GFP-RasV12 cells surrounded by MDCK-pTR Lucifierase-shRNA cells. (l) Quantification of apical extrusion of RasV12 cells mixed with MDCK cells, RasV12 Luc-shRNA cells mixed with MDCK cells or RasV12 cells mixed with MDCK Luc-shRNA cells. Data are mean ± s.e.m. ∗P < 0.05, unpaired two-tailed t-test; n = 3 independent experiments. (m) Immunofluorescence images of xz sections of MDCK-pTRE3G GFP-RasV12 PDK4-sgRNA1 or-sgRNA2 cells surrounded by normal MDCK cells. Arrowheads indicate basal protrusions. (n) Quantification of the apical extrusion of MDCK-pTRE3G GFP-RasV12 PDK4-sgRNA1 or-sgRNA2 cells. Data are mean ± s.e.m. Values are expressed as a ratio relative to MDCK:RasV12 = 50:1. ∗P < 0.05, unpaired two-tailed t-test; n = 3 independent experiments. Scale bars, 10 μm (c–k,m). Statistics source data for l,n are provided in Supplementary Table 2.
Supplementary Figure 4 PDK inhibitor Radicicol restores TMRM incorporation and suppreses apical extrusion of RasV12-transformed cells surrounded by normal cells.
(a) Effect of Radicicol on TMRM incorporation of RasV12-transformed cells. MDCK-pTR GFP-RasV12 cells were co-cultured with normal MDCK cells in the absence or presence of 10 μM Radicicol and incubated with 50 nM TMRM. (b) Immunofluorescence images of xz sections of MDCK-pTR GFP-RasV12 cells surrounded by normal MDCK cells in the absence or presence of Radicicol. (c) Quantification of the effect of Radicicol on apical extrusion. Data are mean ± s.e.m. ∗P < 0.01, unpaired two-tailed t-test; n = 3 independent experiments. (d) Effect of Radicicol on LDHA in RasV12-transformed cells that are surrounded by normal cells. MDCK-pTR GFP-RasV12 cells were mixed with normal MDCK cells in the absence or presence of Radicicol. Cells were stained with Hoechst 33342 (blue) and anti-LDHA antibody (red). (e) Establishment of MDCK-pTR GFP-RasV12 cells stably expressing LDHA-shRNA1 or-shRNA2. Knockdown of LDHA is confirmed by western blotting. (f) Effect of LDHA-knockdown on TMRM incorporation. MDCK-pTR GFP-RasV12 LDHA-shRNA1 or-shRNA2 cells were mixed with normal MDCK cells and loaded with 50 nM TMRM. (g) Quantification of the apical extrusion of MDCK-pTR GFP-RasV12 LDHA-shRNA1 or-shRNA2 cells. Data are mean ± s.e.m. ∗P < 0.05, unpaired two-tailed t-test; n = 3 independent experiments. (h) Establishment of tetracycline-inducible PDH-knockdown MDCK cell lines. Effect of tetracycline on expression of PDH protein is determined by western blotting. (i) TMRM incorporation in PDH-knockdown cells. MDCK-pTR PDH-shRNA1 or-shRNA2 cells were fluorescently labelled with CMFDA dye (green), and co-cultured with normal MDCK cells. Cells were incubated with tetracycline for 48 h, and loaded with 50 nM TMRM (red). (j) Immunofluorescence images of xz sections of MDCK-pTR PDH-shRNA cells surrounded by normal MDCK cells at 72 h after tetracycline addition. Scale bars, 10 μm (a,b,d,f,i,j). For Supplementary Fig. 4e, h, unprocessed original scans of immunoblotting are also shown in Supplementary Fig. 9. Statistics source data for c,g are provided in Supplementary Table 2.
Supplementary Figure 5 Effect of various inhibitors on TMRM incorporation and FRET analyses for ATP.
(a,b) Effect of EPLIN- or Filamin-knockdown on TMRM incorporation (a) or p-PDH and LDHA accumulation (b). MDCK-pTR GFP-RasV12 or MDCK-pTR GFP-RasV12 EPLIN-shRNA2 cells were co-cultured with MDCK or MDCK-pTR Filamin-shRNA2 cells. (c) MDCK-pTR GFP-RasV12 cells were co-cultured with normal MDCK cells in the presence of various inhibitors and loaded with TMRM (red). Each inhibitor inhibits the following molecule or cellular process; Blebbistatin: myosin-II, Cytochalasin D: actin polymerization, Y27632: Rho kinase, NAC: reactive oxygen species, L-NAME: nitrogen oxide synthase, 3-MA: autophagy, KT5720: PKA. (d) Schematics for ATP-FRET (ATeam). (e) ATP-FRET images. MDCK or MDCK-pTRE3G myc-RasV12 cells transiently expressing ATeam were stained with CMTPX and co-cultured with MDCK cells or cultured alone with doxycycline for 16 h in the absence or presence of DCA or 2-DG and then analysed by dual-emission fluorescence microscopy. FRET/CFP ratio images were generated to represent FRET efficiency. (f) Quantification of ATP-FRET efficiency ratio (FRET/CFP). The box plots represent values from the 25th (bottom) to the 75th (top) percentiles, with the median as the horizontal line. ∗P < 0.001, unpaired two-tailed t-test; n = 50, 34, 32, 32, 15 and 15 cells pooled from two independent experiments. (g) Effect of 2-DG on lactate production in RasV12-transformed cells. MDCK-pTRE3G myc-RasV12 cells were incubated with the indicated concentration of 2-DG for 24 h, and lactate concentration in the culture media was measured. Data are mean ± s.e.m. from three independent experiments. (h,i) Effect of 2-DG on apical extrusion of RasV12-transformed cells. MDCK-pTR GFP-RasV12 cells were co-cultured with normal MDCK cells in the absence or presence of 25 mM 2-DG. (h) Immunofluorescence images of xz sections of MDCK-pTR GFP-RasV12 cells surrounded by normal MDCK cells in the absence or presence of 2-DG. (i) Quantification of the effect of 2-DG on apical extrusion. Data are mean ± s.e.m. ∗P < 0.01, unpaired two-tailed t-test; n = 3 independent experiments. Scale bars, 10 μm (a–c,e,h). Statistics source data for f,g,i are provided in Supplementary Table 2.
Supplementary Figure 6 Establishment of conditional RasV12-GFP knock-in mice and tamoxifen-dependent expression of transgenes in intestinal organoids.
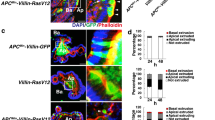
(a) Physical maps of the DNMT1 gene locus and its targeting vector. The targeting vector contains a cDNA encoding H-RasV12 under control of the CAG promoter and downstream of a loxP-flanked stop cassette plus a frt-flanked IRES-eGFP component. Filled-in and open arrowheads indicate the loxP sequences and frt sequences, respectively. The grey rectangles under the line correspond to the probes that were used for Southern blot hybridization. Arrows denote PCR primer sites for the genotyping. (b) Southern blot analysis of genomic DNA prepared from the wild-type ES cells and the targeted ES clones. DNA was digested by BglII (probe 1) or SacI (probe 2) and processed for Southern blotting using the hybridization probes shown in (a). (c) Organoids from Villin-CreERT2;LSL-RasV12-IRES-eGFP mice or Villin-CreERT2;LSL-eGFP mice were incubated with the indicated concentration of tamoxifen for 24 h and stained with Hoechst 33342 (blue). Ba and Ap stand for the basal and apical side, respectively. Asterisks indicate mucin-rich, autofluorescent materials in the apical lumen. Scale bars, 10 μm. The percentage of RasV12-GFP- or GFP-expressing cells relative to total cells in an organoid was quantified. Data are mean ± s.e.m. n = 2 independent experiments. Statistics source data for c are provided in Supplementary Table 2.
Supplementary Figure 7 Cropped images of the time-lapse observation for apically extruding RasV12 cells in intestinal organoids.
(a) Immunofluorescence of EPLIN in intestinal organoids from villin-CreERT2;LSL-RasV12-IRES-eGFP mice at 24 h after treatment of 100 nM or 1 μM tamoxifen. The organoids were stained with Hoechst 33342 (blue) and anti-EPLIN antibody (red). Scale bars, 10 μm. (b) Representative serial images of time-lapse microscopic observation for intestinal organoids from Villin-CreERT2;LSL-eGFP mice (upper panels) or Villin-CreERT2;LSL-RasV12-IRES-eGFP mice (middle and lower panels) at the indicated times after treatment of 100 nM tamoxifen. Note that apical extrusion of RasV12-transformed cells took variable times between 2–20 h. Scale bars, 50 μm. (c) Effect of DCA on TMRM incorporation ex vivo. The intestinal organoids from Villin-CreERT2;LSL-RasV12-IRES-eGFP mice at 24 h after treatment of 100 nM tamoxifen in the absence or presence of 50 mM DCA were loaded with 50 nM TMRM (red). Scale bars, 10 μm. Asterisks indicate mucin-rich, autofluorescent materials in the apical lumen (a,c). Ba and Ap stand for the basal and apical side, respectively (a–c). (d) Quantification of the fluorescence intensity of TMRM. Data are mean ± s.e.m. Values are expressed as a ratio relative to normal DCA (-). ∗P < 0.01, unpaired two-tailed t-test; n = 24, 25, 45 and 47 cells pooled from three independent experiments. (e) HIF1 reporter assay. MDCK cells or MDCK-pTR GFP-RasV12 cells were co-transfected with the HIF1 reporter and Renilla expression vector, and co-cultured with non-transfected MDCK cells or MDCK-pTR GFP-RasV12 cells in the absence or presence of 100 μM HIF1 inhibitor Chrysin. After 16 h incubation with tetracycline, cells were lysed and subjected to measurement of the luciferase activity. Data are mean ± s.e.m. ∗P < 0.05, unpaired two-tailed t-test; n = 3 independent experiments. Statistics source data for d,e are provided in Supplementary Table 2.
Supplementary Figure 8 A schematic model for molecular mechanisms of the Warburg effect-like metabolic changes in transformed cells that are surrounded by normal cells.
(a) Effect of various inhibitors on apical extrusion of RasV12-transformed cells and on TMRM incorporation in RasV12-transformed cells that are surrounded by normal cells. *: statistically significant (unpaired two-tailed t-test); ND: not done; Grey box: our published observations10,11,15,16. (b) Difference between conventional Warburg effect and EDAC-induced Warburg effect-like metabolic changes. (c) A schematic model for molecular mechanisms of the Warburg effect-like metabolic changes in transformed cells that are surrounded by normal cells.
Supplementary Figure 9 Unprocessed scans of original blots.
Uncropped scans of images for all immunoblotting experiments are shown.
Supplementary information
Supplementary Information
Supplementary Information (PDF 8465 kb)
Supplementary Table 1
Supplementary Information (XLSX 16 kb)
Supplementary Table 2
Supplementary Information (XLSX 95 kb)
GFP-expressing cells in the intestinal organoid epithelium.
This movie shows that GFP-expressing cells are not extruded from the intestinal organoid epithelium which is derived from Villin-CreERT2;LSL-eGFP mice. Time-lapse images were captured at 15-min intervals for 24 h. (MOV 212 kb)
RasV12-GFP-expressing cells in the intestinal organoid epithelium.
This movie shows that RasV12-GFP-expressing cells are apically extruded from the intestinal organoid epithelium which is derived from Villin-CreERT2;LSL-RasV12-IRES-eGFP mice. Time-lapse images were captured at 15-min intervals for 24 h. (MOV 447 kb)
RasV12-GFP-expressing cells in the intestinal organoid epithelium.
This movie shows that RasV12-GFP-expressing cells are apically extruded from the intestinal organoid epithelium which is derived from Villin-CreERT2;LSL-RasV12-IRES-eGFP mice. Time-lapse images were captured at 15-min intervals for 24 h. (MOV 743 kb)
RasV12-GFP-expressing cells in the intestinal villi in vivo.
This intravital movie by two-photon imaging shows that RasV12-GFP-expressing cells are apically extruded from the intestinal villi which is derived from Villin-CreERT2;LSL-RasV12-IRES-eGFP mice. Time-lapse images were captured at 15-min intervals for 75 min. (MOV 547 kb)
Rights and permissions
About this article
Cite this article
Kon, S., Ishibashi, K., Katoh, H. et al. Cell competition with normal epithelial cells promotes apical extrusion of transformed cells through metabolic changes. Nat Cell Biol 19, 530–541 (2017). https://doi.org/10.1038/ncb3509
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3509
This article is cited by
-
Epithelial recognition and elimination against aberrant cells
Seminars in Immunopathology (2024)
-
Inhibition of Pyruvate Dehydrogenase Kinase 4 Protects Cardiomyocytes from lipopolysaccharide-Induced Mitochondrial Damage by Reducing Lactate Accumulation
Inflammation (2024)
-
Wnt activation disturbs cell competition and causes diffuse invasion of transformed cells through NF-κB-MMP21 pathway
Nature Communications (2023)
-
Biophysics in tumor growth and progression: from single mechano-sensitive molecules to mechanomedicine
Oncogene (2023)
-
Cell competition in development, homeostasis and cancer
Nature Reviews Molecular Cell Biology (2023)