Abstract
Primordial germ cells (PGCs) in many species associate intimately with endodermal cells, but the significance of such interactions is largely unexplored. Here, we show that Caenorhabditis elegans PGCs form lobes that are removed and digested by endodermal cells, dramatically altering PGC size and mitochondrial content. We demonstrate that endodermal cells do not scavenge lobes PGCs shed, but rather, actively remove lobes from the cell body. CED-10 (Rac)-induced actin, DYN-1 (dynamin) and LST-4 (SNX9) transiently surround lobe necks and are required within endodermal cells for lobe scission, suggesting that scission occurs through a mechanism resembling vesicle endocytosis. These findings reveal an unexpected role for endoderm in altering the contents of embryonic PGCs, and define a form of developmentally programmed cell remodelling involving intercellular cannibalism. Active roles for engulfing cells have been proposed in several neuronal remodelling events, suggesting that intercellular cannibalism may be a more widespread method used to shape cells than previously thought.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Cinalli, R. M., Rangan, P. & Lehmann, R. Germ cells are forever. Cell 132, 559–562 (2008).
Seydoux, G. & Braun, R. E. Pathway to totipotency: lessons from germ cells. Cell 127, 891–904 (2006).
Molyneaux, K. A., Stallock, J., Schaible, K. & Wylie, C. Time-lapse analysis of living mouse germ cell migration. Dev. Biol. 240, 488–498 (2001).
Takamura, K., Fujimura, M. & Yamaguchi, Y. Primordial germ cells originate from the endodermal strand cells in the ascidian Ciona intestinalis. Dev. Genes Evol. 212, 11–18 (2002).
Chihara, D. & Nance, J. An E-cadherin-mediated hitchhiking mechanism for C. elegans germ cell internalization during gastrulation. Development 139, 2547–2556 (2012).
DeGennaro, M. et al. Peroxiredoxin stabilization of DE-cadherin promotes primordial germ cell adhesion. Dev. Cell 20, 233–243 (2011).
Sulston, J. E., Schierenberg, E., White, J. G. & Thomson, J. N. The embryonic cell lineage of the nematode Caenorhabditis elegans. Dev. Biol. 100, 64–119 (1983).
Rohrschneider, M. R. & Nance, J. The union of somatic gonad precursors and primordial germ cells during Caenorhabditis elegans embryogenesis. Dev. Biol. 379, 139–151 (2013).
Owraghi, M., Broitman-Maduro, G., Luu, T., Roberson, H. & Maduro, M. F. Roles of the Wnt effector POP-1/TCF in the C. elegans endomesoderm specification gene network. Dev. Biol. 340, 209–221 (2010).
Strome, S. & Wood, W. B. Immunofluorescence visualization of germ-line-specific cytoplasmic granules in embryos, larvae, and adults of Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 79, 1558–1562 (1982).
Head, B. P., Zulaika, M., Ryazantsev, S. & van der Bliek, A. M. A novel mitochondrial outer membrane protein, MOMA-1, that affects cristae morphology in Caenorhabditis elegans. Mol. Biol. Cell 22, 831–841 (2011).
Rolland, S. G., Lu, Y., David, C. N. & Conradt, B. The BCL-2-like protein CED-9 of C. elegans promotes FZO-1/Mfn1,2- and EAT-3/Opa1-dependent mitochondrial fusion. J. Cell Biol. 186, 525–540 (2009).
Yang, W. & Hekimi, S. A mitochondrial superoxide signal triggers increased longevity in Caenorhabditis elegans. PLoS Biol. 8, e1000556 (2010).
Chen, B. C. et al. Lattice light-sheet microscopy: imaging molecules to embryos at high spatiotemporal resolution. Science 346, 1257998 (2014).
Reddien, P. W. & Horvitz, H. R. CED-2/CrkII and CED-10/Rac control phagocytosis and cell migration in Caenorhabditis elegans. Nat. Cell Biol. 2, 131–136 (2000).
Lundquist, E. A., Reddien, P. W., Hartwieg, E., Horvitz, H. R. & Bargmann, C. I. Three C. elegans Rac proteins and several alternative Rac regulators control axon guidance, cell migration and apoptotic cell phagocytosis. Development 128, 4475–4488 (2001).
Kinchen, J. M. et al. Two pathways converge at CED-10 to mediate actin rearrangement and corpse removal in C. elegans. Nature 434, 93–99 (2005).
Wang, X. & Yang, C. Programmed cell death and clearance of cell corpses in Caenorhabditis elegans. Cell Mol. Life Sci. 73, 2221–2236 (2016).
Shin, N. et al. SNX9 regulates tubular invagination of the plasma membrane through interaction with actin cytoskeleton and dynamin 2. J. Cell Sci. 121, 1252–1263 (2008).
Lundmark, R. & Carlsson, S. R. Sorting nexin 9 participates in clathrin-mediated endocytosis through interactions with the core components. J. Biol. Chem. 278, 46772–46781 (2003).
McMahon, H. T. & Boucrot, E. Molecular mechanism and physiological functions of clathrin-mediated endocytosis. Nat. Rev. Mol. Cell Biol. 12, 517–533 (2011).
Almendinger, J. et al. A conserved role for SNX9-family members in the regulation of phagosome maturation during engulfment of apoptotic cells. PLoS ONE 6, e18325 (2011).
Chen, D. et al. Clathrin and AP2 are required for phagocytic receptor-mediated apoptotic cell clearance in Caenorhabditis elegans. PLoS Genet. 9, e1003517 (2013).
Cheng, S. et al. PtdIns(4,5)P(2) and PtdIns3P coordinate to regulate phagosomal sealing for apoptotic cell clearance. J. Cell Biol. 210, 485–502 (2015).
Lu, N., Shen, Q., Mahoney, T. R., Liu, X. & Zhou, Z. Three sorting nexins drive the degradation of apoptotic cells in response to PtdIns(3)P signaling. Mol. Biol. Cell 22, 354–374 (2011).
Yu, X., Odera, S., Chuang, C. H., Lu, N. & Zhou, Z. C. elegans dynamin mediates the signaling of phagocytic receptor CED-1 for the engulfment and degradation of apoptotic cells. Dev. Cell 10, 743–757 (2006).
Lundmark, R. & Carlsson, S. R. Regulated membrane recruitment of dynamin-2 mediated by sorting nexin 9. J. Biol. Chem. 279, 42694–42702 (2004).
Soulet, F., Yarar, D., Leonard, M. & Schmid, S. L. SNX9 regulates dynamin assembly and is required for efficient clathrin-mediated endocytosis. Mol. Biol. Cell 16, 2058–2067 (2005).
Yarar, D., Waterman-Storer, C. M. & Schmid, S. L. SNX9 couples actin assembly to phosphoinositide signals and is required for membrane remodeling during endocytosis. Dev. Cell 13, 43–56 (2007).
Badour, K. et al. Interaction of the Wiskott-Aldrich syndrome protein with sorting nexin 9 is required for CD28 endocytosis and cosignaling in T cells. Proc. Natl Acad. Sci. USA 104, 1593–1598 (2007).
Taylor, M. J., Perrais, D. & Merrifield, C. J. A high precision survey of the molecular dynamics of mammalian clathrin-mediated endocytosis. PLoS Biol. 9, e1000604 (2011).
Baugh, L. R. To grow or not to grow: nutritional control of development during Caenorhabditis elegans L1 arrest. Genetics 194, 539–555 (2013).
Fukuyama, M. et al. C. elegans AMPKs promote survival and arrest germline development during nutrient stress. Biol. Open. 1, 929–936 (2012).
Wolf, T., Qi, W., Schindler, V., Runkel, E. D. & Baumeister, R. Doxycyclin ameliorates a starvation-induced germline tumor in C. elegans daf-18/PTEN mutant background. Exp. Gerontol. 56, 114–122 (2014).
Wolke, U., Jezuit, E. A. & Priess, J. R. Actin-dependent cytoplasmic streaming in C. elegans oogenesis. Development 134, 2227–2236 (2007).
Lei, L. & Spradling, A. C. Mouse oocytes differentiate through organelle enrichment from sister cyst germ cells. Science 352, 95–99 (2016).
Cox, R. T. & Spradling, A. C. A Balbiani body and the fusome mediate mitochondrial inheritance during Drosophila oogenesis. Development 130, 1579–1590 (2003).
Chu, D. S. & Shakes, D. C. Spermatogenesis. Adv. Exp. Med. Biol. 757, 171–203 (2013).
Kevany, B. M. & Palczewski, K. Phagocytosis of retinal rod and cone photoreceptors. Physiology (Bethesda) 25, 8–15 (2010).
Corty, M. M. & Freeman, M. R. Cell biology in neuroscience: architects in neural circuit design: glia control neuron numbers and connectivity. J. Cell Biol. 203, 395–405 (2013).
Schafer, D. P. et al. Microglia sculpt postnatal neural circuits in an activity and complement-dependent manner. Neuron 74, 691–705 (2012).
Han, C. et al. Epidermal cells are the primary phagocytes in the fragmentation and clearance of degenerating dendrites in Drosophila. Neuron 81, 544–560 (2014).
Wehman, A. M., Poggioli, C., Schweinsberg, P., Grant, B. D. & Nance, J. The P4-ATPase TAT-5 inhibits the budding of extracellular vesicles in C. elegans embryos. Curr. Biol. 21, 1951–1959 (2011).
Treusch, S. et al. Caenorhabditis elegans functional orthologue of human protein h-mucolipin-1 is required for lysosome biogenesis. Proc. Natl Acad. Sci. USA 101, 4483–4488 (2004).
Shirayama, M. et al. piRNAs initiate an epigenetic memory of nonself RNA in the C. elegans germline. Cell 150, 65–77 (2012).
Chen, C. C. RAB-10 is required for endocytic recycling in the Caenorhabditis elegans intestine. Mol. Biol. Cell 17, 1286–1297 (2006).
Kamath, R. S., Martinez-Campos, M., Zipperlen, P., Fraser, A. G. & Ahringer, J. Effectiveness of specific RNA-mediated interference through ingested double-stranded RNA in Caenorhabditis elegans. Genome Biol. 2, RESEARCH0002 (2001).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Minevich, G., Park, D. S., Blankenberg, D., Poole, R. J. & Hobert, O. CloudMap: a cloud-based pipeline for analysis of mutant genome sequences. Genetics 192, 1249–1269 (2012).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform. 14, 178–192 (2013).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Nance, J., Munro, E. M. & Priess, J. R. C. elegans PAR-3 and PAR-6 are required for apicobasal asymmetries associated with cell adhesion and gastrulation. Development 130, 5339–5350 (2003).
Frokjaer-Jensen, C. et al. Single-copy insertion of transgenes in Caenorhabditis elegans. Nat. Genet. 40, 1375–1383 (2008).
Armenti, S. T., Chan, E. & Nance, J. Polarized exocyst-mediated vesicle fusion directs intracellular lumenogenesis within the C. elegans excretory cell. Dev. Biol. 394, 110–121 (2014).
Mello, C. C., Kramer, J. M., Stinchcomb, D. & Ambros, V. Efficient gene transfer in C. elegans extrachromosomal maintenance and integration of transforming sequences. EMBO J. 10, 3959–3970 (1991).
Strange, K., Christensen, M. & Morrison, R. Primary culture of Caenorhabditis elegans developing embryo cells for electrophysiological, cell biological and molecular studies. Nat. Protoc. 2, 1003–1012 (2007).
Lynch, A. M. et al. A genome-wide functional screen shows MAGI-1 is an L1CAM-dependent stabilizer of apical junctions in C. elegans. Curr. Biol. 22, 1891–1899 (2012).
Green, R. A. et al. The midbody ring scaffolds the abscission machinery in the absence of midbody microtubules. J. Cell Biol. 203, 505–520 (2013).
Fukushige, T., Goszczynski, B., Yan, J. & McGhee, J. D. Transcriptional control and patterning of the pho-1 gene, an essential acid phosphatase expressed in the C. elegans intestine. Dev. Biol. 279, 446–461 (2005).
Acknowledgements
We thank the Caenorhabditis Genetics Center (CGC), National Bioresource Project (NBRP), and Z. Zhou (Baylor College of Medicine, USA) for providing worm strains. Caged rhodamine dextran was a gift from K. Oegema (University of California, San Diego). We thank T. Hurd (NYU School of Medicine, USA) for sharing TMRE dye and mitochondrial discussion; J. H. Choi (NYU School of Medicine, USA) for assistance in developing the technique for cell culture experiments; D. McIntyre (NYU School of Medicine, USA) for designing a nitrogen gas immobilization chamber for embryos; and L. Christiaen (New York University, USA), N. Ringstad (NYU School of Medicine, USA), T. Hurd (NYU School of Medicine, USA) and members of the Nance laboratory for comments on the manuscript. Library preparation and sequencing of genomic DNA samples was performed at the NYULMC Genome Technology Center, which is partially supported by a Cancer Center Support Grant (P30CA016087) at the Laura and Isaac Perlmutter Cancer Center. Funding was provided by the NIH (R35GM118081, R21HD084809 to J.N.), NYSTEM (C029561 to J.N.) and HHMI (J.M.H., T.-L.C). Y.A. is an HHMI International Student Research fellow.
Author information
Authors and Affiliations
Contributions
Y.A. and J.N. designed experiments. C.M. performed and analysed cell culture, MOMA-1 imaging, and end-1 end-3 embryo experiments. Y.A. performed and analysed all other experiments, and J.H. and T.-L.C. assisted with experiments on the lattice light-sheet microscope. Y.A. and J.N. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 PGC lobe digestion by endoderm.
(a–a’) A newly hatched L1 larva (body outlined). PGC debris (arrowheads) can be seen inside endodermal cells (intestinal rings V and VI). Scale bar, 5 μm.
Supplementary Figure 2 Characterization of mitochondrial dyes.
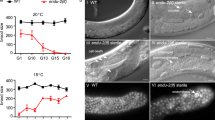
(a–a’) MitoSOX shows stronger labeling in PGCs (one shown, magenta) compared to most somatic cells in the embryo. Asterisk denotes dye accumulation at hole made in eggshell to introduce dye. (b) MitoTracker Green (b’) and MitoSOX (b”) colocalize in embryonic mitochondria. (c) MitoTracker Green (c’) and TMRE (c”) colocalize to mitochondria in embryonic mitochondria. Scale bar, 5 μm.
Supplementary Figure 3 Rhodamine Dextran diffusion from cell bodies into lobes.
(a–b”) Photoactivation of caged Rhodamine Dextran in wild-type L1 larva. Photoactivation in a single PGC (a’,b’) is followed by diffusion into the adjacent PGC (PGCs are connected by a cytoplasmic bridge) (a”, b”, 9/9 embryos). (c–d”) Photoactivation of caged Rhodamine Dextran in end-1 end-3 L1 larva. Photoactivation in a single PGC (c’,d’) is followed by diffusion into persistent lobes and the adjacent PGC (c”, d”, 8/8 embryos). Some caged Rhodamine Dextran becomes uncaged independently of photoactivation, and is visible as stable bright spots in the pre-photoactivation channel. Scale bar, 5 μm.
Supplementary Figure 4 ced-10 in lobe scission.
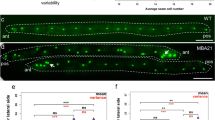
(a–a’) ced-10(n1993) mutant L1 larva with mosaic rescue by ced-10(+)END. The rescuing ced-10(+)END extrachromosomal array, which also expresses nuclear SUR-5-GFP, was lost in the Ea endodermal lineage [inset; lost in cells of intestinal ring V (white arrow), and retained in cells of intestinal ring VI (green arrow)]. (b) Schematic of endodermal cell lineage with placements of cells in intestinal rings shown below and reflecting the mosaic pattern seen in a; adapted from60. Green cells indicate cells with rescue array while grey cells represent loss of array. Two mosaics were found with this loss pattern, and both mosaics showed persistent lobes in intestinal ring V (white arrowhead in inset) and lobe debris in intestinal ring VI. 11/11 intestinal mosaic L1 larvae had persistent lobes. (c) Quantification of PGC volume (cell body + lobes) in wild type, ced-10 mutants and ced-10 mutants with ced-10(+)END (WT, n = 14 embryos/L1 larvae; ced-10 mutants, n = 14 embryos/ L1 larvae; ced-10(+)END,n = 14 embryos/L1 larvae. ∗∗∗P < 0.001, Student’s t-test, mean ± s.d.). Data shown is from a single independent experiment. Source data for repeat experiments is provided in Supplementary Table 3. (d) ced-10(tm597) null mutant L1 larva with mosaic rescue ced-10ALL. Rescue is lost in the intestinal ring V, where a PGC lobe persists (arrowhead). 32/32 intestinal mosaic L1 larvae had persistent lobes. Scale bar, 5 μm.
Supplementary Figure 5 lst-4 in lobe scission.
(a) lst-4( +) rescue of persistent lobes in lst-4(xn45)mutants; arrowheads point to lobe debris (4/4 extrachromosomal arrays completely rescued persistent lobes, n = 13–28 L1 larvae examined per array). (b) Quantification of PGC volume (cell body + lobes) in L1 larvae of lst-4(xn45)mutants (n = 10 L1 larvae) and lst-4(xn45) mutants with lst-4( +)(n = 10 L1 larvae), ∗∗∗P < 0.001, Student’s t-test, mean ± s.d. Data shown is from a single independent experiment. Source data for repeat experiments is provided in Supplementary Table 3. (c) Percent recovered lobes in FRAP experiment on persistent lobes in lst-4(xn45) (n = 18 L1 larvae) and lst-4 (RNAi) (n = 18 L1 larvae) L1 larvae. (d) lst-4 gene structure (isoform c, Wormbase WS252). Gray rectangles are coding exons, white rectangle is the 3′ UTR, and chevrons are introns. Regions of the gene encoding the SH3, PX and BAR domains are indicated. The xn45 lesion mutates a splice donor base within an intron in the region encoding the BAR domain. Scale bar, 5 μm.
Supplementary information
Supplementary Information
Supplementary Information (PDF 949 kb)
Supplementary Table 1
Supplementary Information (XLSX 30 kb)
Supplementary Table 2
Supplementary Information (XLSX 34 kb)
Supplementary Table 3
Supplementary Information (XLSX 40 kb)
PGC lobe formation.
Embryo is oriented posterior to the left, and turns from a ventral view to a lateral view (dorsal up) as the movie progresses. PGC and endodermal cell membranes are labeled. PGC lobes (‘L’) begin forming ∼50 min into the movie and embed into endodermal cells. (AVI 2026 kb)
Rendering of PGC lobes embedded into endodermal cells.
Rendered data from a 2-fold embryo expressing endoderm and PGC surface markers. PGCs (magenta) extend lobes into adjacent endodermal cells (green). (AVI 4401 kb)
Lobe formation in a cultured PGC.
PGC membranes and nucleus are labeled. The nucleus moves to one side of the cell and lobe (‘L’) extends from the opposite side beginning at ∼150 min into the movie. (AVI 119 kb)
P granule movement into PGC lobes.
Embryo is oriented posterior to the left. Before lobe formation, all P granules (PGL-1-RFP) are found at the nuclear periphery. A P granule can be seen detaching from the nuclear periphery and moving into the lobe (‘L’) beginning at 42 min into the movie. (AVI 4342 kb)
Mitochondria in PGC lobes.
Embryo is oriented posterior to the left. All cell membranes are labeled with GFP (PGC membranes are brighter), and mitochondria within PGCs are labeled with mCh-MOMA-1 (green and red channels were switched). Both PGCs are initially visible, then one moves out of the focal plane. Lobe formation begins 16 min into the movie, and mitochondria can be seen localizing preferentially to the lobe (‘L’). (AVI 673 kb)
Rights and permissions
About this article
Cite this article
Abdu, Y., Maniscalco, C., Heddleston, J. et al. Developmentally programmed germ cell remodelling by endodermal cell cannibalism. Nat Cell Biol 18, 1302–1310 (2016). https://doi.org/10.1038/ncb3439
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3439
This article is cited by
-
The transplant rejection response involves neutrophil and macrophage adhesion-mediated trogocytosis and is regulated by NFATc3
Cell Death & Disease (2024)
-
Cell-in-cell phenomena across the tree of life
Scientific Reports (2024)
-
Anticancer chemotherapy and radiotherapy trigger both non-cell-autonomous and cell-autonomous death
Cell Death & Disease (2018)
-
Remodelling germ cells by intercellular cannibalism
Nature Cell Biology (2016)