Abstract
Emerging evidence has shown that GSK3β plays oncogenic roles in multiple tumour types; however, the underlying mechanisms remain largely unknown. Here, we show that nuclear GSK3β is responsible for the accumulation of the histone demethylase KDM1A and critically regulates histone H3K4 methylation during tumorigenesis. GSK3β phosphorylates KDM1A Ser683 upon priming phosphorylation of KDM1A Ser687 by CK1α. Phosphorylation of KDM1A induces its binding with and deubiquitylation by USP22, leading to KDM1A stabilization. GSK3β- and USP22-dependent KDM1A stabilization is required for the demethylation of histone H3K4, thereby repressing BMP2, CDKN1A and GATA6 transcription, which results in cancer stem cell self-renewal and glioblastoma tumorigenesis. In human glioblastoma specimens, KDM1A levels are correlated with nuclear GSK3β and USP22 levels. Furthermore, a GSK3 inhibitor, tideglusib, sensitizes tumour xenografts to chemotherapy in mice via KDM1A downregulation and improves survival. Our findings demonstrate that nuclear GSK3β- and USP22-mediated KDM1A stabilization is essential for glioblastoma tumorigenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Cohen, P. & Frame, S. The renaissance of GSK3. Nat. Rev. Mol. Cell Biol. 2, 769–776 (2001).
Doble, B. W. & Woodgett, J. R. GSK-3: tricks of the trade for a multi-tasking kinase. J. Cell Sci. 116, 1175–1186 (2003).
Hoeflich, K. P. et al. Requirement for glycogen synthase kinase-3β in cell survival and NF-κB activation. Nature 406, 86–90 (2000).
Farago, M. et al. Kinase-inactive glycogen synthase kinase 3β promotes Wnt signaling and mammary tumorigenesis. Cancer Res. 65, 5792–5801 (2005).
Wang, Z. et al. GSK-3 promotes conditional association of CREB and its coactivators with MEIS1 to facilitate HOX-mediated transcription and oncogenesis. Cancer Cell 17, 597–608 (2010).
Naito, S. et al. Glycogen synthase kinase-3β: a prognostic marker and a potential therapeutic target in human bladder cancer. Clin. Cancer Res. 16, 5124–5132 (2010).
Aberle, H., Bauer, A., Stappert, J., Kispert, A. & Kemler, R. β-catenin is a target for the ubiquitin-proteasome pathway. EMBO J. 16, 3797–3804 (1997).
Sears, R. et al. Multiple Ras-dependent phosphorylation pathways regulate Myc protein stability. Genes Dev. 14, 2501–2514 (2000).
Tang, Q. L. et al. Glycogen synthase kinase-3β, NF-κB signaling, and tumorigenesis of human osteosarcoma. J. Natl Cancer Inst. 104, 749–763 (2012).
Miyashita, K. et al. Potential therapeutic effect of glycogen synthase kinase 3β inhibition against human glioblastoma. Clin. Cancer Res. 15, 887–897 (2009).
Ougolkov, A. V. et al. Aberrant nuclear accumulation of glycogen synthase kinase-3β in human pancreatic cancer: association with kinase activity and tumor dedifferentiation. Clin. Cancer Res. 12, 5074–5081 (2006).
Ougolkov, A. V., Bone, N. D., Fernandez-Zapico, M. E., Kay, N. E. & Billadeau, D. D. Inhibition of glycogen synthase kinase-3 activity leads to epigenetic silencing of nuclear factor κB target genes and induction of apoptosis in chronic lymphocytic leukemia B cells. Blood 110, 735–742 (2007).
Reddiconto, G. et al. Targeting of GSK3β promotes imatinib-mediated apoptosis in quiescent CD34 + chronic myeloid leukemia progenitors, preserving normal stem cells. Blood 119, 2335–2345 (2012).
Esteller, M. Cancer epigenomics: DNA methylomes and histone-modification maps. Nat. Rev. Genet. 8, 286–298 (2007).
Hill, E. V. et al. Glycogen synthase kinase-3 controls IL-10 expression in CD4(+) effector T-cell subsets through epigenetic modification of the IL-10 promoter. Eur. J. Immunol. 45, 1103–1115 (2015).
Shi, Y. J. et al. Regulation of LSD1 histone demethylase activity by its associated factors. Mol. Cell 19, 857–864 (2005).
Wang, Y. et al. LSD1 is a subunit of the NuRD complex and targets the metastasis programs in breast cancer. Cell 138, 660–672 (2009).
Lv, S. et al. LSD1 is required for chromosome segregation during mitosis. Eur. J. Cell Biol. 89, 557–563 (2010).
Kauffman, E. C. et al. Role of androgen receptor and associated lysine-demethylase coregulators, LSD1 and JMJD2A, in localized and advanced human bladder cancer. Mol. Carcinog. 50, 931–944 (2011).
Hayami, S. et al. Overexpression of LSD1 contributes to human carcinogenesis through chromatin regulation in various cancers. Int. J. Cancer 128, 574–586 (2011).
Harris, W. J. et al. The histone demethylase KDM1A sustains the oncogenic potential of MLL-AF9 leukemia stem cells. Cancer Cell 21, 473–487 (2012).
Sareddy, G. R. et al. KDM1 is a novel therapeutic target for the treatment of gliomas. Oncotarget 4, 18–28 (2013).
Suva, M. L. et al. Reconstructing and reprogramming the tumor-propagating potential of glioblastoma stem-like cells. Cell 157, 580–594 (2014).
Gong, A. H. et al. FoxM1 drives a feed-forward STAT3-activation signaling loop that promotes the self-renewal and tumorigenicity of glioblastoma stem-like cells. Cancer Res. 75, 2337–2348 (2015).
Rena, G., Bain, J., Elliott, M. & Cohen, P. D4476, a cell-permeant inhibitor of CK1, suppresses the site-specific phosphorylation and nuclear exclusion of FOXO1a. EMBO Rep. 5, 60–65 (2004).
Adamo, A. et al. LSD1 regulates the balance between self-renewal and differentiation in human embryonic stem cells. Nat. Cell Biol. 13, 652–659 (2011).
Sun, G. et al. Histone demethylase LSD1 regulates neural stem cell proliferation. Mol. Cell. Biol. 30, 1997–2005 (2010).
Dominguez, J. M. et al. Evidence for irreversible inhibition of glycogen synthase kinase-3β by tideglusib. J. Biol. Chem. 287, 893–904 (2012).
Bolos, M., Fernandez, S. & Torres-Aleman, I. Oral administration of a GSK3 inhibitor increases brain insulin-like growth factor I levels. J. Biol. Chem. 285, 17693–17700 (2010).
Sereno, L. et al. A novel GSK-3β inhibitor reduces Alzheimer’s pathology and rescues neuronal loss in vivo. Neurobiol. Dis. 35, 359–367 (2009).
Chen, J. et al. A restricted cell population propagates glioblastoma growth after chemotherapy. Nature 488, 522–526 (2012).
Liu, C. et al. Control of β-catenin phosphorylation/degradation by a dual-kinase mechanism. Cell 108, 837–847 (2002).
Gregory, M. A., Qi, Y. & Hann, S. R. Phosphorylation by glycogen synthase kinase-3 controls c-myc proteolysis and subnuclear localization. J. Biol. Chem. 278, 51606–51612 (2003).
Shin, S., Wolgamott, L., Yu, Y., Blenis, J. & Yoon, S. O. Glycogen synthase kinase (GSK)-3 promotes p70 ribosomal protein S6 kinase (p70S6K) activity and cell proliferation. Proc. Natl Acad. Sci. USA 108, E1204–E1213 (2011).
Kotliarova, S. et al. Glycogen synthase kinase-3 inhibition induces glioma cell death through c-MYC, nuclear factor-κB, and glucose regulation. Cancer Res. 68, 6643–6651 (2008).
Beurel, E. & Jope, R. S. Differential regulation of STAT family members by glycogen synthase kinase-3. J. Biol. Chem. 283, 21934–21944 (2008).
Park, B. H., Qiang, L. & Farmer, S. R. Phosphorylation of C/EBPβ at a consensus extracellular signal-regulated kinase/glycogen synthase kinase 3 site is required for the induction of adiponectin gene expression during the differentiation of mouse fibroblasts into adipocytes. Mol. Cell. Biol. 24, 8671–8680 (2004).
Foltz, D. R., Santiago, M. C., Berechid, B. E. & Nye, J. S. Glycogen synthase kinase-3β modulates notch signaling and stability. Curr. Biol. 12, 1006–1011 (2002).
Kubic, J. D., Mascarenhas, J. B., Iizuka, T., Wolfgeher, D. & Lang, D. GSK-3 promotes cell survival, growth, and PAX3 levels in human melanoma cells. Mol. Cancer Res. 10, 1065–1076 (2012).
Schaeffer, V. et al. Binding of OTULIN to the PUB domain of HOIP controls NF-κB signaling. Mol. Cell 54, 349–361 (2014).
Elliott, P. R. et al. Molecular basis and regulation of OTULIN-LUBAC interaction. Mol. Cell 54, 335–348 (2014).
Wu, Y. et al. The deubiquitinase USP28 stabilizes LSD1 and confers stem-cell-like traits to breast cancer cells. Cell Rep. 5, 224–236 (2013).
Zhang, X. Y. et al. The putative cancer stem cell marker USP22 is a subunit of the human SAGA complex required for activated transcription and cell-cycle progression. Mol. Cell 29, 102–111 (2008).
Lin, Z. et al. USP22 antagonizes p53 transcriptional activation by deubiquitinating Sirt1 to suppress cell apoptosis and is required for mouse embryonic development. Mol. Cell 46, 484–494 (2012).
Tolosa, E. et al. A phase 2 trial of the GSK-3 inhibitor tideglusib in progressive supranuclear palsy. Mov. Disord. 29, 470–478 (2014).
Persano, L. et al. BMP2 sensitizes glioblastoma stem-like cells to Temozolomide by affecting HIF-1α stability and MGMT expression. Cell Death Differ. 3, e412 (2012).
Kamnasaran, D., Qian, B., Hawkins, C., Stanford, W. L. & Guha, A. GATA6 is an astrocytoma tumor suppressor gene identified by gene trapping of mouse glioma model. Proc. Natl Acad. Sci. USA 104, 8053–8058 (2007).
Ligon, K. L. et al. Olig2-regulated lineage-restricted pathway controls replication competence in neural stem cells and malignant glioma. Neuron 53, 503–517 (2007).
Hirose, Y., Berger, M. S. & Pieper, R. O. p53 effects both the duration of G2/M arrest and the fate of temozolomide-treated human glioblastoma cells. Cancer Res. 61, 1957–1963 (2001).
Schonberg, D. L. et al. Preferential iron trafficking characterizes glioblastoma stem-like cells. Cancer Cell 28, 441–455 (2015).
Hu, Y. & Smyth, G. K. ELDA: extreme limiting dilution analysis for comparing depleted and enriched populations in stem cell and other assays. J. Immunol. Methods 347, 70–78 (2009).
Zhang, N. et al. FoxM1 promotes β-catenin nuclear localization and controls Wnt target-gene expression and glioma tumorigenesis. Cancer Cell 20, 427–442 (2011).
Shembade, N. & Harhaj, E. W. Elucidating dynamic protein-protein interactions and ubiquitination in NF-κB signaling pathways. Methods Mol. Biol. 1280, 283–295 (2015).
Li, M. et al. Deubiquitination of p53 by HAUSP is an important pathway for p53 stabilization. Nature 416, 648–653 (2002).
Acknowledgements
We thank J. Munch in MD Anderson’s Department of Scientific Publications for editing the manuscript. We thank Y. Shi (Boston Children’s Hospital) for providing the Flag–KDM1A plasmid; J. Yang (Texas Children’s Cancer Center) for providing the USP2, USP4, USP5, USP8, USP13, USP14, USP16, USP18, USP21, USP25, USP26, USP28, USP29, USP30, USP36, USP39, USP46 and USP48 expression plasmids; H.-K. Lin (MD Anderson Cancer Center) for providing the USP1, USP3, USP11, USP15, USP22 and CYLD expression plasmids; and X. He (Boston Children’s Hospital) for providing β-catenin+/+ and β-catenin−/− MEFs. This work was supported in part by US National Cancer Institute grants R01CA157933, R01CA182684, R01CA152309, P50CA127001, R01CA195651 and CA16672 (Cancer Center Support Grant).
Author information
Authors and Affiliations
Contributions
S.H. and A.Z. conceived the project and designed the study; A.Z. and K.L. performed most of the experiments under the supervision of S.H.; S.Z. and Y.C. assisted in some in vitro experiments; J.X. and Z.W. assisted in the mouse experiments; N.Z., K.D.A., K.X. and J.R.W. provided reagents and conceptual advice; S.H. and A.Z. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 GSK3β stabilizes KDM1A by inhibiting the proteasome degradation pathway.
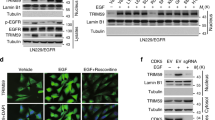
(a) KDM1A protein levels were detected in GSK3β knockout (GSK3β+/+) mouse embryonic fibroblasts (MEFs), GSK3β knockout (GSK3β−/−) MEFs, and GSK3β−/− MEFs with reconstituted expression of GSK3β. (b) Effect of GSK3β knockdown on KDM1A expression. GSC11 cells were transfected with GSK3β siRNAs, and KDM1A expression was detected by immunoblotting. (c) Effect of GSK3α knockdown on KDM1A expression. KDM1A protein levels were detected in GSC11 cells after transfection of GSK3α siRNAs. (d) KDM1A mRNA levels were detected in GSK3β+/+ and GSK3β−/− MEFs. GAPDH was used as an internal control. (e) KDM1A protein levels were detected in β-catenin+/+ and β-catenin−/− MEFs. (f) KDM1A is regulated by the ubiquitin-proteasome pathway. HS683 cells were treated with cycloheximide (CHX) with or without the proteasome inhibitor MG132 for the indicated times and KDM1A levels were detected by immunoblotting. (g) 293T cells were transfected with Flag-KDM1A and the constitutively active form of GSK3β (GSK3β-CA) or the control vector, and then treated with CHX as indicated. Expression of KDM1A was detected by immunoblotting. Fold change of western blot bands was determined as described in Fig. 1b and shown. Source data of Supplementary Fig. 1d can be found in Supplementary Table 3. Uncropped images of blots are shown in Supplementary Fig. 8.
Supplementary Figure 2 GSK3β phosphorylates KDM1A serine 683 after priming phosphorylation by CKIα.
(a) The amino acid sequence of KDM1A shows the sites potentially targeted by GSK3β. (b) Immunofluorescence assays were performed in HFU-251 MG and GSC11 cells using antibodies against KDM1A and GSK3β. Scale bar, 50 μM. (c) Effects of mutations of various Ser/Thr residues on the ubiquitination of KDM1A. 293T cells were transfected with HA-Ubi, Myc-GSK3β, and wild-type Flag-KDM1A or various mutants with serine/threonine-to-alanine mutations. Cells were treated with MG132 for 4 h, harvested, and then lysed for immunoprecipitation (IP) analysis. (d) 293T cells were transfected with wild-type Flag-KDM1A (WT) or mutant with serine 683-to-alanine (S683A) or aspartic acid (S683D) mutation. Cells were treated with CHX for the indicated times. Band intensity of KDM1A from western blots was quantified as described in Fig. 1b (mean ± s.d., n = 3 independent experiments, two-tailed Student’s t-test). ∗P < 0.01. (e) GSC11 or HFU-251 MG cells were treated with the CKI inhibitor D4476 or D4476 plus LiCl for 6 h. Cell lysates were subjected to immunoblotting analysis. Fold change of western blot bands was determined as described in Fig. 1b and shown. (f) Reciprocal interaction between Flag-KDM1A and HA-CK1α. 293T cells were transfected with Flag-KDM1A and HA-CKIα, HA-CKIδ, or HA-CKIɛ plasmids. Cell lysates were immunoprecipitated with the HA-tag (left panel) or Flag-tag antibody (right panel) and then analysed by immunoblotting using the indicated primary antibodies. Blots were incubated with a HRP-conjugated secondary antibody (light chain specific). (g) Reciprocal interaction of endogenous KDM1A with CK1α. GSC11 cell lysates were immunoprecipitated using antibodies against CKIα, CKIδ, and CKIɛ (left panel), or KDM1A (right panel). The immunoprecipitates were subjected to immunoblotting analysis using the indicated primary antibodies and a HRP-conjugated secondary antibody (light chain specific). Inputs correspond to 2% total cell lysates. (h) HF-U251 MG cells were transfected two independent siRNAs against CKIα, CKIδ, or CKIɛ, and then the expression of KDM1A was detected by western blotting. Uncropped images of blots are shown in Supplementary Fig. 8.
Supplementary Figure 3 USP22 deubiquitinates and stabilizes KDM1A.
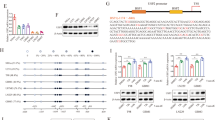
(a) Effect of different deubiquitinating enzymes (DUBs) on the expression of KDM1A. 293T cells were transfected with different DUBs and then lysed for immunoblotting to detect KDM1A expression. (b) Effect of various DUBs on the ubiquitination of KDM1A. 293T cells were transfected with Flag-KDM1A, Myc-Ubi, and USP15, USP21, USP22, or USP28 and then treated with MG132 for 6 h before harvest. Cell lysates were immunoprecipitated using an anti-KDM1A antibody and then subjected to immunoblotting analysis using the indicated antibodies. (c,d) 293T cells were transfected with Flag-KDM1A, Myc-Ubi, and HA-USP22 or Flag-USP28. Cell lysates were immunoprecipitated using antibodies against Myc-tag (c) or KDM1A (d). (e) Knockdown of USP22 does not affect KDM1A mRNA level. GSC11 cell were transfected with control siRNA or USP22 siRNA and KDM1A mRNA were detected by real-time PCR. Values were normalized to that in control (mean ± s.e.m., n = 3 independent experiments, two-tailed Student’s t-test). GAPDH mRNA was used as an internal control. P > 0.05. (f) 293T cells were transfected with HA-USP22 or Flag-USP28 plasmid and then treated with 50 μg ml−1 CHX for the indicated times. Cell lysates were subjected to immunoblotting analysis as indicated. (g) The expression levels of nuclear KDM1A, USP22 and GSK3β in Fig. 3f were determined by quantification of the intensity of western blot bands, using the Lamin B for normalization and the results are expressed as level relative to NHA-E6/E7 cells. Then the correlation of nuclear KDM1A/USP22 and KDM1A/GSK3β levels in different cell lines was analysed (mean ± s.e.m., n = 3 independent experiments, Pearson correlation test). (h) Immunofluorescence assay was used to analyse the co-localization of KDM1A and USP22 in GSC11 cells. Scale bar, 20 μm. Uncropped images of blots are shown in Supplementary Fig. 8.
Supplementary Figure 4 GSK3β promotes the binding of KDM1A with USP22.
293T cells were transfected with Flag-KDM1A, HA-USP22, and GSK3β-CA or GSK3β-KD and then treated with MG132 for 6 h before harvest. Cell lysates were immunoprecipitated with antibodies against (a) HA or (b) KDM1A and then analysed by immunoblotting using the indicated antibodies. Uncropped images of blots are shown in Supplementary Fig. 8.
Supplementary Figure 5 GSK3β and USP22 promote stem cell self-renewal through KDM1A.
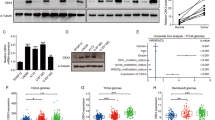
(a) ChIP assays were performed in GSC11 cells transfected with control shRNA, shRNA targeting KDM1A, GSK3β or USP22 using an antibody against KDM1A and primers in the promoter regions of BMP2, CDKNA1, and GATA6. ACTB promoter was used as a negative control. Values are the percentage to input (mean ± s.e.m., n = 3 independent experiments, two-tailed Student’s t-test). ∗P < 0.01. (b) KDM1A expression was detected by immunoblotting in GSC11 cells stably expressing two individual shRNAs targeting KDM1A. (c) Primary (1st) and secondary (2nd) neurosphere formation were assessed in GSC11 and GSC20 cells using two different KDM1A shRNA. Scale bar, 500 μm. (d) Effects of KDM1A knockdown on the expression of stem cell self-renewal and differentiation markers. GSC11 and GSC20 cells expressing KDM1A shRNAs were analysed by immunoblotting using the indicated antibodies. (e) Immunofluorescence assays were performed to detect the expression of self-renewal and differentiation markers in GSC11 cells expressing KDM1A shRNAs. Scale bar, 100 μm. (f) GSC11 cells overexpressing KDM1A shRNAs were reconstituted by the expression of shRNA-resistant KDM1A (ShR), and neurosphere formation was assessed. Scale bar, 500 μm. (g) USP22 knockdown inhibits neurosphere formation through KDM1A. GSC11 cells overexpressing USP22 shRNAs were reconstituted by the expression of wild-type KDM1A, and neurosphere formation was assessed. Scale bar, 500 μm. (h) GSC11 cells stably expressing two different shRNAs targeting GSK3β were analysed by immunoblotting. (i) mRNA levels of KDM1A in GSC11 cells expressing sh-control, sh-GSK3β, sh-GSK3β + KDM1A S683D, and sh-GSK3β + KDM1A S683A were analysed by real-time PCR (mean ± s.e.m., n = 3 independent experiments, two-tailed Student’s t-test). (j) The secondary neurosphere formation efficiency (spheres/cells plated) of GSC11 cells stably expressing the indicated shRNAs or proteins were calculated (mean ± s.e.m., n = 3 independent experiments, two-tailed Student’s t-test). ∗P < 0.05. Uncropped images of blots are shown in Supplementary Fig. 8.
Supplementary Figure 6 Immunohistochemical analysis of GSK3β and KDM1A in mouse brain tumours derived from GSC11 cells expressing Sh-Control, Sh-GSK3β, ShGSK3β + KDM1A S683D, or Sh-GSK3β + KDM1A S683A.
Insets: high magnification images corresponding to the areas marked by yellow dot lines. Scale bar for H&E staining, 50 μm; Scale bar for IHC, 25 μm.
Supplementary Figure 7 Tideglusib inhibits GSC self-renewal and tumour formation.
(a) Effect of tideglusib on Tau phosphorylation. GSC11 cells were treated with the indicated concentrations of tideglusib for 6 h (left panel) or with tideglusib at a final concentration of 2.5 μM for the indicated times (right panel) in the presence of MG132. Cell lysates were analysed by western blotting using antibodies against Tau or phospho-Tau (Ser396). Fold change of western blot bands was determined as described in Fig. 1b and shown. (b) Effect of tideglusib on the expression of stem cell self-renewal and differentiation markers. GSC11 and GSC20 cells were treated with 5 μM tideglusib for 48 h, and Oct4, CD133, and Tuj-1 expression was analysed by immunofluorescence. Scale bar, 50 μm. (c) NHAs were treated with 5 μM tideglusib for the indicated times, and cell viability was analysed using XTT assays. (d) Schedule for animal treatment. For treatment with TMZ (20 mg kg−1 d−1) or tideglusib (25 mg kg−1 d−1) alone, mice were intraperitoneally injected every other day. For combinatorial treatment, mice received injections of TMZ or tideglusib on alternating days for 30 d. Source data of Supplementary Fig. 7c can be found in Supplementary Table 3. Uncropped images of blots are shown in Supplementary Fig. 8.
Supplementary information
Supplementary Information
Supplementary Information (PDF 2194 kb)
Supplementary Table 1
Supplementary Information (XLSX 11 kb)
Supplementary Table 2
Supplementary Information (XLSX 11 kb)
Supplementary Table 3
Supplementary Information (XLSX 12 kb)
Rights and permissions
About this article
Cite this article
Zhou, A., Lin, K., Zhang, S. et al. Nuclear GSK3β promotes tumorigenesis by phosphorylating KDM1A and inducing its deubiquitylation by USP22. Nat Cell Biol 18, 954–966 (2016). https://doi.org/10.1038/ncb3396
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3396
This article is cited by
-
CRISPR/Cas9-based genome-wide screening of the deubiquitinase subfamily identifies USP3 as a protein stabilizer of REST blocking neuronal differentiation and promotes neuroblastoma tumorigenesis
Journal of Experimental & Clinical Cancer Research (2023)
-
GSK3β-driven SOX2 overexpression is a targetable vulnerability in esophageal squamous cell carcinoma
Oncogene (2023)
-
The role of histone H3 lysine demethylases in glioblastoma
Cancer and Metastasis Reviews (2023)
-
PU.1 and MYC transcriptional network defines synergistic drug responses to KIT and LSD1 inhibition in acute myeloid leukemia
Leukemia (2022)
-
GSK3β palmitoylation mediated by ZDHHC4 promotes tumorigenicity of glioblastoma stem cells in temozolomide-resistant glioblastoma through the EZH2–STAT3 axis
Oncogenesis (2022)