Abstract
Microtubules and actin filaments are the two main cytoskeleton networks supporting intracellular architecture and cell polarity. The centrosome nucleates and anchors microtubules and is therefore considered to be the main microtubule-organizing centre. However, recurring, yet unexplained, observations have pointed towards a connection between the centrosome and actin filaments. Here we have used isolated centrosomes to demonstrate that the centrosome can directly promote actin-filament assembly. A cloud of centrosome-associated actin filaments could be identified in living cells as well. Actin-filament nucleation at the centrosome was mediated by the nucleation-promoting factor WASH in combination with the Arp2/3 complex. Pericentriolar material 1 (PCM1) seemed to modulate the centrosomal actin network by regulating Arp2/3 complex and WASH recruitment to the centrosome. Hence, our results reveal an additional facet of the centrosome as an intracellular organizer and provide mechanistic insights into how the centrosome can function as an actin-filament-organizing centre.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Bornens, M. Organelle positioning and cell polarity. Nat. Rev. Mol. Cell Biol. 9, 874–886 (2008).
Etienne-Manneville, S. Actin and microtubules in cell motility: which one is in control? Traffic 5, 470–477 (2004).
Rodriguez, O. C. et al. Conserved microtubule–actin interactions in cell movement and morphogenesis. Nat. Cell Biol. 5, 599–609 (2003).
Coles, C. H. & Bradke, F. Coordinating neuronal actin–microtubule dynamics. Curr. Biol. 25, R677–R691 (2015).
Chesarone, M. A., DuPage, A. G. & Goode, B. L. Unleashing formins to remodel the actin and microtubule cytoskeletons. Nat. Rev. Mol. Cell Biol. 11, 62–74 (2010).
Vaughan, S. & Dawe, H. R. Common themes in centriole and centrosome movements. Trends Cell Biol. 21, 57–66 (2011).
Tang, N. & Marshall, W. F. Centrosome positioning in vertebrate development. J. Cell Sci. 125, 4951–4961 (2012).
Piel, M., Meyer, P., Khodjakov, A., Rieder, C. L. & Bornens, M. The respective contributions of the mother and daughter centrioles to centrosome activity and behavior in vertebrate cells. J. Cell Biol. 149, 317–329 (2000).
Chevrier, V. The Rho-associated protein kinase p160ROCK is required for centrosome positioning. J. Cell Biol. 157, 807–817 (2002).
Euteneuer, U. & Schliwa, M. Evidence for an involvement of actin in the positioning and motility of centrosomes. J. Cell Biol. 101, 96–103 (1985).
Cao, J., Crest, J., Fasulo, B. & Sullivan, W. Cortical actin dynamics facilitate early-stage centrosome separation. Curr. Biol. 20, 770–776 (2010).
Wang, W., Chen, L., Ding, Y., Jin, J. & Liao, K. Centrosome separation driven by actin-microfilaments during mitosis is mediated by centrosome-associated tyrosine-phosphorylated cortactin. J. Cell Sci. 121, 1334–1343 (2008).
Rosenblatt, J., Cramer, L. P., Baum, B. & Mcgee, K. M. Myosin II-dependent cortical movement is required for centrosome separation and positioning during mitotic spindle assembly. Cell 117, 361–372 (2004).
Kunda, P., Pelling, A. E., Liu, T. & Baum, B. Moesin controls cortical rigidity, cell rounding, and spindle morphogenesis during mitosis. Curr. Biol. 18, 91–101 (2008).
Antoniades, I., Stylianou, P. & Skourides, P. A. Making the connection: ciliary adhesion complexes anchor basal bodies to the actin cytoskeleton. Dev. Cell 28, 70–80 (2014).
Pan, J., You, Y., Huang, T. & Brody, S. L. RhoA-mediated apical actin enrichment is required for ciliogenesis and promoted by Foxj1. J. Cell Sci. 120, 1868–1876 (2007).
Kleve, M. G. & Clark, W. H. Association of actin with sperm centrioles: isolation of centriolar complexes and immunofluorescent localization of actin. J. Cell Biol. 86, 87–95 (1980).
Pitaval, A., Tseng, Q., Bornens, M. & Théry, M. Cell shape and contractility regulate ciliogenesis in cell cycle-arrested cells. J. Cell Biol. 191, 303–312 (2010).
Hong, H., Kim, J. & Kim, J. Myosin heavy chain 10 (MYH10) is required for centriole migration during the biogenesis of primary cilia. Biochem. Biophys. Res. Commun. 461, 180–185 (2015).
Dawe, H. R. et al. Nesprin-2 interacts with meckelin and mediates ciliogenesis via remodelling of the actin cytoskeleton. J. Cell Sci. 122, 2716–2726 (2009).
Gomez, T. S. et al. Formins regulate the actin-related protein 2/3 complex-independent polarization of the centrosome to the immunological synapse. Immunity 26, 177–190 (2007).
Chodagam, S., Royou, A., Whitfield, W., Karess, R. & Raff, J. W. The centrosomal protein CP190 regulates myosin function during early Drosophila development. Curr. Biol. 15, 1308–1313 (2005).
Stevenson, V. A., Kramer, J., Kuhn, J. & Theurkauf, W. E. Centrosomes and the Scrambled protein coordinate microtubule-independent actin reorganization. Nat. Cell Biol. 3, 68–75 (2001).
Munro, E., Nance, J. & Priess, J. R. Cortical flows powered by asymmetrical contraction transport PAR proteins to establish and maintain anterior-posterior polarity in the early C. elegans embryo. Dev. Cell 7, 413–424 (2004).
Cowan, C. R. & Hyman, A. a. Centrosomes direct cell polarity independently of microtubule assembly in C. elegans embryos. Nature 431, 92–96 (2004).
Woolner, S., O’Brien, L. L., Wiese, C. & Bement, W. M. Myosin-10 and actin filaments are essential for mitotic spindle function. J. Cell Biol. 182, 77–88 (2008).
Mitsushima, M. et al. Revolving movement of a dynamic cluster of actin filaments during mitosis. J. Cell Biol. 191, 453–462 (2010).
Fink, J. et al. External forces control mitotic spindle positioning. Nat. Cell Biol. 13, 771–778 (2011).
Stinchcombe, J. C., Majorovits, E., Bossi, G., Fuller, S. & Griffiths, G. M. Centrosome polarization delivers secretory granules to the immunological synapse. Nature 443, 462–465 (2006).
Bornens, M. Is the centriole bound to the nuclear membrane? Nature 270, 80–82 (1977).
Burakov, A. V. & Nadezhdina, E. S. Association of nucleus and centrosome: magnet or velcro? Cell Biol. Int. 37, 95–104 (2013).
Bornens, M. & Moudjou, M. Studying the composition and function of centrosomes in vertebrates. Methods Cell Biol. 61, 13–34 (1999).
Andersen, J. S. et al. Proteomic characterization of the human centrosome by protein correlation profiling. Nature 426, 570–574 (2003).
Jakobsen, L. et al. Novel asymmetrically localizing components of human centrosomes identified by complementary proteomics methods. EMBO J. 30, 1520–1535 (2011).
Firat-Karalar, E. N., Sante, J., Elliott, S. & Stearns, T. Proteomic analysis of mammalian sperm cells identifies new components of the centrosome. J. Cell Sci. 127, 4128–4133 (2014).
Bornens, M., Paintrand, M., Berges, J., Marty, M. C. & Karsenti, E. Structural and chemical characterization of isolated centrosomes. Cell Motil. Cytoskeleton 8, 238–249 (1987).
Miranda, A. F., Godman, G. C. & Tanenbaum, S. W. Action of cytochalasin D on cells of established lines. II. Cortex and microfilaments. J. Cell Biol. 62, 406–423 (1974).
Chesarone, M. A. & Goode, B. L. Actin nucleation and elongation factors: mechanisms and interplay. Curr. Opin. Cell Biol. 21, 28–37 (2009).
Hubert, T., Vandekerckhove, J. & Gettemans, J. Actin and Arp2/3 localize at the centrosome of interphase cells. Biochem. Biophys. Res. Commun. 404, 153–158 (2011).
Welch, M. D., DePace, A. H., Verma, S., Iwamatsu, A. & Mitchison, T. J. The human Arp2/3 complex is composed of evolutionarily conserved subunits and is localized to cellular regions of dynamic actin filament assembly. J. Cell Biol. 138, 375–384 (1997).
Nolen, B. J. et al. Characterization of two classes of small molecule inhibitors of Arp2/3 complex. Nature 460, 1031–1034 (2009).
Monfregola, J., Napolitano, G., D’Urso, M., Lappalainen, P. & Ursini, M. V. Functional characterization of Wiskott-Aldrich syndrome protein and scar homolog (WASH), a bi-modular nucleation-promoting factor able to interact with biogenesis of lysosome-related organelle subunit 2 (BLOS2) and gamma-tubulin. J. Biol. Chem. 285, 16951–16957 (2010).
Derivery, E. et al. The Arp2/3 activator WASH controls the fission of endosomes through a large multiprotein complex. Dev. Cell 17, 712–723 (2009).
Bärenz, F., Mayilo, D. & Gruss, O. J. Centriolar satellites: busy orbits around the centrosome. Eur. J. Cell Biol. 90, 983–989 (2011).
Dammermann, A. & Merdes, A. Assembly of centrosomal proteins and microtubule organization depends on PCM-1. J. Cell Biol. 159, 255–266 (2002).
Sillibourne, J. E. et al. Primary ciliogenesis requires the distal appendage component Cep123. Biol. Open 2, 535–545 (2013).
Kubo, A., Sasaki, H., Yuba-Kubo, A., Tsukita, S. & Shiina, N. Centriolar satellites: molecular characterization, ATP-dependent movement toward centrioles and possible involvement in ciliogenesis. J. Cell Biol. 147, 969–980 (1999).
Clark, I. B. & Meyer, D. I. Overexpression of normal and mutant Arp1α (centractin) differentially affects microtubule organization during mitosis and interphase. J. Cell Sci. 112, 3507–3518 (1999).
Patel, H. et al. Kindlin-1 regulates mitotic spindle formation by interacting with integrins and Plk-1. Nat. Commun. 4, 2056 (2013).
Pugacheva, E. N. & Golemis, E. A. The focal adhesion scaffolding protein HEF1 regulates activation of the Aurora-A and Nek2 kinases at the centrosome. Nat. Cell Biol. 7, 937–946 (2005).
Fielding, A. B., Dobreva, I., McDonald, P. C., Foster, L. J. & Dedhar, S. Integrin-linked kinase localizes to the centrosome and regulates mitotic spindle organization. J. Cell Biol. 180, 681–689 (2008).
Hubert, T., Van Impe, K., Vandekerckhove, J. & Gettemans, J. The actin-capping protein CapG localizes to microtubule-dependent organelles during the cell cycle. Biochem. Biophys. Res. Commun. 380, 166–170 (2009).
Bershteyn, M., Atwood, S. X., Woo, W.-M., Li, M. & Oro, A. E. MIM and cortactin antagonism regulates ciliogenesis and hedgehog signaling. Dev. Cell 19, 270–283 (2010).
Epting, D. et al. The Rac1 regulator ELMO controls basal body migration and docking in multiciliated cells through interaction with Ezrin. Development 142, 174–184 (2015).
Sabino, D. et al. Moesin is a major regulator of centrosome behavior in epithelial cells with extra centrosomes. Curr. Biol. 25, 879–889 (2015).
Kaplan, D. D., Meigs, T. E., Kelly, P. & Casey, P. J. Identification of a role for β-catenin in the establishment of a bipolar mitotic spindle. J. Biol. Chem. 279, 10829–10832 (2004).
Ritchey, L., Ottman, R., Roumanos, M. & Chakrabarti, R. A functional cooperativity between Aurora A kinase and LIM kinase1: Implication in the mitotic process. Cell Cycle 11, 296–309 (2012).
Lemullois, M., Klotz, C. & Sandoz, D. Immunocytochemical localization of myosin during ciliogenesis of quail oviduct. Eur. J. Cell Biol. 43, 429–437 (1987).
Espreafico, E. M. et al. Localization of myosin-V in the centrosome. Proc. Natl Acad. Sci. USA 95, 8636–8641 (1998).
Lawo, S., Hasegan, M., Gupta, G. D. & Pelletier, L. Subdiffraction imaging of centrosomes reveals higher-order organizational features of pericentriolar material. Nat. Cell Biol. 14, 1–13 (2012).
Mennella, V. et al. Subdiffraction-resolution fluorescence microscopy reveals a domain of the centrosome critical for pericentriolar material organization. Nat. Cell Biol. 14, 1159–1168 (2012).
Burke, T. A. et al. Homeostatic actin cytoskeleton networks are regulated by assembly factor competition for monomers. Curr. Biol. 24, 579–585 (2014).
Sillibourne, J. E. et al. Autophosphorylation of polo-like kinase 4 and its role in centriole duplication. Mol. Biol. Cell 21, 547–561 (2010).
Dubois, T. et al. Golgi-localized GAP for Cdc42 functions downstream of ARF1 to control Arp2/3 complex and F-actin dynamics. Nat. Cell Biol. 7, 353–364 (2005).
Moudjou, M. & Bornens, M. in Cell Biology: A Laboratory Handbook (ed. Celis, J.) 111–119 (Academic Press, 1998).
Shelanski, M. Chemistry of the filaments and tubules of brain. J. Histochem. Cytochem. 21, 529–539 (1973).
Hyman, A. et al. Preparation of modified tubulins. Methods Enzymol. 196, 478–485 (1991).
Michelot, A. et al. Actin-filament stochastic dynamics mediated by ADF/cofilin. Curr. Biol. 17, 825–833 (2007).
Achard, V. et al. A ‘primer’-based mechanism underlies branched actin filament network formation and motility. Curr. Biol. 20, 423–428 (2010).
Casabona, M. G., Vandenbrouck, Y., Attree, I. & Couté, Y. Proteomic characterization of Pseudomonas aeruginosa PAO1 inner membrane. Proteomics 13, 2419–2423 (2013).
Acknowledgements
This work was supported by the European Research Council (Starting grant 310472), the Agence Nationale pour la Recherche (ANR-12-BSV5-0004-01, ANR-10-INBS-08-01) and the ‘Laboratory of Excellence’ (Grenoble Alliance for Integrated Structural Cell Biology). We are grateful to M. Bornens (Institut Curie, France) for providing pEGFP-centrin1 plasmid, A. Khodjakov (Wadsworth Center, USA) for RPE1 EGFP–centrin-1 cells, R. Tsien (UCSD, USA) for pRSET-B-dTomato plasmid, A. Gautreau (LEBS, France) for WASH antibodies and T. Stradal (Helmholtz Center for Infection Research, Germany) for p34-Arc antibodies. We thank M. Vantard and M. Bornens for constructive discussions.
Author information
Authors and Affiliations
Contributions
J.S., L.B. and M.T. conceived and supervised the project. F.F. and J.S. performed the experimental work. J.G. and C.G. defined the TicTac buffer. Y.C. performed the mass spectrometry analysis. F.F., J.S., L.B. and M.T. analysed the data. M.T. wrote the manuscript, which was edited by F.F., J.S. and L.B.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Actin nucleation and elongation in Brinkley and TicTac buffers.
(a,b) Time-lapse imaging of actin assembly on isolated centrosome (EGFP-centrin1) in Brinkley buffer (a) and TicTac buffer (b). Actin array in the TicTac buffer is denser than in Brinkley buffer. Buffer recipes show in green the common components and in red the different ones. Scale bars represent 10 μm. Images are representative of 2 independent experiments. (c) Pyrene polymerization assay in Brinkley and TicTac buffers performed with 2 μM actin (10% pyrene) in various conditions: in the presence of 2 μM profilin, in the presence of 2 μM actin, 500 nM pWA and 50 nM Arp2/3. Arp2/3 activity is better preserved in TicTac buffer than in Brinkley buffer. Curves are representative of 2 independent experiments. (d) Elongation assay performed with 2 μM actin monomers from phalloidin-stabilized filament seeds in Brinkley (green) and TicTac (purple) buffers. Actin elongation was similar in both conditions (2 independent experiment). (e) The graphic shows the quantification of actin fluorescence intensity integrated over a 2 μm diameter circle around the centrosome, plotted against time and normalized with respect to initial intensity as readout of actin nucleation activity in Brinkley (green, mean of n = 9 actin asters) and TicTac (purple, mean of n = 14 actin asters) buffers. Bars represent the s.d. Actin nucleation activity is reduced in Brinkley buffer.
Supplementary Figure 2 Actin filament polarity on isolated centrosomes.
(a) Growing ends of actin filaments were identified using local photobleaching (green rectangle at t = 5 seconds). The absence of displacement of the photobleached region (see green arrows) indicates that the addition of actin monomers occurred at the aster periphery and not at the center of actin aster. So, filament barbed ends (growing ends) were located at aster periphery. Formins, which are linking filament barbed ends, were probably not located at the centre of the aster. Conversely, Arp2/3 complexes, which are located at filament pointed ends, were more likely to be located at aster center. Scale bar represents 10 μm. (b) Kymograph representing the spatial position over time of the photobleached region (green arrow). Green line indicates the absence of displacement of the photobleached region. (c) Linescans of fluorescence intensity along a radial segment starting from the actin aster center at different time points. Green region indicates the position of the photobleached region. Images are representative of 3 independent experiments.
Supplementary Figure 3 Arp2/3 complex localization on isolated centrosomes.
(a) Immunofluorescent staining of isolated centrosomes (EGFP-centrin1, green) after actin filaments assembly (red). Actin asters were immunostained with three antibodies against distinct subunits of Arp2/3 (Arp2, Arp3 and p34-Arc, green, image are representative of 109, 183 and 104 actin asters for Arp2, Arp3 and p34-Arc respectively from 2 independent experiments). Scale bar represents 10 μm. (b) Western blot of isolated centrosomes immunostained for Arp2. Western blot is representative of 4 independent experiments. (c) Western blot of isolated centrosomes immunostained for WASH1 (1 experiment). (d) Western blot of cell lysates immunostained for WASH1. Cells were treated either with a control siRNA or with a siRNA directed against WASH1 (1 experiment).
Supplementary Figure 4 Specificity of Arp2/3 complex antibodies – in cell competition assay.
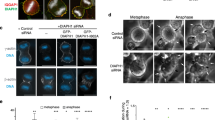
(a,b) Immunofluorescence staining of RPE1 cells fixed in cold methanol with ninein (red) and p34-Arc and Arp2 (green) antibodies, respectively, incubated with increasing quantities of purified Arp2/3 complexes. DNA in blue. Bottom panels show zoom on centrosome region. Reduction of antibody signal confirms the specificity of antigen detection. Scale bars represent 10 μm. (c) Quantification of p34-Arc and Arp2 fluorescence intensity at the centrosome integrated over a 3 μm diameter circle around the centrosome showing the reduction of antibody signal increasing the quantity of purified Arp2/3 complexes. Data are representative of 2 independent experiments: p34-Arc n = 61, 88, 79 and 73 cells and Arp2 n = 42, 29, 37 and 32 cells for 0, 12, 24 and 48 pmol Arp2/3 complex respectively. In red the mean. Unpaired t-test with Welch’s correction (∗∗∗∗ indicates p ≤ 0.0001 for all the conditions).
Supplementary Figure 5 CK666 and SMIFH2 inhibitors – pyrene polymerization assay.
(a) Pyrene polymerization assay to test CK666 inhibitor performed with 2 μM actin (10% pyrene) in various conditions: in the presence of 2 μM actin (DMSO), in the presence of 2 μM actin and 50 nM Arp2/3 (DMSO), in the presence of 2 μM actin, 50 nM Arp2/3 and 500 nM pWA (DMSO), in the presence of 2 μM actin, 50 nM Arp2/3, 500 nM pWA and 60 μM CK666. Arp2/3 activity is reduced in the presence of CK666. Curves are representative of 2 independent experiments. (b) Pyrene polymerization assay to test SMIFH2 inhibitor performed with 2 μM actin (10% pyrene) in various conditions: in the presence of 2 μM actin (DMSO), in the presence of 2 μM actin and 1 nM mDia1 (DMSO), in the presence of 2 μM actin, 1 nM mDia1 and 0.2 mM SMIFH2. SMIFH2 inhibitor reduces mDia1 activity. Curves are representative of 2 independent experiments.
Supplementary Figure 6 CK666 treatment reduces the amount of Arp2/3 complex at the centrosome of HEK293T and RPE1 cells.
(a) and (b) Immunofluorescence staining of DNA (blue), p34-Arc (green) and γ-tubulin (red) of HEK293T and RPE1 cells respectively incubated with DMSO (top panel) or 0.2 mM CK666 (bottom panel) and subsequently fixed with cold methanol. Right panels show the zoom on centrosome region for p34-Arc signal and the merge. Graph represent p34-Arc amount at the centrosome to compare the two conditions. For the HEK293T cells data from 2 independent experiments are pooled: n = 79 and 83 cells for DMSO and CK666 condition respectively. RPE1 cells (1 experiment): n = 21 and 35 cells for DMSO and CK666 condition respectively. In red the mean. Unpaired t-test with Welch’s correction (∗∗∗∗ indicates p ≤ 0.0001 for HEK293T and RPE1 cells). Scale bars represent 10 μm.
Supplementary Figure 7 Involvement of centriolar satellites in Arp2/3 and WASH localization to the centrosome of RPE1 cells.
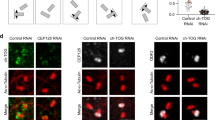
(a–c) Knockdown of PCM1 in RPE1 cells fixed in cold methanol and stained for DNA (blue), ninein (red), p34-Arc (a, green), WASH (b, white) and Arp2 (c, green) respectively. Top row shows cells treated with a negative control siRNA. Bottom row shows cells treated with siRNA against PCM1. Right panels show a zoom on centrosome region. Scale bars; 5 μm. (d) Western blot evaluation of siRNA knockdown for the 2 target sequences, siPCM1-1 and siPCM1-2. GAPDH was used as loading control. (e) Graph show p34-Arc, WASH and Arp2 amount at the centrosome for the control and siPCM1. Data from two distinct siRNA PCM1-targeting sequences are pooled. P34-Arc (1 experiment): n = 41 and 48 cells for control and siPCM1 respectively. For WASH, data from 2 independent experiments are pooled: n = 97 and 100 cells for control and siPCM1 respectively. For Arp2, data from 2 independent experiments are pooled: n = 71 and 172 cells for control and siPCM1 respectively. In red the mean. (f) Immunofluorescence staining of RPE1 cells incubated with DMSO (top panel), nocodazole (central panel) and ciliobrevin D (bottom panel), fixed in cold methanol and stained for DNA (blue), ninein (red), p34-Arc (green) and WASH (white). Right panels show a zoom on centrosome region. Scale bars; 5 μm. Graph show p34-Arc and WASH amount at the centrosome for DMSO, nocodazole and ciliobrevin D conditions. Data from 2 independent experiments are pooled. P34-Arc: n = 54, 83 and 59 cells for DMSO, nocodazole and ciliobrevin D respectively. WASH: n = 129, 192 and 187 cells for DMSO, nocodazole and ciliobrevin D respectively. In red the mean. e,f; unpaired t-test with Welch’s correction was used (∗∗∗∗ indicates p ≤ 0.0001 for all conditions).
Supplementary information
Supplementary Information
Supplementary Information (PDF 1681 kb)
Supplementary Table 1
Supplementary Information (XLSX 101 kb)
Actin assembly in Brinkley and TicTac buffers.
Mix of Actin (1 μM), profilin (3 μM), Arp2/3 (120 nM) and pWA (200 nM) in Brinkley or TicTac buffers were recorded in TIRF microscopy. Note the absence of branches and nucleation of new filaments in the Brinkley buffer. Total elapsed time is 23 minutes and is accelerated 200 times. (MOV 1713 kb)
Dynamic microtubules assembly from centrosomes.
Tubulin dimers were added on top of isolated centrosomes (from Jurkat cells expressing EGFP-centrin1) at 30 μM in TicTac buffer. Microtubule assembly was followed by TIRF microscopy. EGFP-centrin1 is in green and microtubules are in red. Total elapsed time is 22 minutes and is accelerated 300 times. Scale bar is 10 μm. (MOV 376 kb)
Actin filaments assembly from centrosomes.
Actin monomers were added on top of isolated centrosomes (from Jurkat cells expressing EGFP-centrin1) at 1 μM in TicTac buffer. Actin filaments assembly was followed by TIRF microscopy. EGFP-centrin1 is in red and actin filaments are in green. Total elapsed time is 29 minutes and is accelerated 300 times. Scale bar is 10 μm. (MOV 90 kb)
Actin filaments and dynamic microtubules assembly from centrosomes.
Tubulin dimers, 30 μM, and actin monomers, 1 μM, were added on top of isolated centrosomes (from Jurkat cells) in TicTac buffer. Dual-colour time-lapse observation was performed using TIRF microscopy. Microtubules are in red and actin filaments in green. Total elapsed time is 6 minutes and is accelerated 150 times. Scale bar is 10 μm. (MOV 1744 kb)
Actin dynamics in T lymphocytes.
Time-lapse monitoring of actin network and centrosome in Jurkat cells EGFP-centrin1 (green). Cells were transfected with Lifeact-RFP (red) and observed using spinning disk microscopy. Centrosomes were followed over the time using Z stack acquisition. Only the Z plane containing the centrosome is shown. Total elapsed time is 80 minutes and is accelerated 2400 times. Scale bar is 10 μm. (MOV 42 kb)
Actin dynamics in T lymphocytes treated with cytochalasin D.
Time-lapse monitoring of actin network and centrosome in Jurkat cells EGFP-centrin1 (green) in the presence of 10 μg/ml cytochalasin D. Cells were transfected with Lifeact-RFP (red) and observed using spinning disk microscopy. Centrosomes were followed over the time using Z stack acquisition. Only the Z plane containing the centrosome is shown. Total elapsed time is 120 minutes and is accelerated 2400 times. Scale bar is 10 μm. (MOV 55 kb)
Actin incorporation during filament assembly from centrosomes.
Colour switch experiments on isolated centrosomes: Alexa568-actin (red) and Alexa488-actin (green) monomers were added sequentially on isolated centrosomes. Left: merge. Right: Alexa488-actin channel. Total elapsed time is 8 minutes and is accelerated 70 times. Scale bar is 10 μm. (MOV 154 kb)
Actin incorporation during filament assembly from centrosomes in the presence of capping protein.
Colour switch experiments in the presence of capping protein: we introduced sequentially Alexa-568-actin, capping protein and Alexa-488-actin on isolated centrosomes. Left: merge. Right: Alexa-488-actin channel. Total elapsed time is 42 minutes and is accelerated 70 times. Scale bar is 10 μm. (MOV 211 kb)
Inhibition of Arp2/3 during filament assembly from centrosomes.
Colour switch experiments on isolated centrosomes in the absence (top) or presence (bottom) of Arp2/3 complex inhibitor. Alexa568-actin (red) and Alexa488-actin (green) monomers were added sequentially on isolated centrosomes. Left: merge. Right: Alexa488-actin channel. Top: control experiment in the presence of DMSO. Bottom: in the presence of CK666. Total elapsed time is 8 minutes and is accelerated 70 times. Scale bar is 10 μm. (MOV 474 kb)
Inhibition of Arp2/3 during filament assembly from centrosomes in the presence of capping protein.
Colour switch experiments in the presence of capping protein and absence (top) or presence (bottom) of Arp2/3 complex inhibitor. We introduced sequentially Alexa-568-actin, capping protein and Alexa-488-actin on isolated centrosomes. Left: merge. Right: Alexa-488-actin channel. Top: control experiment in the presence of DMSO. Bottom: in the presence of CK666. Total elapsed time is 42 minutes and is accelerated 70 times. Scale bar is 10 μm. (MOV 454 kb)
Rights and permissions
About this article
Cite this article
Farina, F., Gaillard, J., Guérin, C. et al. The centrosome is an actin-organizing centre. Nat Cell Biol 18, 65–75 (2016). https://doi.org/10.1038/ncb3285
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3285
This article is cited by
-
CEP63 upregulates YAP1 to promote colorectal cancer progression through stabilizing RNA binding protein FXR1
Oncogene (2022)
-
A genetically-encoded crosslinker screen identifies SERBP1 as a PKCε substrate influencing translation and cell division
Nature Communications (2021)
-
Spindle positioning and its impact on vertebrate tissue architecture and cell fate
Nature Reviews Molecular Cell Biology (2021)
-
Calponin-3 deficiency augments contractile activity, plasticity, fibrogenic response and Yap/Taz transcriptional activation in lens epithelial cells and explants
Scientific Reports (2020)
-
Acto-myosin force organization modulates centriole separation and PLK4 recruitment to ensure centriole fidelity
Nature Communications (2019)