Abstract
Translation in eukaryotes is followed to detect toxins and virulence factors and coupled to the induction of defence pathways. Caenorhabditis elegans germline-specific mutations in translation components are detected by this system to induce detoxification and immune responses in distinct somatic cells. An RNA interference screen revealed gene inactivations that act at multiple steps in lipid biosynthetic and kinase pathways upstream of MAP kinase to mediate the systemic communication of translation defects to induce detoxification genes. Mammalian bile acids can rescue the defect in detoxification gene induction caused by C. elegans lipid biosynthetic gene inactivations. Extracts prepared from C. elegans with translation deficits but not from the wild type can also rescue detoxification gene induction in lipid-biosynthesis-defective strains. These eukaryotic antibacterial countermeasures are not ignored by bacteria: particular bacterial species suppress normal C. elegans detoxification responses to mutations in translation factors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
11 April 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41556-022-00868-1
References
Xu, C. C., Li, C. Y.-T. C. & Kong, A.-N. T. A. Induction of phase I, II and III drug metabolism/transport by xenobiotics. Arch. Pharm. Res. 28, 249–268 (2005).
Melo, J. A. & Ruvkun, G. Inactivation of conserved C. elegans genes engages pathogen- and xenobiotic-associated defenses. Cell 149, 452–466 (2012).
Dunbar, T. L. T., Yan, Z. Z., Balla, K. M. K., Smelkinson, M. G. M. & Troemel, E. R. E. C. elegans detects pathogen-induced translational inhibition to activate immune signaling. Cell Host Microbe 11, 375–386 (2012).
McEwan, D. L. D., Kirienko, N. V. N. & Ausubel, F. M. F. Host translational inhibition by Pseudomonas aeruginosa Exotoxin A Triggers an immune response in Caenorhabditis elegans. Cell Host Microbe 11, 364–374 (2012).
Waitz, J. A., Sabatelli, F., Menzel, F. & Moss, E. L. Biological activity of antibiotic G-418, a new micromonospora-produced aminoglycoside with activity against protozoa and helminths. Antimicrob. Agents Chemother. 6, 579–581 (1974).
Pittenger, R. C. et al. Hygromycin. I. Preliminary studies on the production and biologic activity of a new antibiotic. Antibiot. Chemother. 3, 1268–1278 (1953).
Maciejowski, J. J. et al. Autosomal genes of autosomal/X-linked duplicated gene pairs and germ-line proliferation in Caenorhabditis elegans. Genetics 169, 1997–2011 (2005).
Hanazawa, M. et al. The Caenorhabditis elegans eukaryotic initiation factor 5A homologue, IFF-1, is required for germ cell proliferation, gametogenesis and localization of the P-granule component PGL-1. Mech. Dev. 121, 213–224 (2004).
Mertenskötter, A., Keshet, A., Gerke, P. & Paul, R. J. The p38 MAPK PMK-1 shows heat-induced nuclear translocation, supports chaperone expression, and affects the heat tolerance of Caenorhabditis elegans. Cell Stress Chaperones 18, 293–306 (2013).
Ferdinandusse, S. et al. Mutations in the gene encoding peroxisomal sterol carrier protein X (SCPx) cause leukencephalopathy with dystonia and motor neuropathy. Am. J. Hum. Genet. 78, 1046–1052 (2006).
Autio, K. J. et al. Role of AMACR (α-methylacyl-CoA racemase) and MFE-1 (peroxisomal multifunctional enzyme-1) in bile acid synthesis in mice. Biochem. J. 461, 125–135 (2014).
Kim, D. H. et al. Integration of Caenorhabditis elegans MAPK pathways mediating immunity and stress resistance by MEK-1 MAPK kinase and VHP-1 MAPK phosphatase. Proc. Natl Acad. Sci. USA 101, 10990–10994 (2004).
Mizuno, T. et al. The Caenorhabditis elegans MAPK phosphatase VHP-1 mediates a novel JNK-like signaling pathway in stress response. EMBO J. 23, 2226–2234 (2004).
Butcher, R. A. et al. Biosynthesis of the Caenorhabditis elegans dauer pheromone. Proc. Natl Acad. Sci. USA 106, 1875–1879 (2009).
Motola, D. L. et al. Identification of ligands for DAF-12 that govern dauer formation and reproduction in C. elegans. Cell 124, 1209–1223 (2006).
Grice, E. A. et al. A diversity profile of the human skin microbiota. Genome Res. 18, 1043–1050 (2008).
Hillion, M. et al. Comparative study of normal and sensitive skin aerobic bacterial populations. Microbiologyopen 2, 953–961 (2013).
Zeeuwen, P. L. et al. Microbiome dynamics of human epidermis following skin barrier disruption. Genome Biol. 13, R101 (2012).
Becker, K. et al. Kocuria rhizophila adds to the emerging spectrum of micrococcal species involved in human infections. J. Clin. Microbiol. 46, 3537–3539 (2008).
Moissenet, D. et al. Persistent bloodstream infection with Kocuria rhizophila related to a damaged central catheter. J. Clin. Microbiol. 50, 1495–1498 (2012).
Azzopardi, E. A. E., Azzopardi, S. M. S., Boyce, D. E. D. & Dickson, W. A. W. Emerging gram-negative infections in burn wounds. J. Burn. Care Res. 32, 570–576 (2011).
Obata, T., Goto, Y., Kunisawa, J. & Sato, S. Indigenous opportunistic bacteria inhabit mammalian gut-associated lymphoid tissues and share a mucosal antibody-mediated symbiosis. Proc. Natl Acad. Sci. USA 107, 7419–7424 (2010).
Sonnenberg, G. F. et al. Innate lymphoid cells promote anatomical containment of lymphoid-resident commensal bacteria. Science 336, 1321–1325 (2012).
Enright, M. R. & Griffin, C. T. Specificity of association between Paenibacillus spp. and the entomopathogenic nematodes, Heterorhabditis spp. Microb. Ecol. 48, 414–423 (2004).
Montalvo-Katz, S., Huang, H. & Appel, M. D. Association with soil bacteria enhances p38-dependent infection resistance in Caenorhabditis elegans. Infect. Immun. 81, 514–520 (2013).
Zhang, R. & Hou, A. Host-microbe interactions in Caenorhabditis elegans. ISRN Microbiol. 2013, 356451 (2013).
Son, S. H., Khan, Z., Kim, S. G. & Kim, Y. H. Plant growth-promoting rhizobacteria, Paenibacillus polymyxa and Paenibacillus lentimorbus suppress disease complex caused by root-knot nematode and fusarium wilt fungus. J. Appl. Microbiol. 107, 524–532 (2009).
Stuart, L. M., Paquette, N. & Boyer, L. Effector-triggered versus pattern-triggered immunity: how animals sense pathogens. Nat. Rev. Immunol. 13, 199–206 (2013).
Durieux, J., Wolff, S. & Dillin, A. The cell-non-autonomous nature of electron transport chain-mediated longevity. Cell 144, 79–91 (2011).
Liu, Y., Samuel, B. S., Breen, P. C. & Ruvkun, G. Caenorhabditis elegans pathways that surveil and defend mitochondria. Nature 508, 406–410 (2014).
Prahlad, V., Cornelius, T. & Morimoto, R. I. Regulation of the cellular heat shock response in Caenorhabditis elegans by thermosensory neurons. Science 320, 811–814 (2008).
Saridaki, A. & Bourtzis, K. Wolbachia: more than just a bug in insects genitals. Curr. Opin. Microbiol. 13, 67–72 (2010).
Fenn, K. & Blaxter, M. Wolbachia genomes: revealing the biology of parasitism and mutualism. Trends Parasitol. 22, 60–65 (2006).
Widmann, B. et al. The kinase activity of human Rio1 is required for final steps of cytoplasmic maturation of 40S subunits. Mol. Biol. Cell 23, 22–35 (2012).
Thomas, C., Pellicciari, R., Pruzanski, M., Auwerx, J. & Schoonjans, K. Targeting bile-acid signalling for metabolic diseases. Nat. Rev. Drug Discov. 7, 678–693 (2008).
Ridlon, J. M., Kang, D.-J. & Hylemon, P. B. Bile salt biotransformations by human intestinal bacteria. J. Lipid Res. 47, 241–259 (2006).
Kawamata, Y. et al. A G protein-coupled receptor responsive to bile acids. J. Biol. Chem. 278, 9435–9440 (2003).
Acknowledgements
Strains were provided by the Caenorhabditis Genetics Center (CGC), which is funded by the NIH Office of Research Infrastructure Programs (P40 OD010440), and S. Mitani of the Japanese National Bioresources Project. This work was supported in part by the National Institutes of Health Grant AG043184-16A (to G.R.). Thanks to D. Coil and J. Eisen for providing Kocuria species.
Author information
Authors and Affiliations
Contributions
J.A.G. and G.R. conceived the project. E.J. and E.M. provided the microbes used in the experiments. E.J. carried out the bacterial suppression experiments. J.L.-F. helped with the data collection and analysis. X.Z. conducted the qRT-PCR experiments. P.B. constructed transgenic pmk-1::mcherry lines. J.A.G. and G.R. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 8 Germline translation mutations induce systemic xenobiotic detoxification response.
(a) pgp-5::gfp is induced in the intestine of animals homozygous for the iff-1(tm483) germline translation defective mutation but weakly in heterozygous iff-1(tm483)/+ animals. Arrow indicates the red pharyngeal marker of the animals heterozygous for iff-1(tm483). Small arrowhead points out iff-1(tm483)homozygous animals. Scale bar, 50 μm. (b) cyp-34A9::gfp is induced in the homozygous iff-1(tm483) mutant but not in wild type. Scale bar, 50 μm. (c) pgp-5::gfp is induced in the homozygous eft-3(q145) mutant but not in wild type. Scale bar, 50 μm. (d) RNAi of rpl-11.2 or eft-4 induce pgp-5::gfp in the intestine. Scale bar, 50 μm. (e) hsp-60::gfp is not induced in eft-3(q145) mutants. Mean ± s.d. of n = 10 worms for each condition. Data represent one out of two independent experiments. (f) Quantification of pgp-5::gfp induction in the eft-3(q145);pgp-5::gfp germline translation defective strain after ablation of germline precursors. (g) Ablation of Z2 and Z3 germline precursors disrupts pgp-5::gfp expression in the intestine in eft-3(q145);pgp-5::gfp. Scale bar, 50 μm. (h) Quantification of the GFP fluorescence after ablation of the germline in eft-3(q145);pgp-5::gfp mutants using imageJ software. Error bars represent SD. Unpaired t test; ∗∗∗P < 0.001. ns denotes no significant difference. (i) Translation-inhibition by eft-4 RNAi only in the neurons or muscle or hypodermis induces pgp-5::gfp gene expression in the intestine. Scale bar, 50 μm. (j) Translation inhibition by eft-4 RNAi only in neurons or hypodermis activates transcription of pgp-5 mRNA as assessed by qPCR. Error bars represent SD. Unpaired t test; ∗∗∗P < 0.001. ns denotes no significant difference. ∼300 worms per condition were washed off 1 plate for each experiment. Mean ± s.d. is from n = 3 independent experiments.
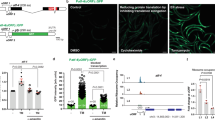
Supplementary Figure 9 Genetic pathways that mediate translation-defect-induced xenobiotic defense response.
(a) Specificity of the anti-p38 MAPK antibody in eft-3(q145);pmk-1(km25) mutant. No nuclear staining is present. Non-specific cytoplasmic staining is present at low levels. Scale bar, 20 μm. (b) PMK-1–p38 MAPK protein is nuclearly localized throughout the germline in wild type as well as the eft-3(q145) germline translation-defective mutant, suggesting that other pathways induce this kinase cascade in the normal germline. Anti-p38 MAPK antibody staining on dissected germline from wildtype showing active p38 in the rachis region. Non-specific cytoplasmic staining is present at low levels. Scale bars, 20 μm. (c) Anti-p38 MAPK antibody staining on dissected germline from wildtype showing active p38 in distal region. Non-specific cytoplasmic staining is present at low levels. Scale bar, 20 μm. (d) Nuclear localization of mCherry::PMK-1 in germline translation defective mutant. Scale bar, 50 μm. (e) Nuclearly localized p38 in the intestine of germline translation defective mutants carrying mCherry::PMK-1 corresponds to the active phosphorylated p38 MAPK. (f) Overlapping sets of genes identified in the distinct screens reported here. The RNAi hits from the screen are shown as circles. The numbers show the overlapping and unique genes in each screen. (g) Mutations in genes required for lipid–bile acid biosynthesis cause hypersensitivity to hygromycin. Scale bar: 1 mm. (h) dhs-28 is required for p38 MAPK phosphorylation and nuclear translocation of active p38 in the intestine of the eft-3(q145) strain. Arrows indicate the nuclear p38 staining. (i) An overview of bile acid biosynthetic pathways.
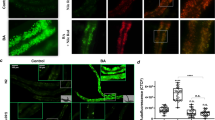
Supplementary Figure 10 Exogenous mammalian bile acids enhance the induction of pgp-5::gfp in response to mild translation inhibition. Mammalian bile acids do not induce pgp-5::gfp expression in the absence of ribosomal defects. Scale bar, 50 μm.
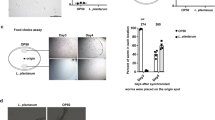
(a) Quantification of gfp fluorescence in animals treated with mammalian bile acids. Error bars represent SD. Unpaired t-test. n = 10 worms for each condition. ns no significant difference. Data represent one out of two independent experiments. (b) Exogenous mammalian bile acids enhance the induction of pgp-5::gfp expression in response to mild translational defect in dilute rpl-7 RNAi-treated animals. Fold change was calculated in comparison to pgp-5::gfp carrying animals. Error bars represent SD. One-way ANOVA; ∗∗∗P < 0.001. ns, P > 0.05. n = 10 worms for each condition. Data represent one out of two independent experiments. (c) Mammalian bile acids are sufficient to induce pgp-5::gfp expression in the vhp-1(sa366);pgp-5::gfp strain without ribosomal defects. Scale bar, 50 μm. (d) Mammalian bile acids do not induce hsp-4::gfp (a ER stress response chaperone gene) in response to low doses of DTT. Scale bar, 50 μm. (e) Mixed bile acid treatment or Kocuria rhizophila feeding does not induce gpdh-1::gfp expression. Error bars represent SD. One-way ANOVA. ns, P > 0.05. n = 10 worms for each condition. Data represent one out of two independent experiments. (f) Mammalian bile acids enhance the response to mild translational defect in dilute eft-4 RNAi-treated animals to induce pgp-5::gfp expression. Scale bar, 50 μm. (g) Lipid extract from eft-4RNAi, iff-2 RNAi or rpl-7 RNAi enhance the mild translational defect in dilute eft-4 RNAi-treated animals to induce pgp-5::gfp expression. Fold change was calculated in comparison to pgp-5::gfp carrying animals. Error bars represent SD. One-way ANOVA; ∗∗∗P < 0.001. ns, P > 0.05. n = 10 worms for each condition. Data represent one out of two independent experiments. (h) pgp-5::gfp expression in wild type animals is not induced by co-culturing with a large excess of eft-3(q145) mutant animals each experiencing germline-translation defects and thus sterility, but not carrying pgp-5::gfp. Thus there is no pheromone for translational stress or bile acid signaling between animals. Scale bar, 50 μm. (i) daf-12 nuclear hormone gene activity is not necessary for pgp-5::gfp induction by hygromycin and addition of the ligand for DAF-12, dafachronic acid, is not sufficient to induce pgp-5::gfp absence of hygromycin. n = 20 worms for each condition. Data represent one out of two independent experiments. (j) Quantification of the GFP fluorescence in worms growing on lawns of Paenibacilli, Kocuria or Alcaligenes bacteria that disrupt induction of pgp-5::gfp in response to germline translation defects in the eft-3(q145);pgp-5::gfp strain, compared to growth on control E. coli OP50. Error bars represent SD. One-way ANOVA; ∗∗∗P < 0.001. n = 10 worms for each condition. Data represent one out of two independent experiments. (k) Animals growing on lawns of Paenibacilli, Kocuria or Alcaligenes bacteria show normal induction of hsp-4::gfp in response to ER stress. Scale bar, 50 μm. (l) Kocuria anti-translation surveillance activity is live cell-associated, inactivated by heat and requires continued exposure to Kocuria. n = 30 worms for each condition. Data represent one out of three independent experiments.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1515 kb)
Supplementary Table 8
Supplementary Information (XLSX 83 kb)
Supplementary Table 9
Supplementary Information (XLSX 44 kb)
Rights and permissions
About this article
Cite this article
Govindan, J., Jayamani, E., Zhang, X. et al. Lipid signalling couples translational surveillance to systemic detoxification in Caenorhabditis elegans. Nat Cell Biol 17, 1294–1303 (2015). https://doi.org/10.1038/ncb3229
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3229