Abstract
The differentiation of stem cells is a tightly regulated process essential for animal development and tissue homeostasis. Through this process, attainment of new identity and function is achieved by marked changes in cellular properties. Intrinsic cellular mechanisms governing stem cell differentiation remain largely unknown, in part because systematic forward genetic approaches to the problem have not been widely used1,2. Analysing genes required for germline stem cell differentiation in the Drosophila ovary, we find that the mitochondrial ATP synthase plays a critical role in this process. Unexpectedly, the ATP synthesizing function of this complex was not necessary for differentiation, as knockdown of other members of the oxidative phosphorylation system did not disrupt the process. Instead, the ATP synthase acted to promote the maturation of mitochondrial cristae during differentiation through dimerization and specific upregulation of the ATP synthase complex. Taken together, our results suggest that ATP synthase-dependent crista maturation is a key developmental process required for differentiation independent of oxidative phosphorylation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Neumuller, R. A. et al. Genome-wide analysis of self-renewal in Drosophila neural stem cells by transgenic RNAi. Cell Stem Cell 8, 580–593 (2011).
Yan, D. et al. A regulatory network of Drosophila germline stem cell self-renewal. Dev. Cell 28, 459–473 (2014).
Gilboa, L. & Lehmann, R. How different is Venus from Mars? The genetics of germ-line stem cells in Drosophila females and males. Development 131, 4895–4905 (2004).
Spradling, A., Fuller, M. T., Braun, R. E. & Yoshida, S. Germline stem cells. Cold Spring Harb. Perspect. Biol. 3, a002642 (2011).
Czech, B., Preall, J. B., McGinn, J. & Hannon, G. J. A transcriptome-wide RNAi screen in the Drosophila ovary reveals factors of the germline piRNA pathway. Mol. Cell 50, 749–761 (2013).
Dietzl, G. et al. A genome-wide transgenic RNAi library for conditional gene inactivation in Drosophila. Nature 448, 151–156 (2007).
Van Doren, M., Williamson, A. L. & Lehmann, R. Regulation of zygotic gene expression in Drosophila primordial germ cells. Curr. Biol. 8, 243–246 (1998).
Vinayagam, A. et al. Protein complex-based analysis framework for high-throughput data sets. Sci. Signal. 6, rs5 (2013).
Walker, J. E. The ATP synthase: the understood, the uncertain and the unknown. Biochem. Soc. Trans. 41, 1–16 (2013).
Wittig, I. et al. Assembly and oligomerization of human ATP synthase lacking mitochondrial subunits a and A6L. Biochim. Biophys. Acta 1797, 1004–1011 (2010).
Gilboa, L., Forbes, A., Tazuke, S. I., Fuller, M. T. & Lehmann, R. Germ line stem cell differentiation in Drosophila requires gap junctions and proceeds via an intermediate state. Development 130, 6625–6634 (2003).
Kai, T. & Spradling, A. An empty Drosophila stem cell niche reactivates the proliferation of ectopic cells. Proc. Natl Acad. Sci. USA 100, 4633–4638 (2003).
Lin, H. & Spradling, A. C. Fusome asymmetry and oocyte determination in Drosophila. Dev. Genet. 16, 6–12 (1995).
McKearin, D. M. & Spradling, A. C. bag-of-marbles: a Drosophila gene required to initiate both male and female gametogenesis. Genes Dev. 4, 2242–2251 (1990).
Lantz, V., Chang, J. S., Horabin, J. I., Bopp, D. & Schedl, P. The Drosophila orb RNA-binding protein is required for the formation of the egg chamber and establishment of polarity. Genes Dev. 8, 598–613 (1994).
Mitchell, P. Chemiosmotic coupling in oxidative and photosynthetic phosphorylation. 1966. Biochim. Biophys. Acta 1807, 1507–1538 (2011).
Neumuller, R. A. et al. Mei-P26 regulates microRNAs and cell growth in the Drosophila ovarian stem cell lineage. Nature 454, 241–245 (2008).
Davies, K. M. et al. Macromolecular organization of ATP synthase and complex I in whole mitochondria. Proc. Natl Acad. Sci. USA 108, 14121–14126 (2011).
Paumard, P. et al. The ATP synthase is involved in generating mitochondrial cristae morphology. EMBO J. 21, 221–230 (2002).
Wittig, I., Karas, M. & Schagger, H. High resolution clear native electrophoresis for in-gel functional assays and fluorescence studies of membrane protein complexes. Mol. Cell. Proteomics 6, 1215–1225 (2007).
Lonergan, T., Bavister, B. & Brenner, C. Mitochondria in stem cells. Mitochondrion 7, 289–296 (2007).
Sathananthan, H., Pera, M. & Trounson, A. The fine structure of human embryonic stem cells. Reprod. Biomed. Online 4, 56–61 (2002).
Kasahara, A., Cipolat, S., Chen, Y., Dorn, G. W. & 2nd Scorrano, L. Mitochondrial fusion directs cardiomyocyte differentiation via calcineurin and Notch signaling. Science 342, 734–737 (2013).
Mitra, K., Rikhy, R., Lilly, M. & Lippincott-Schwartz, J. DRP1-dependent mitochondrial fission initiates follicle cell differentiation during Drosophila oogenesis. J. Cell Biol. 197, 487–497 (2012).
Rieger, B., Junge, W. & Busch, K. B. Lateral pH gradient between OXPHOS complex IV and F(0)F(1) ATP-synthase in folded mitochondrial membranes. Nat. Commun. 5, 3103 (2014).
Strauss, M., Hofhaus, G., Schroder, R. R. & Kuhlbrandt, W. Dimer ribbons of ATP synthase shape the inner mitochondrial membrane. EMBO J. 27, 1154–1160 (2008).
Giorgio, V. et al. Dimers of mitochondrial ATP synthase form the permeability transition pore. Proc. Natl Acad. Sci. USA 110, 5887–5892 (2013).
Fukuoh, A. et al. Screen for mitochondrial DNA copy number maintenance genes reveals essential role for ATP synthase. Mol. Syst. Biol. 10, 734–755 (2014).
Brand, A. H. & Perrimon, N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118, 401–415 (1993).
Barrett, K., Leptin, M. & Settleman, J. The Rho GTPase and a putative RhoGEF mediate a signaling pathway for the cell shape changes in Drosophila gastrulation. Cell 91, 905–915 (1997).
Chen, D. & McKearin, D. M. A discrete transcriptional silencer in the bam gene determines asymmetric division of the Drosophila germline stem cell. Development 130, 1159–1170 (2003).
Lee, T. & Luo, L. Mosaic analysis with a repressible cell marker for studies of gene function in neuronal morphogenesis. Neuron 22, 451–461 (1999).
Ohlstein, B. & McKearin, D. Ectopic expression of the Drosophila Bam protein eliminates oogenic germline stem cells. Development 124, 3651–3662 (1997).
LaJeunesse, D. R. et al. Three new Drosophila markers of intracellular membranes. BioTechniques 36, 784–788 (2004).
Zaccai, M. & Lipshitz, H. D. Differential distributions of two adducin-like protein isoforms in the Drosophila ovary and early embryo. Zygote 4, 159–166 (1996).
Liu, N., Han, H. & Lasko, P. Vasa promotes Drosophila germline stem cell differentiation by activating mei-P26 translation by directly interacting with a (U)-rich motif in its 3′ UTR. Genes Dev. 23, 2742–2752 (2009).
Ramadan, N., Flockhart, I., Booker, M., Perrimon, N. & Mathey-Prevot, B. Design and implementation of high-throughput RNAi screens in cultured Drosophila cells. Nat. Protoc. 2, 2245–2264 (2007).
Celniker, S. E. et al. Unlocking the secrets of the genome. Nature 459, 927–930 (2009).
Suhai, T., Heidrich, N. G., Dencher, N. A. & Seelert, H. Highly sensitive dection of ATPase activity in native gels. Electrophoresis 20, 3622–3625 (2009).
Luft, J. H. Improvements in epoxy resin embedding methods. J. Biophys. Biochem. Cytol. 9, 409–414 (1961).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Acknowledgements
We thank C. Malone, M. Murphy, J. Carroll, A. Sfeir, E. Skolnik, R. Cinalli, A. Zamparini and A. Blum for comments on the manuscript and advice. We thank F. Liang, C. Petzold and K. Dancel of the NYULMC OCS Microscopy Core for their assistance with transmission electron microscopy, and the NYULMC Immune Monitoring Core supported in part by NCATS NIH grant UL1 TR00038 and NCI NIH grant P30CA016087. We acknowledge the DGRC supported by NIH 2P40OD010949-10A1 and Bloomington Stock Center for reagents. F.K.T. was supported by EMBO and HFSP long-term fellowships, C.G.S. by NIH F31/HD080380, T.R.H. by CIHR, J.R.K.S. by NIH F32/GM082169, B.C. by a PhD fellowship from the Boehringer Ingelheim Fonds and J.B.P. by ACS award 121614-PF-11-277-01-RMC. This work was supported by the NIH (5R01GM062534) and a kind gift from K. W. Davis to G.J.H. G.J.H. is an investigator of the HHMI. R.L. is an HHMI investigator and is supported by NIH R01/R37HD41900.
Author information
Authors and Affiliations
Contributions
F.K.T., T.R.H., R.L., C.G.S. and J.R.K.S. designed the experiments. C.G.S. carried out most of the Drosophila crosses, dissections and immunofluorescence stainings. J.R.K.S. and F.K.T. acquired the majority of the confocal microscopy images, with T.R.H. and C.G.S. also contributing. F.K.T. made the dsRNA, carried out the bioinformatics analysis and carried out the RNA expression analysis. T.R.H. carried out the cell culture transfections and CN-PAGE analysis. T.R.H., F.K.T. and R.L. wrote the manuscript, with all authors approving the final version. B.C., J.B.P. and G.J.H. provided the initial list of genes required for oogenesis. R.L., T.R.H., F.K.T., C.G.S. and J.R.K.S. contributed to the discussion of the results.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
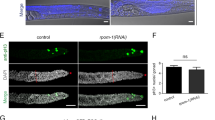
Supplementary Figure 1 Mitochondrial transcription, translation and protein import machinery components are required for germ cell differentiation.
The following VDRC UAS-RNAi strains were expressed in the germline: 105995/KK for mRpL18, 104412/KK for mRpL21, 101506/KK for mRpL27, 101442/KK for mRpL40, 101656/KK for mTTF, and 13794/GD for Blp. Knockdown germaria were immunostained with α-Vasa (green) and α-1B1 (red). Images are representative of at least 100 ovarioles analyzed per genotype. Scale bar, 20 μm.
Supplementary Figure 2 Transfection of S2R+ cells with dsRNA effectively silences the expression of target genes as measured by RT-qPCR.
The data are the means of three technical replicates.
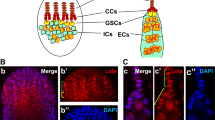
Supplementary Figure 3 ATP synthase subunit β is up-regulated in differentiating cysts.
Germaria expressing a mitochondrially-targeted EYFP (mitoEYFP) were immunostained with α-mei-P26 (right),  GFP, and
GFP, and  ATP synthase β. The ratio of ATP synthase β to mitoEYFP is shown. Image is representative of at least 100 ovarioles analyzed per genotype. Scale bar, 20 μm.
ATP synthase β. The ratio of ATP synthase β to mitoEYFP is shown. Image is representative of at least 100 ovarioles analyzed per genotype. Scale bar, 20 μm.
Supplementary Figure 4 ATP synthase subunit g is not required for ATP synthase stability in S2R+ cells.
Immunofluorescence staining of S2R+ cells transfected with dsRNA that silences complex III, IV and ATP synthase subunits. Cells were immunostained with α-ATP synthase α (green top), α-ATP synthase β (green middle), and α-PDH E1α (green bottom). Images are representative of approximately 300 cells assessed from two fields. Scale bar, 100 μm.
Supplementary Figure 5 Uncropped images of electrophoretic separation techniques in Fig. 5.
(a) CN-PAGE of S2R+ cells treated with dsRNA targeting lacZ or ATP synthase subunits g, e, α or β. ATP synthase was detected by immunoblotting with α-ATP synthase β. Image is representative of 3 independent experiments. (b) SDS-PAGE of the same samples followed by immunoblotting with α-porin served as a sample processing control. (c) CN-PAGE of S2R+ cells treated with dsRNA targeting lacZ or ATP synthase subunits g, e, α or β. ATPase activity was measured in gel. Image is representative of 2 independent experiments.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1687 kb)
Supplementary Table 1
Supplementary Information (XLSX 13 kb)
Supplementary Table 2
Supplementary Information (XLSX 31 kb)
Supplementary Table 3
Supplementary Information (XLSX 46 kb)
Supplementary Table 4
Supplementary Information (XLSX 45 kb)
Supplementary Table 5
Supplementary Information (XLSX 41 kb)
Rights and permissions
About this article
Cite this article
Teixeira, F., Sanchez, C., Hurd, T. et al. ATP synthase promotes germ cell differentiation independent of oxidative phosphorylation. Nat Cell Biol 17, 689–696 (2015). https://doi.org/10.1038/ncb3165
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3165
This article is cited by
-
Intestine-enriched apolipoprotein b orthologs are required for stem cell progeny differentiation and regeneration in planarians
Nature Communications (2022)
-
Autophagy is required for spermatogonial differentiation in the Drosophila testis
Biologia Futura (2022)
-
Formyl tetrahydrofolate deformylase affects hydrogen peroxide accumulation and leaf senescence by regulating the folate status and redox homeostasis in rice
Science China Life Sciences (2021)
-
Cellular metabolic reprogramming controls sugar appetite in Drosophila
Nature Metabolism (2020)
-
Protein signatures of seminal plasma from bulls with contrasting frozen-thawed sperm viability
Scientific Reports (2020)