Abstract
Epithelial organs develop through tightly coordinated events of cell proliferation and differentiation in which endocytosis plays a major role. Despite recent advances, how endocytosis regulates the development of vertebrate organs is still unknown. Here we describe a mechanism that facilitates the apical availability of endosomal SNARE receptors for epithelial morphogenesis through the developmental upregulation of plasmolipin (pllp) in a highly endocytic segment of the zebrafish posterior midgut. The protein PLLP (Pllp in fish) recruits the clathrin adaptor EpsinR to sort the SNARE machinery of the endolysosomal pathway into the subapical compartment, which is a switch for polarized endocytosis. Furthermore, PLLP expression induces apical Crumbs internalization and the activation of the Notch signalling pathway, both crucial steps in the acquisition of cell polarity and differentiation of epithelial cells. We thus postulate that differential apical endosomal SNARE sorting is a mechanism that regulates epithelial patterning.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Rodriguez-Boulan, E. & Macara, I. G. Organization and execution of the epithelial polarity programme. Nat. Rev. Mol. Cell Biol. 15, 225–242 (2014)
Eaton, S. & Martin-Belmonte, F. Cargo sorting in the endocytic pathway: a key regulator of cell polarity and tissue dynamics. Cold Spring Harb. Perspect. Biol. 6 (2014).
Fabrowski, P. et al. Tubular endocytosis drives remodelling of the apical surface during epithelial morphogenesis in Drosophila. Nat. Commun. 4, 2244 (2013)
Richardson, E. C. & Pichaud, F. Crumbs is required to achieve proper organ size control during Drosophila head development. Development 137, 641–650 (2010).
Lu, H. & Bilder, D. Endocytic control of epithelial polarity and proliferation in Drosophila. Nat. Cell Biol. 7, 1232–1239 (2005)
Bokel, C. & Brand, M. Endocytosis and signaling during development. Cold Spring Harb. Perspect. Biol. 6 (2014).
Bagnat, M., Cheung, I. D., Mostov, K. E. & Stainier, D. Y. Genetic control of single lumen formation in the zebrafish gut. Nat. Cell Biol. 9, 954–960 (2007)
Alvers, A. L., Ryan, S., Scherz, P. J., Huisken, J. & Bagnat, M. Single continuous lumen formation in the zebrafish gut is mediated by smoothened-dependent tissue remodeling. Development 141, 1110–1119 (2014).
Sanchez-Pulido, L., Martin-Belmonte, F., Valencia, A. & Alonso, M. A. MARVEL: a conserved domain involved in membrane apposition events. Trends Biochem. Sci. 27, 599–601 (2002).
Navis, A., Marjoram, L. & Bagnat, M. Cftr controls lumen expansion and function of Kupffer’s vesicle in zebrafish. Development 140, 1703–1712 (2013).
Cocchiaro, J. L. & Rawls, J. F. Microgavage of zebrafish larvae. J. Visualized Experiments: JoVE e4434 (2013)
Cermak, T. et al. Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res. 39, e82 (2011)
Golachowska, M. R., Hoekstra, D. & van, I. S. C. Recycling endosomes in apical plasma membrane domain formation and epithelial cell polarity. Trends Cell Biol. 20, 618–626 (2010).
Galvez-Santisteban, M. et al. Synaptotagmin-like proteins control the formation of a single apical membrane domain in epithelial cells. Nat. Cell Biol. 14, 838–849 (2012).
Saint-Pol, A. et al. Clathrin adaptor epsinR is required for retrograde sorting on early endosomal membranes. Dev. Cell 6, 525–538 (2004).
Mills, I. G. et al. EpsinR: an AP1/clathrin interacting protein involved in vesicle trafficking. J. Cell Biol. 160, 213–222 (2003).
Hirst, J., Motley, A., Harasaki, K., Peak Chew, S. Y. & Robinson, M. S. EpsinR: an ENTH domain-containing protein that interacts with AP-1. Mol. Biol. Cell 14, 625–641 (2003).
Miller, S. E., Collins, B. M., McCoy, A. J., Robinson, M. S. & Owen, D. J. A SNARE-adaptor interaction is a new mode of cargo recognition in clathrin-coated vesicles. Nature 450, 570–574 (2007).
Chidambaram, S., Zimmermann, J. & von Mollard, G. F. ENTH domain proteins are cargo adaptors for multiple SNARE proteins at the TGN endosome. J. Cell Sci. 121, 329–338 (2008).
Chidambaram, S., Mullers, N., Wiederhold, K., Haucke, V. & von Mollard, G. F. Specific interaction between SNAREs and epsin N-terminal homology (ENTH) domains of epsin-related proteins in trans-Golgi network to endosome transport. J. Biol. Chem. 279, 4175–4179 (2004).
Leventis, P. A. et al. Liquid facets-related (lqfR) is required for egg chamber morphogenesis during Drosophila oogenesis. PloS ONE 6, e25466 (2011)
Lee, J. H., Overstreet, E., Fitch, E., Fleenor, S. & Fischer, J. A. Drosophila liquid facets-Related encodes Golgi epsin and is an essential gene required for cell proliferation, growth and patterning. Dev. Biol. 331, 1–13 (2009).
Leuchowius, K. J., Weibrecht, I. & Soderberg, O. In situ proximity ligation assay for microscopy and flow cytometry. in Current Protocols in Cytometry (eds Robinson, J. P. et al.) Ch. 9 (Wiley, 2011)
Thompson, B. J., Pichaud, F. & Roper, K. Sticking together the Crumbs—an unexpected function for an old friend. Nat. Rev. Mol. Cell Biol. 14, 307–314 (2013).
Fre, S., Bardin, A., Robine, S. & Louvard, D. Notch signaling in intestinal homeostasis across species: the cases of Drosophila, Zebrafish and the mouse. Exp. Cell Res. 317, 2740–2747 (2011).
Harder, J. L., Whiteman, E. L., Pieczynski, J. N., Liu, C. J. & Margolis, B. Snail destabilizes cell surface Crumbs3a. Traffic 13, 1170–1185 (2012).
Boncompain, G. et al. Synchronization of secretory protein traffic in populations of cells. Nat. Methods 9, 493–498 (2012).
VanDussen, K. L. et al. Notch signaling modulates proliferation and differentiation of intestinal crypt base columnar stem cells. Development 139, 488–497 (2012).
Van Es, J. H. et al. Notch/gamma-secretase inhibition turns proliferative cells in intestinal crypts and adenomas into goblet cells. Nature 435, 959–963 (2005).
Vaccari, T., Lu, H., Kanwar, R., Fortini, M. E. & Bilder, D. Endosomal entry regulates Notch receptor activation in Drosophila melanogaster. J. Cell Biol. 180, 755–762 (2008).
Wang, W. & Struhl, G. Distinct roles for Mind bomb, Neuralized and Epsin in mediating DSL endocytosis and signaling in Drosophila. Development 132, 2883–2894 (2005).
Wang, W. & Struhl, G. Drosophila Epsin mediates a select endocytic pathway that DSL ligands must enter to activate Notch. Development 131, 5367–5380 (2004).
Crosnier, C. et al. Delta-Notch signalling controls commitment to a secretory fate in the zebrafish intestine. Development 132, 1093–1104 (2005).
Martin-Belmonte, F. & Perez-Moreno, M. Epithelial cell polarity, stem cells and cancer. Nat. Rev. Cancer 12, 23–38 (2012).
Yamamoto, M. et al. Mib-Jag1-Notch signalling regulates patterning and structural roles of the notochord by controlling cell-fate decisions. Development 137, 2527–2537 (2010).
Wibowo, I., Pinto-Teixeira, F., Satou, C., Higashijima, S. & Lopez-Schier, H. Compartmentalized Notch signaling sustains epithelial mirror symmetry. Development 138, 1143–1152 (2011).
Liu, Y., Pathak, N., Kramer-Zucker, A. & Drummond, I. A. Notch signaling controls the differentiation of transporting epithelia and multiciliated cells in the zebrafish pronephros. Development 134, 1111–1122 (2007).
Bosse, F., Hasse, B., Pippirs, U., Greiner-Petter, R. & Muller, H. W. Proteolipid plasmolipin: localization in polarized cells, regulated expression and lipid raft association in CNS and PNS myelin. J. Neurochem. 86, 508–518 (2003).
Schier, A. F. et al. Mutations affecting the development of the embryonic zebrafish brain. Development 123, 165–178 (1996).
Doyle, E. L. et al. TAL Effector-Nucleotide Targeter (TALE-NT) 2.0: tools for TAL effector design and target prediction. Nucleic Acids Res. 40, W117–W122 (2012).
Rodriguez-Fraticelli, A. E. et al. The Cdc42 GEF Intersectin 2 controls mitotic spindle orientation to form the lumen during epithelial morphogenesis. J. Cell Biol. 189, 725–738 (2010).
Roux, K. J., Kim, D. I., Raida, M. & Burke, B. A promiscuous biotin ligase fusion protein identifies proximal and interacting proteins in mammalian cells. J. Cell Biol. 196, 801–810 (2012).
Acknowledgements
We thank C. M. Ruiz-Jarabo for her comments on the manuscript and members of the Martin-Belmonte laboratory and Bagnat laboratories for helpful discussions. We thank A. Alvers (Duke University, North Carolina, USA) for helping in the isolation of gut cells, J. Cocchiaro and L. Marjoram (Duke University, North Carolina, USA) for help in gavaging experiments, B. Margolis (University of Michigan, Michigan, USA) for the Crb3/panCrb antibody, R. Jahn (Max Planck Institute for Biophysical Chemistry, Germany) for Stx7 plasmids, M. Robinson (University of Cambridge, UK) for EpsR plasmids, and R. Kopan and J. L. de la Pompa (CNIC, Spain) for Notch plasmids and mib1ta52b embryos. We also thank M. Guerra at the EM Unit for skillful technical assistance. This work was supported by grants from the MINECO (BFU2011-22622) and CONSOLIDER (CSD2009-00016) to F.M-B, by grant SAF2013-44857-R to M.L.T. by NIH innovator grant 1DP2OD006486 to M.B., and by grant AGL2013-48998-C2-2-R to G.A. A.E.R-F was supported by a CSIC JAE PhD fellowship. M.B-F. is a recipient of a Fundación Obra Social ‘La Caixa’ PhD fellowship. G.A. was supported by the Amarouto Program for senior researchers from the Comunidad Autónoma de Madrid.
Author information
Authors and Affiliations
Contributions
A.E.R-F., M.B. and F.M-B. designed the experiments; A.E.R-F., J.B., M.B-F. and G.A. carried out the experiments; A.E.R-F. and F.M-B. wrote the manuscript; G.B. and F.P. designed and constructed RUSH experimental tools; M.J.G-L. and M.L.T. designed and constructed the Notch-ligand tools; N.R-R., M.A.A. and J.M. produced and characterized the mammalian PLLP antibody; A.E.R-F. and G.A. designed and carried out the electron microscopy experiments.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Additional characterization of pllp expression and pllp loss-of-function
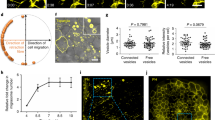
(A) In situ hybridization of pllp probe at 48–72 hpf. Embryos and larvae were collected and fixed at different time points, and incubated with a DIG-labelled PLLP antisense RNA probe and AP-linked anti-DIG antibody. ISH from all different time points were performed at the same time and developed for 2 h before fixation and cleanup. Arrow indicates gut. (B) TgBAC(pllp-GFP) transgenic zebrafish at 48 hpf. Transgenic fish were bred and embryos and larvae were analysed by epifluorescence microscopy. (C) Transverse sections of TgBAC(pllp-GFP) fish at 48–96-hpf. Sections from the posterior midgut (about 1/3 total gut length before the cloaca) were stained to analyse GFP expression using phalloidin (which labels F-actin in apical microvilli) and DAPI (for DNA). Arrows indicate apical localization. Scale bars, 10 μm (magnification, 5 μm). (D) TALEN-generated pllppd1116 mutant null allele. TALENs were generated to target the first exon of zebrafish pllp and injected into 1-cell embryos. After raising the founders, we cloned an allele, pd1116 that harbours a null mutation. We confirmed RNA nonsense mediated decay of the pllp mRNA in the homozygous mutant embryos by measuring pllpmRNA levels in pllppd1116 mutant larvae at 120 hpf by RT-qPCR. Results are mean ± s.d.% expression relative to control and normalized with rRNA 18S expression (n = 3 RNA extracts; ∗P < 0.05 (Student’s t test); Statistic source data can be found in Supplementary Table 3). (E) Live whole-mount images of WT and pllppd1116 larvae at 144 hpf. (F) Quantification of total number of intestinal cells per section of 6 dpf pllppd1116 larvae. 144 hpf larvae were fixed, sectioned and stained with DAPI and F-actin to quantify the total number of intestinal cells per section. Results are represented as mean ± s.d. total number of nuclei in a 2 μm-thick cross-sections (n = 10 sections from 5 WT and 5 pllppd1116 fish selected from 3 independent experiments). (G) Toluidin-stained EM sections of pllppd1116 mutant larvae (400x magnification). Yellow arrows indicate enlarged immature endocytic compartments. Red bars are shown to compare the difference in cell height. Scale bars, 5 μm. (H) Silencing of TgBAC(pllp-GFP) in morpholino-injected 4 dpf larvae. TgBAC(pllp-GFP) embryos were injected with MO1 and MO2 morpholinos at one-cell stage and allowed to grow until 4 dpf. Whole animal lysates from 20 larvae were analysed by Western blot, using anti-GFP antibodies. (I) Quantification of % of morpholino-injected larvae presenting disrupted epithelial columnar organization. WT embryos were injected with MO1 and MO2 morpholinos at one-cell stage and grown until 4 dpf. Posterior gut sections were stained with Phalloidin (green) and DAPI (red) and analysed by confocal microscopy. Data are presented as % of total larvae with disrupted columnar organization (n = 58 control, 25 MO1 and 20 MO2-injected larvae pooled from three independent morpholino injection experiments; ∗P < 0.05 (Student’s t test)). (J) Confocal images showing the phenotype of morpholino-injected larvae quantified in (I). L, lumen. Scale bars, 5 μm. (K) Quantification of microvilli length measured in TEM at 6000x magnification. Results are represented as mean ± s.d. length (in μm) (n = 20 microvilli length averages per cell from 5 WT and 5 mutant cells, randomly selected from 2 independent experiments; ∗P < 0.05 (Student’s t test)). (L) Quantification of apical endosomes in WT and pllppd1116 larvae. Diameters of apical endosomes (within 0-3 μm from the apical surface) were measured from TEM images. Results are represented as mean ± s.d. number of endosomes per cell (n = 10 cells from 5 WT and 5 mutant fish, randomly selected from 2 independent experiments; ∗P < 0.05 (Student’s t test)) (M) Quantification of apical membrane expansion in pllppd1116 larvae. Perimeter of the whole luminal apical membrane (red) was divided by the perimeter of the basal membrane (blue). Results are represented as mean ± s.d. apical/basal perimeter ratios per cell (n = 20 cells per condition, from 4 control and 4 mutant fish, randomly selected from 4 independent experiments; ∗P < 0.05 (Student’s t test)). (N) Quantification of larvae survival in the first 18 dpf. WT and pllppd1116 5 dpf larvae (n = 25) were raised in 1L tanks with limited food supply and were assessed by observing heartbeat every day until 18 dpf. Solid lines indicate WT larvae. Dotted lines indicate pllppd1116 larvae. (O) Rescue of dextran endocytosis in pllppd1116 larvae carrying the TgBAC(pllp-GFP)pd1115 transgene. 144 hpf larvae were gavaged with Dextran-TR, incubated for 2 h, sectioned and stained with DAPI (blue, DNA). Scale bars, 10 μm. (P) Quantification of phenotype rescue. 144 hpf larvae from a cross of TgBAC(pllp-GFP) pllp+/pd1116 with pllppd1116/pd1116 fish were gavaged with Dextran-TR, incubated for 2 h, sorted by transgenic GFP expression, and then the percentage of larvae with disrupted endocytosis (<50%) and disrupted cell height (<15μm) phenotypes were quantified in each case (GFP+, N = 39; GFP-, N = 85). (Q) Rescue of Rab11 subapical localization in pllppd1116 larvae carrying the TgBAC(pllp-GFP) transgene. 144 hpf WT larvae were fixed, sectioned and stained with the anti-Rab11 antibody (red) and DAPI (blue, DNA). Scale bars, 10 μm.
Supplementary Figure 2 PLLP is required for epithelial morphogenesis in 3D-MDCK.
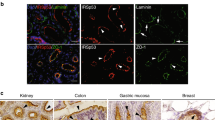
(A) Tubular sorting endosomes in apical endosomes in WT and pllppd1116 larvae. 144 hpf larval gut sections were analysed by TEM and the number of connected tubular structures as depicted in the micrograph (orange) was quantified. Scale, 100 nm. (B) Localization of endogenous PLLP in mouse intestinal cells. Agarose-embedded sections of 1 month-old BL6 mouse small intestines were stained with the PLLP antibody (green), Phalloidin (red) and DAPI (blue, DNA). A magnification of cells from a villar tip is shown. Scale bars, 10 μm. (C) Localization of endogenous PLLP in mouse kidney cells. Agarose-embedded sections of 1 month-old BL6 mouse kidneys were stained with the PLLP antibody (green), Phalloidin (red) and DAPI (blue, DNA). Magnifications of a medullar and a cortical kidney tubule are shown. Scale bars, 5 μm (magnification, 2 μm). (D) Silencing of PLLP expression in MDCK cysts. MDCK cells transfected with control or PLLP-specific siRNAs were grown to form cysts and lysed after 72 h. Westernblot analysis was performed to quantify PLLP protein levels using control or different PLLP-specific siRNAs oligos. (E) Phenotype of PLLP-KD in MDCK cysts. MDCK cells transfected with control or PLLP-specific siRNAs were grown to form cysts and fixed after 72 h. MDCK cysts were labelled with anti-PLLP antibody (green) and analysed by confocal microscopy with DIC. Notice the disruption of PLLP antibody signal in PLLP-KD cysts. Scale bars, 5 μm. (F) PLLP and Rab11a colocalization in MDCK cysts. MDCK cells stably expressing PLLP-GFP were grown to form cysts and fixed after 72 h. MDCK cysts were labelled with anti-Rab11 antibody (red), anti-βcatenin (blue) and analysed by confocal microscopy. Arrow indicates apical recycling endosome. Scale bars, 5 μm. (G) Rab11 localization in PLLP-KD cysts. MDCK cells transfected with control or PLLP-specific siRNAs were grown to form cysts and fixed after 72 h. MDCK cysts were labelled with anti-Rab11 antibody (red), anti-Podxl (green) DAPI (DNA, blue) and analysed by confocal microscopy with DIC. Right panels show whole z-axis projection of the Rab11 signal surrounding a single lumen. Arrows indicates apical recycling endosome. Scale bars, 5 μm. (H) Subcellular EM localization of PLLP-GFP aggregates in overexpressing cells. PLLP-GFP overexpressing cells were FACS-sorted at 48 h, pelleted, seeded into monolayers, and fixed after 24 h. Then, cell pellets were processed for immunogold-EM using anti-GFP and 15 nm protein-A gold beads. Scale bars, 200 nm (magnification, 100 nm). (I) In vivo overexpression of Pllp-GFP in TgBAC(pllp-GFP)pd1114 at 5 dpf. Posterior gut sections of 5 dpf larvae carrying a TgBAC(pllp-GFP)pd1114 overexpressing allele were fixed and stained with Phalloidin (red) and DAPI (blue, DNA). Scale bars, 5μm.
Supplementary Figure 3 EpsR and Stx7 are required for epithelial morphogenesis in 3D-MDCK.
(A) EpsR localization in PLLP-Cherry cells. MDCK cells stably expressing PLLP-Cherry were fixed after 72 h, labelled with anti-EpsR antibody (green) and analysed by confocal microscopy. An x-y section of the apical region of a single cell is shown. Arrows indicate colocalization of PLLP and EpsR. Scale bars, 2 μm. (B) Quantification of disrupted Rab7-GFP and EpsR fluorescent staining in PLLP-KD cysts. GFP-Rab7 MDCK cells were transfected with control or PLLP-specific siRNA, grown to form cysts and fixed after 72 h. MDCK cysts were labelled with anti-EpsR and PLLP antibodies and analysed by confocal microscopy to measure fluorescent intensity. Data represented are fold-reduction values of GFP-Rab7 and EpsR staining compared to control (n = 25 control and 25 PLLP-KD cells from 3 independent experiments; ∗P < 0.05 (Student’s t test)). Only cells with less than 10% of control PLLP expression were counted. (C) Silencing of EpsR expression using siRNA. MDCK cells were transfected with control or EpsR-specific siRNAs, grown to form cysts and lysed after 72 h. Westernblot analysis was performed to quantify endogenous EpsR protein levels. (D) Rab11 localization in EpsR-KD cysts. MDCK cells were transfected with control or EpsR-specific siRNAs, grown to form cysts and fixed after 72 h. MDCK cysts were labelled with anti-Rab11 antibody (red), E-cadherin (green) and DNA (blue) and analysed by confocal microscopy. Arrows indicate dispersed Rab11 endosomes in EpsR-KD cysts. Scale bars, 5 μm. (E) Requirement of EpsR for oversized Rab11 compartment in PLLP-GFP overexpression phenotype. PLLP-GFP overexpressing cells were co-transfected with control or EpsR siRNAs, grown for 48 h, fixed and stained to detect Rab11 (red). Scale bars, 5 μm. (F) Endogenous subapical Stx7 localization in mouse small intestinal villi. Agarose-embedded sections of 1 month-old BL6 mice small intestines were stained with the Stx7 antibody (green), Phalloidin (red) and DAPI (blue, DNA). A magnification of cells from a villar tip is shown. Scale bars, 20 μm (magnification, 10 μm). (G) Stx7 and PLLP probe ligation assay (PLA). MDCK cells stably expressing WT untagged PLLP were transiently transfected with GFP mock, Rab11-GFP or Stx7-GFP plasmids. PLA was performed with mouse anti-GFP (green) and rabbit anti-PLLP antibodies and whole-cell maximum z-stack projections of the PLA signal (red) were analysed by confocal microscopy Arrow indicates colocalization of PLA signal and Stx7-GFP. Scale bars, 10 μm. (H) Quantification of PLLP PLA. Integrated density of the PLA fluorescent signal was quantified in 16-bit images of maximum z-stack projections and represented as fold-increase over GFP background signal for GFP-Rab11 and GFP-Stx7. Data are average PLA intensity shown as base-2 logarithm of the signal-to-background ratio (n = 11 GFP, 15 GFP-Rab11 and 12 GFP-Stx7 cells selected from 2 independent experiments). (I) Phenotype rescue of Stx7-KD MDCK cysts. MDCK cells stably expressing siRNA resistant Stx7-GFP were transfected with control or Stx7-specific siRNAs, grown to form cysts and fixed after 72 h. MDCK cysts were labelled with anti-Podxl antibody (red) and βcatenin (blue) and analysed by confocal microscopy. Scale bars, 5 μm. (J) Silencing of Stx7 expression using siRNA. WT MDCK cells or MDCK cells stably expressing Stx7-GFP were transfected with control or Stx7-specific siRNAs, grown to form cysts and lysed after 72 h. Westernblot analysis was performed to quantify Stx7 protein levels. ∗ Unspecific bands from the antibody. (K) Quantification of lumen formation phenotypes in Stx7-KD MDCK cysts and phenotype rescue. Measurements are normalized to WT MDCK cells (control) and expressed as mean ± s.d. percentage relative to control of single lumen-forming cysts (Control, 100 ± 5.3%; Stx7-KD, 52.3 ± 6.4%; Control GFP-Stx7, 97.0 ± 0.2%; Stx7-KD GFP-Stx7, 98.4 ± 5.0%; n = 3 independent siRNA experiments; ∗∗∗P < 0.005 (Student’s t test); Statistic source data can be found in Supplementary Table 3). (L) Rab11 localization in Stx7-KD cysts. MDCK cells were transfected with control or Stx7-specific siRNAs, grown to form cysts and fixed after 72 h. MDCK cysts were labelled with anti-Rab11 antibody (red), Phalloidin (green) and E-cadherin (blue) and analysed by confocal microscopy. Arrows indicate apical recycling endosomes. Scale bars, 5 μm. (M) Stx7 localization in 44 hpf pllppd1116 mutant larvae. 96 hpf WT or pllppd1116 mutant larvae were fixed with acetone, sectioned and stained with the anti-Stx7 antibody (green), and DAPI (red, DNA). Arrows indicate apical Stx7 localization. Notice the mislocalized dispersed Stx7 in pllppd1116 larvae. Scale bars, 10 μm.
Supplementary Figure 4 Additional analysis on Crb and Notch signaling.
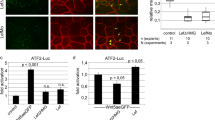
(A) Quantification of apical membrane localization of Crb in cysts at different time points. Pearson colocalization coefficients were calculated by staining Podxl and Crb3 at different time points. Data represented are averaged Pearson coefficients ± s.d. (n = 5 averaged colocalization coefficients from 10 cells per condition, selected from 2 independent experiments; ∗P < 0.05 (Student’s t test)). (B) Quantification of linear correlation of Crb3 and ZO1 linear profiles. Linear intensity profiles were drawn at the apical plasma membrane and across 4 tight junctions. Data represented are averaged linear (monodimensional) Pearson correlation coefficients ± s.d. (n = 6 linear profiles taken from 3 cysts per condition, randomly selected from 3 independent siRNA experiments; ∗P < 0.05 (Student’s t test)). (C) Example ZO1 and Crb3 linear profiles of control-transfected and PLLP siRNA-transfected cysts. Red line, ZO1; green line, Crb3. Data are represented as % of maximum signal intensity (for each channel). TJ, position of tight junction (determined by peak in ZO1 staining). (D) Western blot of ligand-mediated Notch-1a transactivation assays in MDCK-II/OP9 cocultures. MDCK cells stably expressing Notch1a-myc are polarized over a monolayer of mesenchymal OP9 cells stably expressing GFP or Jagged-1 or Delta-like 1. Activated Notch1a is quantified using anti-Cleaved Notch1a (Val1744). Tubulin is used as loading control. (E) MDCK coculture system to analyse Notch1 transactivation by Jagged-1. Stable MDCK cell lines expressing Notch1 and Jagged-1 are mixed (4:1) and seeded to grow polarized cysts for 48 h. (F) Western blot of ligand-mediated Notch-1a transactivation assays in MDCK cocultures. MDCK cells stably expressing Notch1a-myc are mixed (4:1) with GFP or Jagged-1 expressing MDCK cells and grown in cysts (100,000 cells/ml) for 48 h. Activated Notch1a is quantified using anti-Cleaved Notch1a (Val1744). Tubulin is a loading control. (G) Effect of PLLP-KD or EpsR-KD in Notch1 transactivation by Jagged-1. MDCK cells stably expressing Notch1 or Jagged-1 were transfected with control, PLLP or EpsR siRNA, mixed as indicated and seeded to grow polarized cysts for 48 h. Cleaved Notch1 protein levels were analysed by Western blot. GAPDH is a loading control. (H) Quantification of Notch1 transactivation by Jagged-1 on PLLP-KD or EpsR-KD. Bars are colour coded to indicate control (black), PLLP (white) or EpsR (grey) silencing in either Notch (N) or Jagged-1 (J) MDCK cells. Data are expressed as mean ± s.d. as % of control (n = 5 extracts from 5 independent experiments; ∗P < 0.05 (Student’s t test)).
Supplementary Figure 5 Unedited Western-blot scans.
Cropped regions are indicated with a red rectangule as appropriate. Antibodies and concentrations are provided for each Western-blot.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1672 kb)
Supplementary Table 1
Supplementary Information (XLSX 55 kb)
Supplementary Table 2
Supplementary Information (XLS 628 kb)
Supplementary Table 3
(XLSX 55 kb)
Stx7/PLLP FRAP experiment.
MDCK cells stably expressing PLLP-Cherry were transfected with Stx7-GFP and imaged using confocal microscopy every 4 s. The outlined region of interest was photobleached using the 488 nm and 564 nm lasers for 12 rounds after the 4th frame. Frame rate, 10 s−1. (MOV 7700 kb)
Crb3 RUSH in MDCK cysts.
MDCK cells stably expressing Streptavidin-KDEL and EGFP-SBP-Crb3a were cultured to grow cysts for 72 h. Biotin was added at t = 0 and cysts were imaged using confocal microscopy (pinhole = 1 μm) every 1 min. Frame rate, 3 s−1. (AVI 1355 kb)
Crb3 RUSH in control cells.
MDCK cells stably expressing Streptavidin-KDEL and EGFP-SBP-Crb3a were transfected with control siRNA and cultured to grow cysts for 72 h. Biotin was added at t = 0 and cysts were imaged using confocal microscopy (pinhole = 1 μm) every 1 min. Frame rate, 3 s−1. (AVI 326 kb)
Crb3 RUSH in PLLP-KD cells.
MDCK cells stably expressing Streptavidin-KDEL and EGFP-SBP-Crb3a were transfected with PLLP-specific siRNA and cultured to grow cysts for 72 h. Biotin was added at t = 0 and cysts were imaged using confocal microscopy (pinhole = 1 μm) every 1 min. Frame rate, 3 s−1. (AVI 595 kb)
Crb3 RUSH in Dynasore-treated cells.
MDCK cells stably expressing Streptavidin-KDEL and EGFP-SBP-Crb3a were cultured to grow cysts for 72 h. Biotin was added at t = 0 and cysts were imaged using confocal microscopy (pinhole = 1 μm) every 1 min. Dynasore was added at 50 μM after 40 min. Frame rate, 3 s−1. (AVI 460 kb)
Rights and permissions
About this article
Cite this article
Rodríguez-Fraticelli, A., Bagwell, J., Bosch-Fortea, M. et al. Developmental regulation of apical endocytosis controls epithelial patterning in vertebrate tubular organs. Nat Cell Biol 17, 241–250 (2015). https://doi.org/10.1038/ncb3106
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3106
This article is cited by
-
Voltage-sensing phosphatase (Vsp) regulates endocytosis-dependent nutrient absorption in chordate enterocytes
Communications Biology (2022)
-
Plasmolipin regulates basolateral-to-apical transcytosis of ICAM-1 and leukocyte adhesion in polarized hepatic epithelial cells
Cellular and Molecular Life Sciences (2022)
-
MALL, a membrane-tetra-spanning proteolipid overexpressed in cancer, is present in membraneless nuclear biomolecular condensates
Cellular and Molecular Life Sciences (2022)
-
Transgenic fluorescent zebrafish lines that have revolutionized biomedical research
Laboratory Animal Research (2021)
-
Proximity interactions of the ubiquitin ligase Mind bomb 1 reveal a role in regulation of epithelial polarity complex proteins
Scientific Reports (2019)