Abstract
The cell-biological program termed the epithelial–mesenchymal transition (EMT) confers on cancer cells mesenchymal traits and an ability to enter the cancer stem cell (CSC) state. However, the interactions between CSCs and their surrounding microenvironment are poorly understood. Here we show that tumour-associated monocytes and macrophages (TAMs) create a CSC niche through juxtacrine signalling with CSCs. We performed quantitative proteomic profiling and found that the EMT program upregulates the expression of CD90, also known as Thy1, and EphA4, which mediate the physical interactions of CSCs with TAMs by directly binding with their respective counter-receptors on these cells. In response, the EphA4 receptor on the carcinoma cells activates Src and NF-κB. In turn, NF-κB in the CSCs induces the secretion of a variety of cytokines that serve to sustain the stem cell state. Indeed, admixed macrophages enhance the CSC activities of carcinoma cells. These findings underscore the significance of TAMs as important components of the CSC niche.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Huber, M. A., Kraut, N. & Beug, H. Molecular requirements for epithelial-mesenchymal transition during tumor progression. Curr. Opin. Cell Biol. 17, 548–558 (2005).
Thiery, J. P., Acloque, H., Huang, R. Y. & Nieto, M. A. Epithelial-mesenchymal transitions in development and disease. Cell 139, 871–890 (2009).
Guo, W. et al. Slug and Sox9 cooperatively determine the mammary stem cell state. Cell 148, 1015–1028 (2012).
Mani, S. A. et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 133, 704–715 (2008).
Morel, A. P. et al. Generation of breast cancer stem cells through epithelial-mesenchymal transition. PLoS ONE 3, e2888 (2008).
Buck, E. et al. Loss of homotypic cell adhesion by epithelial-mesenchymal transition or mutation limits sensitivity to epidermal growth factor receptor inhibition. Mol. Cancer Ther. 6, 532–541 (2007).
Creighton, C. J. et al. Residual breast cancers after conventional therapy display mesenchymal as well as tumor-initiating features. Proc. Natl Acad. Sci. USA 106, 13820–13825 (2009).
Gupta, P. B. et al. Identification of selective inhibitors of cancer stem cells by high-throughput screening. Cell 138, 645–659 (2009).
Li, H. J., Reinhardt, F., Herschman, H. R. & Weinberg, R. A. Cancer-stimulated mesenchymal stem cells create a carcinoma stem cell niche via prostaglandin E2 signaling. Cancer Discov. 2, 840–855 (2012).
Malanchi, I. et al. Interactions between cancer stem cells and their niche govern metastatic colonization. Nature 481, 85–89 (2011).
Merlos-Suarez, A. et al. The intestinal stem cell signature identifies colorectal cancer stem cells and predicts disease relapse. Cell Stem Cell 8, 511–524 (2011).
Wynn, T. A., Chawla, A. & Pollard, J. W. Macrophage biology in development, homeostasis and disease. Nature 496, 445–455 (2013).
Coussens, L. M., Zitvogel, L. & Palucka, A. K. Neutralizing tumor-promoting chronic inflammation: a magic bullet? Science 339, 286–291 (2013).
Gyorki, D. E., Asselin-Labat, M. L., Rooijen, N. v., Lindeman, G. J. & Visvader, J. E. Resident macrophages influence stem cell activity in the mammary gland. Breast Cancer Res. 11, R62 (2009).
Elenbaas, B. et al. Human breast cancer cells generated by oncogenic transformation of primary mammary epithelial cells. Genes Dev. 15, 50–65 (2001).
Ong, S. E. et al. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol. Cell. Proteom. 1, 376–386 (2002).
Udeshi, N. D. et al. Methods for quantification of in vivo changes in protein ubiquitination following proteasome and deubiquitinase inhibition. Mol. Cell. Proteomics 11, 148–159 (2012).
Barker, T. H. & Hagood, J. S. Getting a grip on Thy-1 signaling. Biochim. Biophys. Acta. 1793, 921–923 (2009).
Wetzel, A. et al. Human Thy-1 (CD90) on activated endothelial cells is a counterreceptor for the leukocyte integrin Mac-1 (CD11b/CD18). J. Immunol. 172, 3850–3859 (2004).
Lim, E. et al. Transcriptome analyses of mouse and human mammary cell subpopulations reveal multiple conserved genes and pathways. Breast Cancer Res. 12, R21 (2010).
Shipitsin, M. et al. Molecular definition of breast tumor heterogeneity. Cancer Cell 11, 259–273 (2007).
Dontu, G. et al. In vitro propagation and transcriptional profiling of human mammary stem/progenitor cells. Genes Dev. 17, 1253–1270 (2003).
Tsuchiya, S. et al. Establishment and characterization of a human acute monocytic leukemia cell line (THP-1). Int. J. Cancer 26, 171–176 (1980).
Donnenberg, V. S. et al. Localization of CD44 and CD90 positive cells to the invasive front of breast tumors. Cytometry B Clin. Cytom. 78, 287–301 (2010).
Donnenberg, A. D., Zimmerlin, L., Landreneau, R. J., Luketich, J. D. & Donnenberg, V. S. KIT (CD117) expression in a subset of non-small cell lung carcinoma (NSCLC) patients. PLoS ONE 7, 1–11 (2012).
Medrek, C., Ponten, F., Jirstrom, K. & Leandersson, K. The presence of tumor associated macrophages in tumor stroma as a prognostic marker for breast cancer patients. BMC Cancer 12, 306 (2012).
Tsutsui, S. et al. Macrophage infiltration and its prognostic implications in breast cancer: the relationship with VEGF expression and microvessel density. Oncol. Rep. 14, 425–431 (2005).
Ch’ng, E. S., Tuan Sharif, S. E. & Jaafar, H. In human invasive breast ductal carcinoma, tumor stromal macrophages and tumor nest macrophages have distinct relationships with clinicopathological parameters and tumor angiogenesis. Virchows Arch. 462, 257–267 (2013).
Qian, B. Z. & Pollard, J. W. Macrophage diversity enhances tumor progression and metastasis. Cell 141, 39–51 (2010).
Murray, P. J. & Wynn, T. A. Protective and pathogenic functions of macrophage subsets. Nat. Rev. Immunol. 11, 723–737 (2011).
Lin, E. Y., Nguyen, A. V., Russell, R. G. & Pollard, J. W. Colony-stimulating factor 1 promotes progression of mammary tumors to malignancy. J. Exp. Med. 193, 727–740 (2001).
Ohri, C. M., Shikotra, A., Green, R. H., Waller, D. A. & Bradding, P. The tissue microlocalisation and cellular expression of CD163, VEGF, HLA-DR, iNOS, and MRP 8/14 is correlated to clinical outcome in NSCLC. PLoS ONE 6, e21874 (2011).
Gabrilovich, D. I., Ostrand-Rosenberg, S. & Bronte, V. Coordinated regulation of myeloid cells by tumours. Nat. Rev. Immunol. 12, 253–268 (2012).
Hartman, Z. C. et al. Growth of triple-negative breast cancer cells relies upon coordinate autocrine expression of the proinflammatory cytokines IL-6 and IL-8. Cancer Res. (2013).
Korkaya, H., Liu, S. & Wicha, M. S. Regulation of cancer stem cells by cytokine networks: attacking cancer’s inflammatory roots. Clin. Cancer Res. 17, 6125–6129 (2011).
Wyckoff, J. et al. A paracrine loop between tumor cells and macrophages is required for tumor cell migration in mammary tumors. Cancer Res. 64, 7022–7029 (2004).
Tam, W. L. et al. Protein Kinase C alpha is a central signaling node and therapeutic target for breast cancer stem cells. Cancer Cell 24, 347–364 (2013).
Thomson, S., Petti, F., Sujka-Kwok, I., Epstein, D. & Haley, J. D. Kinase switching in mesenchymal-like non-small cell lung cancer lines contributes to EGFR inhibitor resistance through pathway redundancy. Clin. Exp. Metastasis 25, 843–854 (2008).
Gentile, A., Lazzari, L., Benvenuti, S., Trusolino, L. & Comoglio, P. M. Ror1 is a pseudokinase that is crucial for Met-driven tumorigenesis. Cancer Res. 71, 3132–3141 (2011).
Pasquale, E. B. Eph receptors and ephrins in cancer: Bidirectional signalling and beyond. Nat. Rev. Cancer 10, 165–180 (2010).
Lamberto, I. et al. Distinctive binding of three antagonistic peptides to the ephrin-binding pocket of the EphA4 receptor. Biochem. J. 445, 47–56 (2012).
Zhou, L. et al. EphA4 signaling regulates phospholipase Cgamma1 activation, cofilin membrane association, and dendritic spine morphology. J. Neurosci. 27, 5127–5138 (2007).
Griner, E. M. & Kazanietz, M. G. Protein kinase C and other diacylglycerol effectors in cancer. Nat. Rev. Cancer 7, 281–294 (2007).
Kalo, M. S. & Pasquale, E. B. Signal transfer by eph receptors. Cell Tissue Res. 298, 1–9 (1999).
Lu, Z. G., Liu, H., Yamaguchi, T., Miki, Y. & Yoshida, K. Protein kinase Cdelta activates RelA/p65 and nuclear factor-kappaB signaling in response to tumor necrosis factor-alpha. Cancer Res. 69, 5927–5935 (2009).
Hoffmann, E., Dittrich-Breiholz, O., Holtmann, H. & Kracht, M. Multiple control of interleukin-8 gene expression. J. Leukoc. Biol. 72, 847–855 (2002).
Sasaki, C. Y., Ghosh, P. & Longo, D. L. Recruitment of RelB to the Csf2 promoter enhances RelA-mediated transcription of granulocyte-macrophage colony-stimulating factor. J. Biol. Chem. 286, 1093–1102 (2011).
Yoon, S. et al. NF-kappaB and STAT3 cooperatively induce IL6 in starved cancer cells. Oncogene. 31, 3467–3481 (2012).
Ryan, K. M., Ernst, M. K., Rice, N. R. & Vousden, K. H. Role of NF-kappaB in p53-mediated programmed cell death. Nature 404, 892–897 (2000).
Li, S. et al. TWIST1 associates with NF-kappaB subunit RELA via carboxyl-terminal WR domain to promote cell autonomous invasion through IL8 production. BMC Biol. 10, 73 (2012).
Gisina, A. M. et al. Detection of minor subpopulations of colorectal adenocarcinoma cells expressing cancer stem cell markers. Bull. Exp. Biol. Med. 151, 234–238 (2011).
He, J. et al. CD90 is identified as a candidate marker for cancer stem cells in primary high-grade gliomas using tissue microarrays. Mol. Cell. Proteomics 11, M111 010744 (2012).
Jiang, J. et al. Trastuzumab (herceptin) targets gastric cancer stem cells characterized by CD90 phenotype. Oncogene. 31, 671–682 (2012).
Yang, Z. F. et al. Significance of CD90 + cancer stem cells in human liver cancer. Cancer Cell 13, 153–166 (2008).
Oki, M. et al. Overexpression of the receptor tyrosine kinase EphA4 in human gastric cancers. World J. Gastroenterol. 14, 5650–5656 (2008).
Oshima, T. et al. Overexpression of EphA4 gene and reduced expression of EphB2 gene correlates with liver metastasis in colorectal cancer. Int. J. Oncol. 33, 573–577 (2008).
Binda, E. et al. The EphA2 receptor drives self-renewal and tumorigenicity in stem-like tumor-propagating cells from human glioblastomas. Cancer Cell 22, 765–780 (2012).
Gyorffy, B. et al. An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res. Treat. 123, 725–731 (2010).
Orimo, A. et al. Stromal fibroblasts present in invasive human breast carcinomas promote tumor growth and angiogenesis through elevated SDF-1/CXCL12 secretion. Cell 121, 335–348 (2005).
Yu, M. et al. A developmentally regulated inducer of EMT, LBX1, contributes to breast cancer progression. Genes Dev. 23, 1737–1742 (2009).
Shibue, T. & Weinberg, R. A. Integrin beta1-focal adhesion kinase signaling directs the proliferation of metastatic cancer cells disseminated in the lungs. Proc. Natl Acad. Sci. USA 106, 10290–10295 (2009).
Tergaonkar, V., Pando, M., Vafa, O., Wahl, G. & Verma, I. p53 stabilization is decreased upon NFkappaB activation: A role for NFkappaB in acquisition of resistance to chemotherapy. Cancer Cell 1, 493–503 (2002).
Stewart, S. A. et al. Lentivirus-delivered stable gene silencing by RNAi in primary cells. RNA 9, 493–501 (2003).
Han, J. et al. Tbx3 improves the germ-line competency of induced pluripotent stem cells. Nature 463, 1096–1100 (2010).
Hu, Y. & Smyth, G. K. ELDA: extreme limiting dilution analysis for comparing depleted and enriched populations in stem cell and other assays. J. Immunol. Methods 347, 70–78 (2009).
Hammond, S. L., Ham, R. G. & Stampfer, M. R. Serum-free growth of human mammary epithelial cells: rapid clonal growth in defined medium and extended serial passage with pituitary extract. Proc. Natl Acad. Sci. USA 81, 5435–5439 (1984).
Naba, A. et al. The matrisome: In silico definition and in vivo characterization by proteomics of normal and tumor extracellular matrices. Mol. Cell. Proteomics 11, M111 014647 (2012).
Acknowledgements
We thank L. Ma and T. Shibue for critical reviews of the manuscript, R. Bronson for evaluating the histopathology, D. R. Mani and P. Thiru for statistical analyses and C. Baty for assistance in imaging. We also thank A. Oberle, J. Karlsson and E. Procopio for technical assistance. This research was supported by the MIT Ludwig Center for Molecular Oncology and by grants from the Breast Cancer Research Foundation and National Institutes of Health (NIH; R01-CA078461, P01-CA080111 and U54-CA163109) to R.A.W. Additional support comes in part from the Department of Defense (BC032981 and BC044784), NIH (P30CA047904), the Hillman Foundation and the Glimmer of Hope Foundation to V.S.D. This work was also supported in part by the Broad Institute of MIT and Harvard, and by grants from NIH (U24CA160034) to S.A.C. R.A.W. is an American Cancer Society and D. K. Ludwig Foundation Cancer Research Professor. Postdoctoral fellowship support for H.L. was from the Cancer Research Institute (New York) and the MIT Ludwig Center for Molecular Oncology.
Author information
Authors and Affiliations
Contributions
H.L. designed the project, conducted experiments and analysed data under the guidance of R.A.W. K.R.C. and S.A.C. performed the proteomics profiling and data analysis. V.S.D. and R.B. performed patient sample staining, imaging and quantification. W.L.T. performed gene microarray, chromatin immunoprecipitation and data analysis. H.L. and X.Y. performed immunofluorescence and imaging of mouse tumour tissues. J.F. contributed to signalling pathway analysis and E.N.E. provided technical expertise. F.R. performed mouse surgeries. H.L. and R.A.W. wrote the manuscript. All authors critically read and contributed to the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Proteomics profiling of EMT-induced changes in membrane proteins.
(a) For stable isotope labeling, the HMLE-Twist cells and control HMLE-vector cells were maintained in the same medium, except that the medium for one of the two cell types contained the C13N15 Arg and Lys amino acids (heavy) instead of the standard C12N14 Arg and Lys (light). A replicate experiment was subsequently performed in which the light/heavy labels were swapped in order to reduce noise from proteins present in the culture medium. Following culture, the HMLE-vector and HMLE-Twist cells were mixed in equal numbers; a membrane protein fraction was prepared from the mixture (panel b) and subjected to tryptic digestion. The resulting peptides were further resolved by off-gel electrophoresis and analyzed by LC-MS/MS (liquid chromatography-tandem mass spectrometry). (b) A flow chart of the membrane protein fractionation strategy. (c) Western blot detecting protein enrichment and contamination in different fractions. E-cadherin and N-cadherin, plasma membrane proteins; Large T antigen, nuclear protein; β-actin, cytosolic and cytoskeletal protein. Numbers on the gels indicate molecular weight markers in kDa.
Supplementary Figure 2 CD90 expression is enriched in the MaSCs and CSCs.
(a) CD90 mRNA levels in HMLE-derived cell lines detected by expression array. HMLE and HMLE-vector are epithelial control cells, while Twist, Snail, Slug and Zeb1 cells have undergone EMT driven by these individual EMT-TFs. (b) Enrichment in gene expression of THY1 (CD90) and three other EMT-related genes in the human MaSC-enriched subset; plotted based on data from Lim et.al (2010)20. (c) Expression of CD90 in CD44+/PROCR+ stem/progenitor cells or CD24+ differentiated cells purified from human breast tumors; plotted based on data from Shipitsin et.al (2007)21. Results are presented as the counts of SAGE tags for THY1 (CD90) in 10 SAGE libraries, The ratio and P value shown is between CD44+/PROCR+ (n = 5; 1 normal, 4 tumors) and CD24+ (n = 5; 2 normal, 3 tumors) purified cells. Student’s t-test performed. Error bars indicate S.D. (d) Tumor incidence of sorted CD90hiCD24− HMLER cells in Nude mice compared to the bulk, CD90medCD24+/− or CD90lo24+ populations. (e) MB-MDA-435 cells sorted based on CD90 expression. (f) Tumor weight and (g) H&E staining of xenograft tumors from sorted CD90low and CD90high MB-MDA-435 cells. Data from two independent experiments were pooled (f). Error bar indicate S.E.M. ∗∗p < 0.01, ∗∗∗p < 0.001 (Student’s t-test). n = 10, 10, 8, 10, 9, 10 mice (left to right). The CD90high cells formed tumors with more aggressive histopathological characteristics, including larger nuclei and elevated stromal cell infiltration. (h) Flow cytometry profile showing CD90 and CD24 levels for the CD90hi, shCD90 in CD90hi, and CD90lo HMLER cells.
Supplementary Figure 3 CD90 expression in patient breast cancer tissue.
(a) Lower magnification images of the tumors 042 (25 ×) and 032 (20 ×) as in Fig. 3G. CSC (red)-TAM (green) pairs were highlighted with red boxes in tumor 032. Tumor 042 has higher contents of CSCs and TAMs and extensive CSC-TAM contacts were observed. (b) CTK+CD90− cancer cells (green) rarely showed contact with nearby CD68+ TAMs (blue; ER + BrCA041), whereas CD90+ CSCs (red) that have partial loss of CTK (yellow arrowheads) can be observed adjacent to TAMs (ER- BrCA042). No CD68+CTK+ cells were observed, indicating that the CD68 antibody is not staining non-specifically for cancer cells. (c) Flow cytometry profiles of breast cancer cells from freshly excised ER − tumor BrCA042. Aneuploid cells were gated by DAPI > 2n (control 2n defined using diploid lymphocytes). The CD90 and CD44 flow cytometry profiles of both cytokeratin + (CTK+) aneuploid cells (green panel) and CTKneg aneuploid cells (blue panel) were shown. (d) Expression of CD44 and CD90 on cytokeratin + (iCTK+) breast cancer cells in ER + tumor BrCA032. 33.7% of hematopoietic lineage negative cells (Non-Heme) were CTK+, about a third of which were aneuploid. 8.6% of all CTK+ cells coexpressed CD90 and CD44. (e) THY1 (CD90) expression correlates with CD68 expression in patient breast cancer samples. Pearson’s correlation was obtained using TCGA breast invasive carcinoma gene expression data set downloaded from the UCSC Cancer Browser.
Supplementary Figure 4 TAMs promote tumor growth.
(a) TAMs were isolated by MACS from xenograft tumors derived from HMLER90hi cells. The purity of the TAMs is indicated by the percentage of F4/80+ cells. The viability of the TAMs determined by DAPI staining is typically between 75% and 87%. (b) TAMs from HMLER90hi-CSC cells (C1, C2) and CSC + TAM coinjections (CT1-3) and naturally arising tumors in MMTV-PyMT mice (P1–P7) were sorted by F4/80 positivity. Gene expression of the M1/M2 related genes were measured by Q-PCR. (c) Tumor sections from breast cancer patients were stained with CD68, CD163 and HLA-DR. Almost all the CD68+ cells show CD163+HLA-DRlo M2 macrophage phenotype (white arrowheads). Note that there are background signals which are spotted rather than homogenous staining of the cell surface and these cells are not counted as CD68+ TAMs in any of the quantifications in main Fig. 3. (d–l) Naturally arising MMTV-PyMT tumors were dissociated and tumor cells were sorted based on CD90 expression, whereas TAMs were isolated by F4/80 expression. 3 × 105 CD90+ or CD90− tumor cells were orthotopically transplanted into syngeneic wildtype Fvb mice with or without comixed 1.5 × 105 TAMs. Typical images are shown. (d) Primary tumor weight after draining fluid; data were pooled from 3 independent experiments. n = 15 mice. T lymphocyte infiltration was shown by CD3 staining of naturally arising tumor (e) and transplanted primary tumors (f). (g) H&E staining of primary tumors. Ne, necrosis; Cy, fluid-filled cyst. (h) Percentage of necrotic area (including necrosis and cysts) was quantified using Fiji software. n = 9 (3 fields were randomly chosen of 3 tumors for each group). (i) Ki67 staining of primary tumors. (j) Quantification of Ki67+ proliferating cells per field at 20× magnification, n = 15 (5 fields were randomly chosen of 3 tumors for each group). (k) Experimental scheme of the secondary transplantation experiment in main Fig. 4g, h. Error bars indicate S.D. ∗∗p < 0.01; ∗∗∗∗p < 0.0001 (One way ANOVA).
Supplementary Figure 5 The cytokines IL-6 and IL-8 secreted by the CSC-like cells promote CSC survival.
(a) The coculture experiment followed by FACS and RNA harvesting. (b) Knockdown of IL-6 and IL-8 in unsorted epithelial HMLER cells. (c) Proliferation of CD90hi or unsorted HMLER cells with IL-6 or IL-8 knockdown. (d) Doxycyclin-induced IL-8 secretion in the HMLER90hi cells with the TetO-IL8 expression vector +/− the rtTA vector, which is required for induction. (e) Doxycyclin-induced IL-8 secretion resulted in enhanced tumorsphere formation in the HMLER90hi cells. (f) Treatment with recombinant cytokines IL-6 (0.8 ng ml−1) and/or IL-8 (10 ng ml−1) increased tumorsphere formation by the HMLER90hi cells. (b–f) Data plotted are average (b,c) or individual (d–f) values of 3 technical replicates from one of two independent experiments. Source data provided in Supplementary Table 7.
Supplementary Figure 6 Contact with monocytes activates EphA4 and downstream proteins in the stem-like cells.
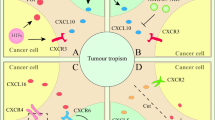
(a) Western blots comparing the protein expression of EphA4 and PDGFRβ in the HMLE-Twist, HMLE-Slug CD90hi and HMLER90hi mesenchymal cells with the HMLE and HMLER90lo epithelial cells. (b) Separation of HMLER90hi or HMLE-Twist cells from cocultured monocytes for western blot analysis. (c) Phospho- and total-protein levels of EphA4, PLCγ1 and PKCδ in HMLE-Twist cells (+/−monocyte-coculture) were detected by western blot. β-actin served as loading control. (d) Q-PCR verifying the knockdown of EphA4 by the shRNA. (e) Cytokine mRNA induction (fold-change relative to corresponding no-monocyte control) in the HMLER90hi cells with sh-Luciferase or sh-EphA4 after 3 h of coculture with the monocytes. Data plotted in d,e are average values of 3 technical replicates from one out of two biological repeats. Source data provided in Supplementary Table 7. (f) Phospho- and total-protein levels of EphA4 in HMLER90hi cells with or without shRNA knockdown of CD90 (+/−monocyte-coculture) were detected by western blot. (g) Western blots of phospho- and total EphA4, EphA2, PLCγ1, PKCδ and Src in the CSC-like cells cocultured for 30 min (30’) with monocytes. The monocytes were pre-treated with either PBS only or PBS with 1 unit ml−1 PI-PLC. (h) Tumor-associated monocytes and macrophages (TAMs) interact with cancer stem cells (CSCs) via CD90- and EphA4- mediated juxtacrine signaling, which activates a network of cytokine signaling. The cytokines can act in autocrine to maintain the stem-cell state, and in paracrine on the inflammatory cells to perpetuate the TAM-CSC interactions. As critical components of the CSC-niche, TAMs enhance tumor initiation and CSC expansion.
Supplementary information
Supplementary Information
Supplementary Information (PDF 2986 kb)
Supplementary Table 1
Supplementary Information (XLSX 1659 kb)
Supplementary Table 2
Supplementary Information (XLSX 15 kb)
Supplementary Table 3
Supplementary Information (XLSX 10 kb)
Supplementary Table 4
Supplementary Information (XLSX 9 kb)
Supplementary Table 5
Supplementary Information (XLSX 9 kb)
Supplementary Table 6
Supplementary Information (XLSX 12 kb)
Supplementary Table 7
Supplementary Information (XLSX 18 kb)
Rights and permissions
About this article
Cite this article
Lu, H., Clauser, K., Tam, W. et al. A breast cancer stem cell niche supported by juxtacrine signalling from monocytes and macrophages. Nat Cell Biol 16, 1105–1117 (2014). https://doi.org/10.1038/ncb3041
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3041
This article is cited by
-
MAGOH promotes gastric cancer progression via hnRNPA1 expression inhibition-mediated RONΔ160/PI3K/AKT signaling pathway activation
Journal of Experimental & Clinical Cancer Research (2024)
-
Eph receptors and ephrins in cancer progression
Nature Reviews Cancer (2024)
-
Tumor Microenvironment: A Niche for Cancer Stem Cell Immunotherapy
Stem Cell Reviews and Reports (2024)
-
Exploring the dynamic interplay between cancer stem cells and the tumor microenvironment: implications for novel therapeutic strategies
Journal of Translational Medicine (2023)
-
Cancer stem cells and their niche in cancer progression and therapy
Cancer Cell International (2023)