Abstract
Metastasis remains the most common cause of death in most cancers, with limited therapies for combating disseminated disease. While the primary tumour microenvironment is an important regulator of cancer progression, it is less well understood how different tissue environments influence metastasis. We analysed tumour–stroma interactions that modulate organ tropism of brain, bone and lung metastasis in xenograft models. We identified a number of potential modulators of site-specific metastasis, including cathepsin S as a regulator of breast-to-brain metastasis. High cathepsin S expression at the primary site correlated with decreased brain metastasis-free survival in breast cancer patients. Both macrophages and tumour cells produce cathepsin S, and only the combined depletion significantly reduced brain metastasis in vivo. Cathepsin S specifically mediates blood–brain barrier transmigration through proteolytic processing of the junctional adhesion molecule, JAM-B. Pharmacological inhibition of cathepsin S significantly reduced experimental brain metastasis, supporting its consideration as a therapeutic target for this disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Quail, D. F. & Joyce, J. A. Microenvironmental regulation of tumor progression and metastasis. Nat. Med. 19, 1423–1437 (2013).
Nguyen, D. X., Bos, P. D. & Massague, J. Metastasis: from dissemination to organ-specific colonization. Nat. Rev. Cancer 9, 274–284 (2009).
Goss, P. E. & Chambers, A. F. Does tumour dormancy offer a therapeutic target? Nat. Rev. Cancer 10, 871–877 (2010).
Affara, N. I., Andreu, P. & Coussens, L. M. Delineating protease functions during cancer development. Methods Mol. Biol. 539, 1–32 (2009).
Mason, S. D. & Joyce, J. A. Proteolytic networks in cancer. Trends Cell Biol. 21, 228–237 (2011).
Gocheva, V. et al. Distinct roles for cysteine cathepsin genes in multistage tumorigenesis. Genes Dev. 20, 543–556 (2006).
Murphy, G. The ADAMs: signalling scissors in the tumour microenvironment. Nat. Rev. Cancer 8, 929–941 (2008).
Butler, G. S. & Overall, C. M. Proteomic identification of multitasking proteins in unexpected locations complicates drug targeting. Nat. Rev. Drug Discov. 8, 935–948 (2009).
Lopez-Otin, C. & Hunter, T. The regulatory crosstalk between kinases and proteases in cancer. Nat. Rev. Cancer 10, 278–292 (2010).
Kessenbrock, K., Plaks, V. & Werb, Z. Matrix metalloproteinases: regulators of the tumor microenvironment. Cell 141, 52–67 (2010).
Allinen, M. et al. Molecular characterization of the tumor microenvironment in breast cancer. Cancer Cell 6, 17–32 (2004).
Finak, G. et al. Stromal gene expression predicts clinical outcome in breast cancer. Nat. Med. 14, 518–527 (2008).
Erez, N., Truitt, M., Olson, P., Arron, S. T. & Hanahan, D. Cancer-associated fibroblasts are activated in incipient neoplasia to orchestrate tumor-promoting inflammation in an NF-κB-dependent manner. Cancer Cell 17, 135–147 (2010).
Katz, A. M. et al. Astrocyte-specific expression patterns associated with the PDGF-induced glioma microenvironment. PLoS ONE 7, e32453 (2012).
Seaman, S. et al. Genes that distinguish physiological and pathological angiogenesis. Cancer Cell 11, 539–554 (2007).
Ojalvo, L. S., Whittaker, C. A., Condeelis, J. S. & Pollard, J. W. Gene expression analysis of macrophages that facilitate tumor invasion supports a role for Wnt-signaling in mediating their activity in primary mammary tumors. J. Immunol. 184, 702–712 (2010).
Paget, S. The distribution of secondary growths in cancer of the breast. 1889. Cancer Metastasis Rev. 8, 98–101 (1989).
Disibio, G. & French, S. W. Metastatic patterns of cancers: results from a large autopsy study. Arch. Pathol. Lab. Med. 132, 931–939 (2008).
Lee, Y. T. Breast carcinoma: pattern of metastasis at autopsy. J. Surg. Oncol. 23, 175–180 (1983).
Bos, P. D. et al. Genes that mediate breast cancer metastasis to the brain. Nature 459, 1005–1009 (2009).
Kang, Y. et al. A multigenic program mediating breast cancer metastasis to bone. Cancer Cell 3, 537–549 (2003).
Minn, A. J. et al. Genes that mediate breast cancer metastasis to lung. Nature 436, 518–524 (2005).
Schwartz, D. R. et al. Hu/Mu ProtIn oligonucleotide microarray: dual-species array for profiling protease and protease inhibitor gene expression in tumors and their microenvironment. Mol. Cancer Res. 5, 443–454 (2007).
Ponomarev, V. et al. A novel triple-modality reporter gene for whole-body fluorescent, bioluminescent, and nuclear noninvasive imaging. Eur. J. Nucl. Med. Mol. Imaging 31, 740–751 (2004).
Shi, G. P. et al. Cathepsin S required for normal MHC class II peptide loading and germinal center development. Immunity 10, 197–206 (1999).
Lorger, M. & Felding-Habermann, B. Capturing changes in the brain microenvironment during initial steps of breast cancer brain metastasis. Am. J. Pathol. 176, 2958–2971 (2010).
Small, D. M. et al. Cathepsin S from both tumor and tumor-associated cells promote cancer growth and neovascularization. Int. J. Cancer 133, 2102–2112 (2013).
Abbott, N. J., Ronnback, L. & Hansson, E. Astrocyte-endothelial interactions at the blood–brain barrier. Nat. Rev. Neurosci. 7, 41–53 (2006).
Eugenin, E. A. & Berman, J. W. Chemokine-dependent mechanisms of leukocyte trafficking across a model of the blood-brain barrier. Methods 29, 351–361 (2003).
Jia, W., Martin, T. A., Zhang, G. & Jiang, W. G. Junctional adhesion molecules in cerebral endothelial tight junction and brain metastasis. Anticancer Res. 33, 2353–2359 (2013).
Liu, W. Y., Wang, Z. B., Zhang, L. C., Wei, X. & Li, L. Tight junction in blood-brain barrier: an overview of structure, regulation, and regulator substances. CNS Neurosci. Ther. 18, 609–615 (2012).
Gallagher, F. A. et al. Magnetic resonance imaging of pH in vivo using hyperpolarized 13C-labelled bicarbonate. Nature 453, 940–943 (2008).
Kirschke, H., Wiederanders, B., Bromme, D. & Rinne, A. Cathepsin S from bovine spleen. Purification, distribution, intracellular localization and action on proteins. Biochem. J. 264, 467–473 (1989).
Nolan, D. J. et al. Molecular signatures of tissue-specific microvascular endothelial cell heterogeneity in organ maintenance and regeneration. Dev. Cell 26, 204–219 (2013).
Choe, Y. et al. Substrate profiling of cysteine proteases using a combinatorial peptide library identifies functionally unique specificities. J. Biol. Chem. 281, 12824–12832 (2006).
Oliveira, M. et al. Improvement of cathepsin S detection using a designed FRET peptide based on putative natural substrates. Peptides 31, 562–567 (2010).
Biniossek, M. L., Nagler, D. K., Becker-Pauly, C. & Schilling, O. Proteomic identification of protease cleavage sites characterizes prime and non-prime specificity of cysteine cathepsins B, L, and S. J. Proteome Res. 10, 5363–5373 (2011).
Oskarsson, T. et al. Breast cancer cells produce tenascin C as a metastatic niche component to colonize the lungs. Nat. Med. 17, 867–874 (2011).
Chapman, H. A. Endosomal proteases in antigen presentation. Curr. Opin. Immunol. 18, 78–84 (2006).
Gupta, S., Singh, R. K., Dastidar, S. & Ray, A. Cysteine cathepsin S as an immunomodulatory target: present and future trends. Expert Opin. Ther. Targets 12, 291–299 (2008).
Madsen, C. D. & Sahai, E. Cancer dissemination–lessons from leukocytes. Dev. Cell 19, 13–26 (2010).
Steeg, P. S., Camphausen, K. A. & Smith, Q. R. Brain metastases as preventive and therapeutic targets. Nat. Rev. Cancer 11, 352–363 (2011).
Eichler, A. F. et al. The biology of brain metastases-translation to new therapies. Nat. Rev. Clin. Oncol. 8, 344–356 (2011).
Park, E. S. et al. Cross-species hybridization of microarrays for studying tumor transcriptome of brain metastasis. Proc. Natl Acad. Sci. USA 108, 17456–17461 (2011).
Turk, B. Targeting proteases: successes, failures and future prospects. Nat. Rev. Drug Discov. 5, 785–799 (2006).
Palermo, C. & Joyce, J. A. Cysteine cathepsin proteases as pharmacological targets in cancer. Trends Pharmacol. Sci. 29, 22–28 (2008).
Drag, M. & Salvesen, G. S. Emerging principles in protease-based drug discovery. Nat. Rev. Drug Discov. 9, 690–701 (2010).
Fingleton, B. Matrix metalloproteinases as valid clinical targets. Curr. Pharm. Des. 13, 333–346 (2007).
Seidah, N. G. & Prat, A. The biology and therapeutic targeting of the proprotein convertases. Nat. Rev. Drug Discov. 11, 367–383 (2012).
Guy, C. T., Cardiff, R. D. & Muller, W. J. Induction of mammary tumors by expression of polyomavirus middle T oncogene: a transgenic mouse model for metastatic disease. Mol. Cell. Biol. 12, 954–961 (1992).
Shree, T. et al. Macrophages and cathepsin proteases blunt chemotherapeutic response in breast cancer. Genes Dev. 25, 2465–2479 (2011).
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004).
Gautier, L., Cope, L., Bolstad, B. M. & Irizarry, R. A. Affy–analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20, 307–315 (2004).
Smyth, G. K. in Bioinformatics and Computational Biology Solutions Using R and Bioconductor (eds Gentleman, R., Carey, V., Dudoit, S., Irizarry, R. A. & Huber, W.) 397–420 (Springer, 2005).
Therneau, T. A Package for Survival Analysis in S. R package version 2.37-7, (2014); http://cran.r-project.org/package=survival
Gocheva, V. et al. IL-4 induces cathepsin protease activity in tumor-associated macrophages to promote cancer growth and invasion. Genes Dev. 24, 241–255 (2010).
Acknowledgements
We thank K. Simpson and X. Chen for excellent technical support, and members of the J.A.J. laboratory for insightful discussion. We thank L. Akkari, O. Olson and D. Yan for reading the manuscript. We are grateful to P. Bos for advice on the BBB assays and experimental brain metastasis model. We thank the MSKCC Core Facilities of Genomics, Molecular Cytology, Small Animal Imaging and Monoclonal Antibody Production for technical assistance. We thank H. Chapman (UCSF) for providing cathepsin S KO mice. This research was supported by the following: US National Cancer Institute program grants of the Integrative Cancer Biology Program (CA148967; J.A.J., C.S.L.) and Tumour Microenvironment Network (CA126518; J.A.J., J.M.), the Health Research Science Board of New York, and the Alan and Sandra Gerry Metastasis Research Initiative (J.A.J.), Deutsche Forschungsgemeinschaft (SE2234/1-1; L.S.), US National Cancer Institute F31 fellowship CA167863 and Gerstner Sloan Kettering graduate program (R.L.B.), US National Cancer Institute F32 fellowship CA130329 (S.D.M.), and Canadian Institutes of Health Research (D.F.Q.).
Author information
Authors and Affiliations
Contributions
L.S., S.D.M. and J.A.J. designed experiments. L.S., S.D.M., D.F.Q. and B.T.E. performed experiments and analysed data. R.L.B., F.R. and C.S.L. performed computational analyses. E.B., P.K.B., W.C.H., L.J.H. and J.M. provided patient samples or reagents. J.A.J., L.S. and R.L.B. wrote the manuscript. All authors edited or commented on the manuscript. J.A.J. conceived and supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Characterization of the stromal cell types in early- and late-stage brain, bone and lung metastasis.
(a–c) Quantification and representative images of in vivo BLI intensity are shown for (a) brain metastases (n = 7 and 9 mice), (b) bone metastases (n = 6 and 7 mice), and (c) lung metastases (n = 4 mice per group) for early and late stages. Circles represent individual mice and horizontal lines represent the mean ± s.e.m. (d–f) Representative images of control tissue and early- and late-stage brain, bone, and lung metastases stained for GFP (tumour cells) and the endothelial marker CD34 or the macrophage markers CD68 (lung and bone) or Iba1 (brain). DAPI staining was used to visualize cell nuclei. Brain metastasis sections were also stained with the astrocyte-specific marker GFAP. Images are representative of three independent samples per stage. Scale bar indicates 50 μm.
Supplementary Figure 2 Tissue- and stage-specific gene expression changes in tumour and stroma.
(a) Venn diagram of the tumour-derived genes that are significantly different between early and late metastases across different metastatic sites (Fig. 1c). Of the 308 genes significantly different between early and late stages in either brain, bone or lung metastases, 176 genes change by stage in all three sites. (b) Venn diagram depicting the overlap of the 75 stroma-derived genes that are significantly different between early- and late-stage metastases across brain, bone and lung (Fig. 1d). Unlike the tumour-derived genes depicted in a, there were no stromal-derived genes that were significantly different between early- and late-stage metastases in all three tissues investigated. (c) Heatmap depicting tissue-specific gene expression in the brain, bone and lung stroma. No tissue-specific genes were identified for tumour-derived genes. Proteases are denoted in purple, endogenous inhibitors in red, and their interacting partners in black. (d) qPCR confirmed the tissue-enriched expression pattern of Htra1 for brain, Mmp13 for bone, and Mmp11 for lung in control tissue, early- and late-stage metastases. n = 6, 4, 4 samples for control tissues, n = 5, 6, 4 samples for early stages, and n = 6, 6, 4 samples for late stages in brain, bone and lung respectively. Graphs represent mean + s.e.m. P values were obtained using two-tailed unpaired t-test: ∗P < 0.05,∗∗P < 0.01, and ∗∗∗P < 0.001.
Supplementary Figure 3 Independent validation of differentially expressed genes in experimental brain, bone and lung metastases.
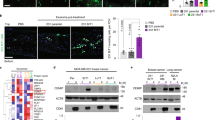
qPCR was used to confirm stage-differential expression changes, and to determine if the expression changes in stromal cells in brain or bone metastasis are a general response to tumour cell colonization of the respective organ, or if the expression changes depend on the tumour cell variant. Hatched bars represent ‘mismatched’ samples (Br-M in bone and Bo-M in brain). Filled bars indicate the ‘matched’ samples. (a,b) qPCR for Ctsh, Cst7, Pcsk1n, Serpini1 (brain) in a and Adamts4, Adamts12, Casp2, Ctse (bone) in b confirmed stage-differential expression changes in a tissue-dependent manner. ‘Mismatched’ samples did not show significant changes. (c,d) qPCR for ADAM21, CTSZ, FAU, TIMP2 (brain) in c and ADAM17, CASP3, DPP8 (bone) in d confirmed stage-differential expression changes in a tissue-dependent manner. ‘Mismatched’ samples (Br-M to bone and Bo-M to brain) revealed that tumour cells underwent similar significant expression changes between early and late stages as identified for matched samples in the respective tissue. (e) qPCR for SERPINE2 confirmed stage-differential expression changes in Lu-M tumour cells in lung metastasis, while those changes were not present in Br-M and Bo-M cells in brain or bone metastasis, respectively. (f) qPCR for Serpina3n and Timp1 confirmed stage-differential expression changes (control versus late) in the stroma of lungs from xenografted animals as well as lungs from the immunocompetent, syngeneic MMTV-PyMT breast cancer model. (g) Immunofluorescence staining for Serpina3n and Timp1 in lungs of MMTV-PyMT breast cancer model (upper panels) and Ctss (red) in co-staining with GFP (tumour cells; green) or Iba1 (macrophages; white) in the syngeneic PyMT-BrM model (lower panels). Images are representative of three independent samples per stage. Scale bar indicates 50 μm. For qPCR validation: n = 11, 15, 15, 13, 7, 15 samples in a, n = 11, 15, 15, 13, 7, 5 samples in b, n = 8, 15, 7, 9, 5, 15 samples in c, n = 8, 15, 7, 9, 5, 5 samples in d, n = 8, 6, 15, 12, 5, 3 samples in e. n = 5 and 9 samples for control lung and late-stage metastases (xenograft model), and n = 6 and 9 for control lung and late-stage metastases (syngeneic model). mRNA expression is depicted relative to early-stage metastases in a–e and relative to control tissue in f. All assays were performed in triplicate and gene expression was normalized to Ubc for all stromal genes, B2M for tumour cell-derived genes. P values were obtained using two-tailed unpaired t-test: NS = not significant, ∗P < 0.05,∗∗P < 0.01, and ∗∗∗P < 0.001.
Supplementary Figure 4 Identification of genes associated with metastasis-free survival (MFS) and differentially expressed between the tumour and stroma.
(a–b) As shown in Fig. 3a for brain metastasis, cross-species scatter plots depict expression changes in the tumour and stromal gene space during the transition from early- to late-stage metastasis for (a) bone and (b) lung metastasis. Differentially expressed genes in the stroma (mouse) or in the tumour (human) gene space are shown in pink or black respectively. Genes that are differentially expressed in both the stroma and the tumour gene space are shown in purple. (c–e) Differentially expressed genes shown in Fig. 1c were analysed for association with MFS for either (c) brain, (d) bone, or (e) lung metastasis, depending on the tissue in which the gene was differentially expressed. Scaled gene expression values were used in a Cox proportional hazards model as described in the methods. Hazard ratios (HR) and 95% confidence intervals (CI) are shown for each organ site. HR < 1.0 is associated with better patient prognosis, whereas HR > 1.0 is associated with poor patient prognosis. For CTSS, the HR for brain MFS was 1.4 (95% CI: 1.05–1.89; P = 0.0209). (f) Genes depicted in c–e are shown in the Venn diagram, where few genes were found to be significantly associated with MFS in multiple tissues. A single gene, SLPI, was significantly associated with MFS in all three tissues, and high expression of levels of SLPI correlated with poor patient prognosis. Hazard ratio significance was determined using Wald’s test with a nominal P value cutoff of <0.05.
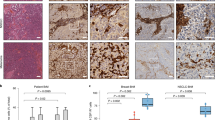
Supplementary Figure 5 Tumour cells and macrophages are the major constituent cell types of patient brain metastases and express high levels of CTSS.
(a,b) Representative images of (a) primary breast cancer and (b) brain metastases patient samples stained for CTSS (red) in combination with the macrophage marker CD68 (white) or a pan-cytokeratin (CK) antibody to visualize tumour cells (green). DAPI staining was used to visualize cell nuclei (blue). All patient samples listed in Supplementary Table 3 were stained and analysed and representative images are shown. The patient samples shown here represent different subtypes of breast cancer based on ER/PR/Her2 status. Scale bar indicates 100 μm in the upper panel rows and 20 μm in the lower panel rows. (c) Quantification of proportions of CK+ tumour cells and CD68+ macrophages in brain metastases samples (n = 7 samples from patients for which there was no matched primary breast tumour tissue available). (d) Combined quantification of proportions of CK+ tumour cells, CD68+ macrophages, and the remaining CK−CD68− cell population in primary breast cancer (BC) (n = 6 samples) and brain metastasis samples (BrM) (n = 13; either matched to the primary, or unmatched samples). (e) Quantification of the CTSS index as a measure of relative CTSS protein levels in tumour cells and macrophages (n = 7 samples). Data in c and e are presented as box plots. Boxes: values between the 25th and 75th percentile; whiskers: minimum and maximum values; horizontal line: median. Data in d are presented as stacked bars + s.e.m.
Supplementary Figure 6 Cathepsin S deficiency does not affect viability of Br-M cells or vessel formation in mice.
(a) Quantification of CTSS mRNA expression in Br-M Ctrl and Br-M CTSS KD tumour cell lines (n = 7 samples per group). Expression is depicted relative to Br-M Ctrl cells. All assays were run in triplicate and gene expression was normalized to B2M. (b) Western blot analysis of CTSS expression levels in cell lysates and conditioned media (CM) from Br-M Ctrl and Br-M CTSS KD cells. Western blots shows representative results from 3 independent experiments. qPCR and western blotting revealed a 90% knockdown efficiency for CTSS at both the mRNA and protein levels. (c) Representative images of CTSS immunofluorescence staining of Br-M Ctrl and Br-M CTSS KD cell lines. DAPI staining was used to visualize cell nuclei (blue). Scale bar indicates 20 μm. (n = 3 replicates). (d) CTSS KD does not affect cell viability in culture as determined by MTT assays (n = 4 independent experiments). (e) Quantification of tumour cell apoptosis (percentage of TUNEL+GFP+ cells) on day 35 post-tumour cell inoculation. Ctrl; Ctss WT (n = 3 mice), CTSS KD; Ctss WT (n = 4 mice), Ctrl; Ctss KO (n = 6 mice), and CTSS KD; Ctss KO (n = 3 mice). (f) Brain metastases size and vessel density were defined as the area covered by GFP (tumour cells) or CD34 (endothelial cells) respectively, and the GFP-covered area was quantified relative to CD34-covered area using Metamorph image analysis. Ctrl; Ctss WT (n = 4 mice), CTSS KD; Ctss WT (n = 4 mice), Ctrl; Ctss KO (n = 6 mice), and CTSS KD; Ctss KO (n = 4 mice) (g) Quantification of vessel density in the brain, defined as the ratio of Texas red lectin+ area to total DAPI area (n = 4 for WT mice, n = 6 for Ctss KO mice), and (h) assessment of vessel permeability by intraveneous injection of Evan’s blue dye (n = 8 mice for each group). Each western blot shows the representative result of three independent experiments. Uncropped images of blots are shown in Supplementary Fig. 9.
Supplementary Figure 7 Cathepsin S deficiency alters the kinetics of brain metastasis formation, though not bone metastasis, but does not affect tumour growth in a regression trial.
(a) Quantification of BLI intensity in vivo at the indicated time points after tumour cell inoculation. Ctrl; Ctss WT (n = 16 mice), CTSS KD; Ctss WT (n = 15 mice), Ctrl; Ctss KO (n = 11 mice), and CTSS KD; Ctss KO (n = 10 mice). (b) Schematic of the regression trial experimental design. (c) Quantification of the BLI intensity from d0-d35 after Br-M tumour cell inoculation (n = 7 mice per group) in the regression trial setting. (d) Quantification of the BLI intensity in the bone and spine region of vehicle-treated and VBY-999-treated animals from the prevention trial, at d35 following Br-M tumour cell inoculation (n = 23 mice for vehicle group and n = 21 mice for VBY-999 group). (e) Quantification of the BLI intensity in the bone and spine region at d35 following Br-M tumour cell inoculation, in the four experimental groups. Ctrl; Ctss WT (n = 10 mice), CTSS KD; Ctss WT (n = 8 mice), Ctrl; Ctss KO (n = 10 mice), and CTSS KD; Ctss KO (n = 11 mice). (f) Quantification of the BLI intensity in the head region (which may arise from skull and/or brain lesions) of vehicle-treated and VBY-999-treated animals from the prevention trial, at d35 following Bo-M tumour cell inoculation (n = 12 mice for vehicle group and n = 13 mice for VBY-999 group). Graphs represent mean ± s.e.m. or circles represent individual mice and horizontal lines represent the mean ± s.e.m. P values were obtained using two-tailed unpaired t-test: NS = not significant, ∗P < 0.05.
Supplementary Figure 8 Cathepsin S deficiency impairs transmigration in an in vitro BBB assay, and sequence analysis identifies a putative cleavage site for cathepsin S.
(a) Pharmacological inhibition of cathepsin S with increasing concentrations of the cathepsin S-specific inhibitor VBY-999 (0 μM (Vehicle) to 100 μM) did not affect Br-M cell viability as determined by MTT assays (n = 3 independent experiments). (b) Quantification of the number of transmigrated Br-M Ctrl and CTSS KD cells. n = 15 samples per condition with 200 fields of view analysed per sample in three independent experiments. (c) Expression of tight junction and adherens junction proteins was confirmed in an independent data set (GSE47067) of FACS sorted endothelial cells. Only Jam-B and Ocln are significantly enriched in brain endothelial cells compared to either lung or bone endothelial cells. n = 3, 3, 6 samples analysed per organ. (d) Western blot analysis of recombinant JAM-A, -B and-C cleavage by CTSS, in the presence or absence of VBY-999 (10 μM), using an antibody that detects the IgG1 domain in the recombinant JAM fusion proteins. (e) Alignment of the amino acid sequence of JAM-A, -B, and-C. Motifs that are conserved in all 3 family members are highlighted in dark purple, and motifs that are conserved in 2 of the 3 JAM family members are depicted in light purple. The putative cleavage location for cathepsin S is indicated by the red box. (f) Quantification of JAM-B cell based cleavage experiments (n = 3 independent experiments). Graphs represent mean ± s.e.m. in a and b, and box plots in c. Boxes: values between the 25th and 75th percentile; whiskers: minimum and maximum values; horizontal line: median. Circles represent individual samples and horizontal lines represent the mean ± s.e.m. in f. P values were obtained using two-tailed unpaired t-test: NS = not significant, ∗P < 0.05,∗∗P < 0.01, and ∗∗∗P < 0.001. Each western blot shows the representative result of three independent experiments. Uncropped images of blots are shown in Supplementary Fig. 9.
Supplementary Figure 9 Uncropped western blots.
Uncropped images of immunoblots displayed in the main and supplementary figures. Dashed boxes indicate regions that were cropped.
Supplementary information
Supplementary Information
Supplementary Information (PDF 2363 kb)
Supplementary Table 1
Supplementary Information (XLSX 87 kb)
Supplementary Table 2
Supplementary Information (XLSX 81 kb)
Supplementary Table 3
Supplementary Information (XLSX 48 kb)
Supplementary Table 4
Supplementary Information (XLSX 50 kb)
Supplementary Table 5
Supplementary Information (XLSX 46 kb)
Supplementary Table 6
Supplementary Information (XLSX 44 kb)
Rights and permissions
About this article
Cite this article
Sevenich, L., Bowman, R., Mason, S. et al. Analysis of tumour- and stroma-supplied proteolytic networks reveals a brain-metastasis-promoting role for cathepsin S. Nat Cell Biol 16, 876–888 (2014). https://doi.org/10.1038/ncb3011
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3011
This article is cited by
-
A rare non-gadolinium enhancing sarcoma brain metastasis with microenvironment dominated by tumor-associated macrophages
Acta Neuropathologica Communications (2024)
-
Water channel protein AQP1 in cytoplasm is a critical factor in breast cancer local invasion
Journal of Experimental & Clinical Cancer Research (2023)
-
Liver metastasis from colorectal cancer: pathogenetic development, immune landscape of the tumour microenvironment and therapeutic approaches
Journal of Experimental & Clinical Cancer Research (2023)
-
The role and regulation of Maf proteins in cancer
Biomarker Research (2023)
-
3D bioprinted multilayered cerebrovascular conduits to study cancer extravasation mechanism related with vascular geometry
Nature Communications (2023)