Abstract
With the prevalence of end-stage renal disease rising 8% per annum globally1, there is an urgent need for renal regenerative strategies. The kidney is a mesodermal organ that differentiates from the intermediate mesoderm (IM) through the formation of a ureteric bud (UB) and the interaction between this bud and the adjacent IM-derived metanephric mesenchyme2 (MM). The nephrons arise from a nephron progenitor population derived from the MM (ref. 3). The IM itself is derived from the posterior primitive streak4. Although the developmental origin of the kidney is well understood2, nephron formation in the human kidney is completed before birth5. Hence, there is no postnatal stem cell able to replace lost nephrons. In this study, we have successfully directed the differentiation of human embryonic stem cells (hESCs) through posterior primitive streak and IM under fully chemically defined monolayer culture conditions using growth factors used during normal embryogenesis. This differentiation protocol results in the synchronous induction of UB and MM that forms a self-organizing structure, including nephron formation, in vitro. Such hESC-derived components show broad renal potential ex vivo, illustrating the potential for pluripotent-stem-cell-based renal regeneration.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Little, M. H. & McMahon, A. P. Mammalian kidney development: principles, progress, and projections. Cold Spring Harb. Perspect. Biol. 4 (5) (2012).
Tam, P. P. & Loebel, D. A. Gene function in mouse embryogenesis: get set for gastrulation. Nat. Rev. Genet. 8, 368–381 (2007).
Kobayashi, A. et al. Six2 defines and regulates a multipotent self-renewing nephron progenitor population throughout mammalian kidney development. Cell Stem Cell 3, 169–181 (2008).
Rumballe, B. A. et al. Nephron formation adopts a novel spatial topology at cessation of nephrogenesis. Dev. Biol. 360, 110–122 (2011).
Lusis, M., Li, J., Ineson, J. & Little, M. H. Isolation of clonogenic, long-term self-renewing renal stem cells. Stem Cell Res. 5, 23–39 (2010).
Takasato, M., Maier, B. & Little, M. H. Recreating kidney progenitors from pluripotent cells. Pediatr. Nephrol.http://dx.doi.org/10.1007/s00467-013-2592-7 (2013).
Gadue, P., Huber, T. L., Paddison, P. J. & Keller, G. M. Wnt and TGF-beta signaling are required for the induction of an in vitro model of primitive streak formation using embryonic stem cells. Proc. Natl Acad. Sci. USA 103, 16806–16811 (2006).
Soares, M. L. et al. Functional studies of signaling pathways in peri-implantation development of the mouse embryo by RNAi. BMC Dev. Biol. 5, 28 (2005).
Lu, C. C. & Robertson, E. J. Multiple roles for Nodal in the epiblast of the mouse embryo in the establishment of anterior–posterior patterning. Dev. Biol. 273, 149–159 (2004).
Sumi, T., Tsuneyoshi, N., Nakatsuji, N. & Suemori, H. Defining early lineage specification of human embryonic stem cells by the orchestrated balance of canonical Wnt/beta-catenin, Activin/Nodal and BMP signaling. Development 135, 2969–2979 (2008).
Davis, R. P. et al. Targeting a GFP reporter gene to the MIXL1 locus of human embryonic stem cells identifies human primitive streak-like cells and enables isolation of primitive hematopoietic precursors. Blood 111, 1876–1884 (2008).
Ng, E. S., Davis, R., Stanley, E. G. & Elefanty, A. G. A protocol describing the use of a recombinant protein-based, animal product-free medium (APEL) for human embryonic stem cell differentiation as spin embryoid bodies. Nat. Protoc. 3, 768–776 (2008).
Mae, S. et al. Monitoring and robust induction of nephrogenic intermediate mesoderm from human pluripotent stem cells. Nat. Commun. 4, 1367 (2013).
James, R. G. et al. Odd-skipped related 1 is required for development of the metanephric kidney and regulates formation and differentiation of kidney precursor cells. Development 133, 2995–3004 (2006).
Tsang, T. E. et al. Lim1 activity is required for intermediate mesoderm differentiation in the mouse embryo. Dev. Biol. 223, 77–90 (2000).
Torres, M., Gómez-Pardo, E., Dressler, G. R. & Gruss, P. Pax-2 controls multiple steps of urogenital development. Development 121, 4057–4065 (1995).
Crossley, P. H. & Martin, G. R. The mouse Fgf8 gene encodes a family of polypeptides and is expressed in regions that direct outgrowth and patterning in the developing embryo. Development 121, 439–451 (1995).
Colvin, J. S. et al. Genomic organization and embryonic expression of the mouse fibroblast growth factor 9 gene. Dev. Dynam. 216, 72–88 (1999).
Dudley, A. T., Godin, R. E. & Robertson, E. J. Interaction between FGF and BMP signaling pathways regulates development of metanephric mesenchyme. Genes Dev. 13, 1601–1613 (1999).
Barak, H. et al. FGF9 and FGF20 maintain the stemness of nephron progenitors in mice and man. Dev. Cell 22, 1191–1207 (2012).
James, R. G. & Schultheiss, T. M. Bmp signaling promotes intermediate mesoderm gene expression in a dose-dependent, cell-autonomous and translation-dependent manner. Dev. Biol. 288, 113–125 (2005).
Rosselot, C. et al. Non-cell-autonomous retinoid signaling is crucial for renal development. Development 137, 283–292 (2010).
Kim, D. & Dressler, G. R. Nephrogenic factors promote differentiation of mouse embryonic stem cells into renal epithelia. J Am. Soc. Nephrol. 16, 3527–3534 (2005).
Mugford, J. W. et al. Hoxd11 specifies a program of metanephric kidney development within the intermediate mesoderm of the mouse embryo. Dev. Biol. 319, 396–405 (2008).
Pachnis, V., Mankoo, B. & Costantini, F. Expression of the c-ret proto-oncogene during mouse embryogenesis. Development 119, 1005–1017 (1993).
Srinivas, S. et al. Expression of green fluorescent protein in the ureteric bud of transgenic mice: a new tool for the analysis of ureteric bud morphogenesis. Dev. Genet. 24, 241–251 (1999).
Mendelsohn, C. et al. Stromal cells mediate retinoid-dependent functions essential for renal development. Development 126, 1139–1148 (1999).
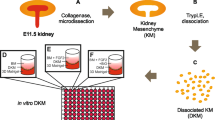
Davies, J. A. et al. Dissociation of embryonic kidney followed by re-aggregation as a method for chimeric analysis. Methods Mol. Biol. 886, 135–146 (2012).
Hendry, C. E. & Vanslambrouck, J. M. et al. Direct transcriptional reprogramming of adult cells to embryonic nephron progenitors. J. Am. Soc. Nephrol. 24, 1424–1434 (2013).
Trinkaus, J. P. & Groves, P. W. Differentiation in culture of mixed aggregates of dissociated tissue cells. Proc. Natl Acad. Sci. USA 41, 787–795 (1955).
Suga, H. et al. Self-formation of functional adenohypophysis in three-dimensional culture. Nature 480, 57–62 (2011).
Eiraku, M. et al. Self-organizing optic-cup morphogenesis in three-dimensional culture. Nature 472, 51–56 (2011).
Spence, J. R. et al. Directed differentiation of human pluripotent stem cells into intestinal tissue in vitro. Nature 470, 105–108 (2011).
Narayanan, K. et al. Human embryonic stem cells differentiate into functional renal proximal tubular-like cells. Kidney Int. 83, 593–603 (2013).
Song, B. et al. The directed differentiation of human iPS cells into kidney podocytes. PLoS ONE 7, e46453 (2012).
Brenner-Anantharam, A. et al. Tailbud-derived mesenchyme promotes urinary tract segmentation via BMP4 signaling. Development 134, 1967–1975 (2007).
Guillaume, R., Bressan, M. & Herzlinger, D. Paraxial mesoderm contributes stromal cells to the developing kidney. Dev. Biol. 329, 169–175 (2009).
Uchiyama, Y. et al. Kif26b, a kinesin family gene, regulates adhesion of the embryonic kidney mesenchyme. Proc. Natl Acad. Sci. USA 107, 9240–9245 (2010).
Linton, J. M., Martin, G. R. & Reichardt, L. F. The ECM protein nephronectin promotes kidney development via integrin alpha8beta1-mediated stimulation of Gdnf expression. Development 134, 2501–2509 (2007).
Acknowledgements
We are grateful to E. Wolvetang for providing an iPSC line and D. Titmarsh, A. Hidalgo-Gonzalez and J. Cooper-White for supportive comments. This work was supported by the Queensland State Government through a National/International Research Alliance Project, the Australian Research Council as part of the Stem Cells Australia Strategic Research Initiative (SRI110001002) and the National Health and Medical Research Council of Australia (APP1041277). M.H.L. is a Senior Principal Research Fellow of the NHMRC.
Author information
Authors and Affiliations
Contributions
M.T. and M.H.L. conceived and planned the project and wrote the manuscript. M.B. and P.X.E. provided technical assistance with hESC culture, histology, microscopy and differentiation protocols. E.G.S. and A.G.E. provided targeted hESC lines and advised on design and execution. J.M.V. provided technical advice, support and analysis for ex vivo recombination assays.
Corresponding author
Ethics declarations
Competing interests
M.H.L. consults for Organovo Inc.
Integrated supplementary information
Supplementary Figure 1 Posterior primitive streak induction.
a, Time course quantitative RT-PCR for pluripotent markers, OCT4 and NANOG after induction with BMP4/ActivinA (30/10 ng/ml), showing a reduction in pluripotent gene expression with time. Error bars are s.d. (n = 3 experiments). The source data for the graph are provided in Supplementary Table 1. b, IF for markers of ES cells, NANOG and ECAD, before (hESCs) and after (day 2) primitive streak induction using CHIR99021. (scale = 100 μm) c, IF for markers of posterior primitive streak, T and MIXL1 (GFP), after the primitive streak induction (day 2) using CHIR99021. MIXL1 was detected as GFP expression driven by the MIXL1 endogenous promoter. (scale = 100 μm) d, Levels of spontaneous OSR1 expression induced across time after culture if 3 different ratios of BMP4 and Activin A (ng/mL). hESCs were formed embryoid bodies with 3 different ratios of BMP4 and Activin A for 3 days then spontaneously differentiated under no growth factor condition until day 14. This demonstrates improved OSR1 expression in cells induced with high BMP4 and low Activin A (30/10). OSR1 marks IM and LPM.
Supplementary Figure 2 Influence of FGF signaling on induction of IM proteins.
a, IF for PAX2 protein on hESC cultures at day 6 treated with BMP4/Activin A to day 2 and FGF2 (200 ng/ml), FGF8 (200 ng/ml), FGF9 (200ng/ml) or no growth factors (no GFs) from day 2 to 6 in the presence or absence of the FGF signaling inhibitor, PD173074. (scale = 200 μm) b, Quantitative RT-PCR to examine the relative expression level of PAX2, LHX1 and OSR1 at day 6 of the same protocol as IF (a). Shaded bars show the effect of addition of the FGF inhibitor, PD173074. Error bars are s.d. (n = 3 experiments). The source data for graphs are provided in Supplementary Table 1. c, IF for the IM marker PAX2 and the marker of both LPM and IM, OSR1, on hESC cultures at day 6 treated with BMP4/Activin A (+FGF9 (B/A)) or 8 μM CHIR99021 (+FGF9 (CHIR)) to day 2 followed by 200 ng/mL FGF9 or no growth factors (no GFs) from day 2 to 6. Secondary antibody only control was used as a negative control (2° Ab only) (scale = 100 μm). The source data for graphs are provided in Supplementary Table 1. d, A table showing the percent of PAX2− and PAX2+ cells in total (total) or together with LHX1− and LHX1+ cells on hESC cultures at day 6 treated with 8 μM CHIR99021 to 2 days followed by 200 ng/mL FGF9 from day 2 to 6. Errors are s.d. (n = 5 fields in total from 3 experiments).
Supplementary Figure 3 The effect of BMP signaling on lateral-medial patterning of early mesoderm.
a, IFfor DAPI (blue) and PAX2 (red) at day 6 in the presence of 200ng/mL FGF9 with or without BMP4 (5 or 50 ng/mL) or the BMP antagonist NOG (25 or 250ng/mL) from day 2 to day 6. (scale = 200 μm) b, qRT-PCR to investigate the effect of this BMP/NOG gradient on the expression of PM (PARAXIS and TBX6) and LPM (FOXF1 and OSR1)markers at day 6. Error bars are s.d. (n = 3 experiments). The source data for graphs are provided in Supplementary Table 1.
Supplementary Figure 4 Schematic illustrating the anticipated gene expression of distinct progenitor and derivative cell populations during early kidney development.
PS, primitive streak; IM, intermediate mesoderm; MM, metanephric mesenchyme; NP, nephron progenitor / nephrogenic mesenchyme; RV, renal vesicle; DT, distal convoluted tubule; PT, proximal convoluted tubule; Pod, podocyte; ND, nephric duct; UB, ureteric bud/ureteric epithelium; CD, collecting duct; MET, mesenchymal to epithelial transition. All genes are indicated in italics. Shaded boxes indicate the timing and duration of expression for adjacent labeled genes. Specific genes marking DT, PT and Pod are indicated next to each cell type. The reciprocal induction of differentiation known to occur between the UB and NP is supported by the expression of FGF9 (nephrogenic mesenchyme survival) and Wnt9b (MET) and from the UB and GDNF (ureteric branching) by the NP.
Supplementary Figure 5 The positive effect of RA on ureteric epithelium formation.
a,EdU incorporation assay at day 12 of differentiation. 30 min exposure by EdU revealed that not only PAX2+ pre-epithelium structures but also PAX2 negative cells are proliferating. White arrowheads indicate EdU incorporation in PAX2+ cell. (scale = 100 μm) b, IM cells at day 6 after primitive streak induction using BMP4/Activin A were cultured for 11 days with FGF9 together with different RA concentrations. IF for UE markers, PAX2+ECAD+, showed UE structures were induced in a RA dose-dependent manner. (scale = 200 μm) c, RT-PCR at day 22 of differentiation using BMP4:Activin A/FGF9/FGF9:BMP7:RA protocol revealed the expression of genes indicative of differentiation into mature renal cell types, including SYNPO, NPHS1and WT1 for podocyte; AQP2 and SCNNB1 for distal tubule or collecting duct and AQP1 and SLC3A1 for proximal tubule. NC, negative control with no DNA template. g, IF of day 22 differentiation using BMP4/Activin A showing co-expression of two key podocyte markers; the slit-diaphragm protein SYNPO (green) and nuclear WT1 (red). Nuclei are also stained with DAPI (blue). (scale = 50 μm) The source data for graphs are provided in Supplementary Table 1.
Supplementary Figure 6 Differentiation of H9 hES cell line and iPS cell line towards renal lineages.
a, b, Immunofluorescence for DAPI (blue), PAX2 (red) or SIX2 (red) at Day 6 and Day 14 of differentiation on H9 hESC (a) and CRL2429 C11 iPS cells (b). (scale = 200 μm).
Supplementary Figure 7 The effect of 3D culture environment on self-organisation events.
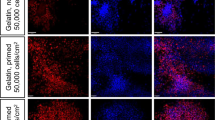
a, Schematic of the replating assay. IM cells at day 6 were harvested and re-plated at high density or low density. Then cells were cultured for 12 days (6 days with 200 ng/mL FGF9 then another 6 days without growth factors). Cells plated at high density formed a uniform layer of cells while those plated at low density formed domed colonies. b, Induced IM cells at day 6 were re-plated to form monolayer or domed colonies at day 18. Cells were stained with ECAD for UE and WT1 for MM. More advanced structures are seen within domed colonies possibly due to the proximity of reciprocally inductive cell populations. (scale = 100 μm).
Supplementary information
Supplementary Information
Supplementary Information (PDF 1827 kb)
Supplementary Table 1
Supplementary Information (XLSX 58 kb)
Supplementary Table 2
Supplementary Information (XLSX 46 kb)
Rights and permissions
About this article
Cite this article
Takasato, M., Er, P., Becroft, M. et al. Directing human embryonic stem cell differentiation towards a renal lineage generates a self-organizing kidney. Nat Cell Biol 16, 118–126 (2014). https://doi.org/10.1038/ncb2894
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2894
This article is cited by
-
Innovative explorations: unveiling the potential of organoids for investigating environmental pollutant exposure
Environmental Science and Pollution Research (2024)
-
Review on kidney diseases: types, treatment and potential of stem cell therapy
Renal Replacement Therapy (2023)
-
Liver organoid culture methods
Cell & Bioscience (2023)
-
Kidney tissue regeneration using bioactive scaffolds incorporated with differentiating extracellular vesicles and intermediate mesoderm cells
Biomaterials Research (2023)
-
Replacing renal function using bioengineered tissues
Nature Reviews Bioengineering (2023)