Abstract
The size of a typical eukaryotic cell is of the order of ∼10 μm. However, some cell types grow to very large sizes, including oocytes (immature eggs) of organisms from humans to starfish. For example, oocytes of the frog Xenopus laevis grow to a diameter ≥1 mm. They have a correspondingly large nucleus (germinal vesicle) of ∼450 μm in diameter, which is similar to smaller somatic nuclei, but contains a significantly higher concentration of actin. The form and structure of this nuclear actin remain controversial, and its potential mechanical role within these large nuclei is unknown. Here, we use a microrheology and quantitative imaging approach to show that germinal vesicles contain an elastic F-actin scaffold that mechanically stabilizes these large nuclei against gravitational forces, which are usually considered negligible within cells. We find that on actin disruption, ribonucleoprotein droplets, including nucleoli and histone locus bodies, undergo gravitational sedimentation and fusion. We develop a model that reveals how gravity becomes an increasingly potent force as cells and their nuclei grow larger than ∼10 μm, explaining the requirement for a stabilizing nuclear F-actin scaffold in large Xenopus oocytes. All life forms are subject to gravity, and our results may have broad implications for cell growth and size control.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Dahl, K. N., Ribeiro, A. J. & Lammerding, J. Nuclear shape, mechanics, and mechanotransduction. Circ. Res. 102, 1307–1318 (2008).
Lammerding, J., Dahl, K. N., Discher, D. E. & Kamm, R. D. in Methods in Cell Biology Vol. 83 (eds Yu-Li, W. & Discher Dennis, E.) 269–294 (Academic, 2007).
Ye, J., Zhao, J., Hoffmann-Rohrer, U. & Grummt, I. Nuclear myosin I acts in concert with polymeric actin to drive RNA polymerase I transcription. Gene Dev. 22, 322–330 (2008).
Hofmann, W. A. & de Lanerolle, P. Nuclear actin: to polymerize or not to polymerize. J. Cell Biol. 172, 495–496 (2006).
Rando, O. J., Zhao, K. & Crabtree, G. R. Searching for a function for nuclear actin. Trends Cell Biol. 10, 92–97 (2000).
Belin, B. J., Cimini, B. A., Blackburn, E. H. & Mullins, R. D. Visualization of actin filaments and monomers in somatic cell nuclei. Mol. Biol. Cell 24, 982–994 (2013).
Stuven, T., Hartmann, E. & Gorlich, D. Exportin 6: a novel nuclear export receptor that is specific for profilin.actin complexes. EMBO J. 22, 5928–5940 (2003).
Bohnsack, M. T., Stuven, T., Kuhn, C., Cordes, V. C. & Gorlich, D. A selective block of nuclear actin export stabilizes the giant nuclei of Xenopus oocytes. Nat. Cell Biol. 8, 257–263 (2006).
Clark, T. G. & Merriam, R. W. Diffusible and bound actin in nuclei of Xenopus laevis oocytes. Cell 12, 883–891 (1977).
Hofmann, W. et al. Cofactor requirements for nuclear export of rev response element (Rre) and constitutive transport element (Cte) containing retroviral rnas: an unexpected role for actin. J. Cell Biol. 152, 895–910 (2001).
Gall, J. G. Exporting actin. Nat. Cell Biol. 8, 205–207 (2006).
Brangwynne, C. P., Koenderink, G. H., MacKintosh, F. C. & Weitz, D. A. Cytoplasmic diffusion: molecular motors mix it up. J. Cell Biol. 183, 583–587 (2008).
Wong, I. Y. et al. Anomalous diffusion probes microstructure dynamics of entangled F-actin networks. Phys. Rev. Lett. 92, 178101 (2004).
Spector, I., Shochet, N. R., Kashman, Y. & Groweiss, A. Latrunculins: novel marine toxins that disrupt microfilament organization in cultured cells. Science 219, 493–495 (1983).
Riedl, J. et al. Lifeact: a versatile marker to visualize F-actin. Nat. Methods 5, 605–607 (2008).
Baarlink, C., Wang, H. & Grosse, R. Nuclear actin network assembly by formins regulates the SRF coactivator MAL. Science 340, 864–867 (2013).
Aebi, U., Cohn, J., Buhle, L. & Gerace, L. The nuclear lamina is a meshwork of intermediate-type filaments. Nature 323, 560–564 (1986).
Dahl, K. N., Kahn, S. M., Wilson, K. L. & Discher, D. E. The nuclear envelope lamina network has elasticity and a compressibility limit suggestive of a molecular shock absorber. J. Cell Sci. 117, 4779–4786 (2004).
Jockusch, B. M., Schoenenberger, C-A., Stetefeld, J. r. & Aebi, U. Tracking down the different forms of nuclear actin. Trends Cell Biol. 16, 391–396 (2006).
Nizami, Z., Deryusheva, S. & Gall, J. G. The cajal body and histone locus body. Csh Perspect Biol. 2, a000653 (2010).
Wu, Z. & Gall, J. G. ‘Micronucleoli’ in the Xenopus germinal vesicle. Chromosoma 105, 438–443 (1997).
Maslova, A. & Krasikova, A. Nuclear actin depolymerization in transcriptionally active avian and amphibian oocytes leads to collapse of intranuclear structures. Nucleus 3, 300–311 (2012).
Brangwynne, C. P., Mitchison, T. J. & Hyman, A. A. Active liquid-like behavior of nucleoli determines their size and shape in Xenopus laevis oocytes. Proc. Natl Acad. Sci. USA 108, 4334–4339 (2011).
Brangwynne, C. P. et al. Germline P granules are liquid droplets that localize by controlled dissolution/condensation. Science 324, 1729–1732 (2009).
Weber, S. C. & Brangwynne, C. P. Getting RNA and protein in phase. Cell 149, 1188–1191 (2012).
Berg, H. C. Random Walks in Biology (Princeton Univ. Press, 1983).
Handwerger, K. E., Cordero, J. A. & Gall, J. G. Cajal bodies, nucleoli, and speckles in the Xenopus oocyte nucleus have a low-density, sponge-like structure. Mol. Biol. Cell 16, 202–211 (2005).
Chan, Y. H. M. & Marshall, W. F. How cells know the size of their organelles. Science 337, 1186–1189 (2012).
Dumont, J. N. Oogenesis in Xenopus laevis (Daudin). I. Stages of oocyte development in laboratory maintained animals. J. Morphol. 136, 153–179 (1972).
Schulz, H. N. & Jorgensen, B. B. Big bacteria. Annu. Rev. Microbiol. 55, 105–137 (2001).
Marshall, W. F. et al. What determines cell size? BMC Biol. 10, 101–123 (2012).
Daniels, B. R., Masi, B. C. & Wirtz, D. Probing single-cell micromechanics in vivo: the microrheology of C. elegans developing embryos. Biophys. J. 90, 4712–4719 (2006).
Valentine, M. T. et al. Colloid surface chemistry critically affects multiple particle tracking measurements of biomaterials. Biophys. J. 86, 4004–4014 (2004).
Gall, J. G. & Wu, Z. Examining the contents of isolated Xenopus germinal vesicles. Methods 51, 45–51 (2010).
Kiseleva, E. et al. Actin- and protein-4.1-containing filaments link nuclear pore complexes to subnuclear organelles in Xenopus oocyte nuclei. J. Cell Sci. 117, 2481–2490 (2004).
Crocker, J. C. G. & David, G. Methods of digital video microscopy for colloidal studies. J. Colloid Inter. Sci. 179, 298–310 (1996).
Taylor, J. R. An Introduction to Error Analysis (Univ. Science Books, 1997).
Acknowledgements
We thank T. Mitchison, S. Weber, E. Wieschaus, C. Broedersz and C. Sosa for discussions and suggestions, D. Mullins (UC-San Francisco, USA) for providing the utrophin constructs, D. Görlich (MPI-Biophysical Chemistry, Goettingen, Germany) for the XPO6 protein, J. Gall (Carnegie Institution, USA) for the GFP::coilin construct and G. Koenderink (AMOLF, Netherlands) for fascin. We are grateful to A. Pozniakovsky for help with cloning and D. Wang for help with frog surgeries, oocyte preparation and some experiments. This work was supported by a Searle Scholar Award (C.P.B.), and an NIH New Innovator Award, 1DP2GM105437-01 (C.P.B.).
Author information
Authors and Affiliations
Contributions
C.P.B. and M.F. designed the study, discussed results, and wrote the paper. M.F. performed the experiments and analysed the data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Actin disruption with latrunculin A or Xpo6.
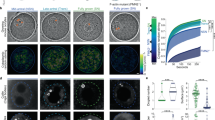
a, Row shows how actin network, visualized with Lifeact::GFP, is disrupted at 15 minute intervals after incubation with latrunculin A. Scale bar = 10 μm. Each image is from a different GV. b, Row shows how nuclear actin structure is disrupted due to actin export at 15-minute intervals after Xpo6 microinjection. Each image is from a different GV. Scale bar = 10 μm. c, d, The probability distribution of F-actin mesh size for GVs under latrunculin A, c, and Xpo-6, d, conditions, for each time point shown above (blue: 15 minutes, yellow: 30 minutes, and red: 45 minutes). Black data points are for the intact Lifeact::GFP structure with no actin disruption (13 z-stacks from 9 GVs). The exponential behavior of the distributions is consistent with a Poisson interval distribution, where the mesh size is ∼1 μm for untreated GVs and ∼10 μm for actin-disrupted GVs after 45 minutes of treatment.
Supplementary Figure 2 Visualization of the nuclear actin network.
a, Image of Lifeact::GFP labeled network within the GV. b, Image of Utrophin-261::GFP labeled network within the GV, showing similar structure as Lifeact::GFP. Scale bar = 10 μm.
Supplementary Figure 3 Expression of Lifeact::GFP does not alter microrheology of the GV.
a, MSD versus lag time of R = 0.1 μm (green) (n = 24 z-positions from 9 GVs, 10,648 particles identified), R = 0.25 μm (blue) (n = 16 z-positions from 8 GVs, 2,053 particles identified), R = 0.5 μm (black) n = 19 z-positions from 6 GVs, 1,867 particles identified), and R = 1 μm (red) (n = 35 z-positions from 14 GVs, 3,011 particles identified) microspheres in native GV (circles) compared with MSD versus lag time of R = 0.1 μm (green) (n = 4 z-positions from 2 GVs, 7,639 particles identified), R = 0.25 μm (blue) (n = 18 z-positions from 6 GVs, 7,250 particles identified), R = 0.5 μm (black) n = 10 z-positions from 4 GVs, 702 particles identified), and R = 1 μm (red) (n = 5 z-positions from 4 GVs, 237 particles identified) microspheres in Lifeact::GFP expressing GVs (triangles). b, Diffusive exponent as a function of microsphere radius, with untreated case in blue and Lifeact::GFP in green. c,MSD at 5 s for each bead size, with untreated case in blue and Lifeact::GFP in green. Error bars = s.e.m.
Supplementary Figure 4 Actin disruption leads to nucleolar sedimentation and fusion.
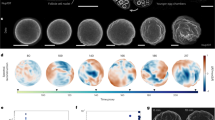
Top images show a maximum intensity projection of a 100-micron thick section of nucleoli (labeled with NPM1::GFP & Fibrillarin::GFP) and bottom images show a 3-D rendering in the X-Z plane. a, Nucleoli are suspended in an untreated GV. For b-d, time-lapse images are from the same GV after Lat-A disruption of actin. e, Large nuclear bodies that form overnight after Lat-A treatment. Scale bar = 50 μm and grid size = 50 μm.
Supplementary Figure 5 Actin disruption after Xpo6 microinjection also leads to formation of a few massive nucleoli at the bottom of the GV.
Top row shows XY maximum intensity projection of an untreated GV (left) and one after overnight incubation after Xpo6 microinjection (right). Bottom row shows the XZ projection of a 100-μm thick section for the corresponding GVs. Scale bar = 50 μm and grid size = 50 μm.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1491 kb)
Diffusion of R = 0.1 μm red microspheres within the Lifeact::GFP actin meshwork.
These beads were the smallest bead size probed and showed diffuse-like behavior. Time reported as min:sec. (MOV 1476 kb)
Diffusion of R = 0.25 μm red microspheres within the Lifeact::GFP actin meshwork.
These intermediate beads showed cage-hopping behaviour, during which the beads diffuse inside a pore and, after some time, jump to a new pore. Time reported as min:sec. (MOV 1386 kb)
Diffusion of R = 1 μm red microspheres within Lifeact::GFP actin meshwork.
These beads were much larger than the average mesh size and exhibited highly-subdiffusive behaviour, leading to their trapped dynamics. Time reported as min:sec. (MOV 336 kb)
Diffusion of NPM1::RFP micronucleoli within Lifeact::GFP actin meshwork.
The diameter of these micronucleoli was approximately equal to or smaller than the pore size, leading to intermittent dynamics and cage-hopping behavior. Time reported as min:sec. (MOV 2505 kb)
Increased mobility of GFP::coilin-labelled HLBs following actin disruption by latrunculin A.
Top panel shows the XY projection of a 100-μm thick section. HLBs are more motile and show more diffusive-like behavior-r than in unperturbed GVs. Bottom panel shows XZ projection of a 100-μm thick section. HLBs sediment to the bottom of the GV on the scale of ∼ 1 h. Time reported as min:sec. (MOV 714 kb)
Sedimentation and fusion of NPM1::GFP- and Fibrillarin::GFP-labelled nucleoli following actin disruption by latrunculin A.
Top panel shows the XY projection of a 100-μm thick section. Nucleoli are more motile and show more diffusive-like behaviour than in unperturbed GVs. Bottom panel shows XZ projection of a 100-μm thick section. Nucleoli rapidly sediment to the bottom of the GV on the scale of ∼ 15 min. Time reported as min:sec. (MOV 1592 kb)
Sedimentation of NPM1::RFP-labelled nucleoli and GFP::coilin-labelled HLBs following actin disruption by latrunculin A.
Top panel shows the XY projection of a 100-μm thick section. Nucleoli and HLBs are more motile and show more diffusive-like behavior than in unperturbed GVs. Bottom panel shows XZ projection of a 100-μm thick section. Nucleoli rapidly sediment to the bottom of the GV on the scale of ∼ 5–10 min, whereas HLBs sediment on a longer time scale of ∼ 30 min. Time reported as min:sec. (MOV 1215 kb)
Actin disruption after 75-min treatment with cytochalasin D.
Actin is labelled in green with Lifeact::GFP and nucleoli are labelled in red with NPM1::RFP. Cyo-D causes the network to become disrupted and results in puncta formation of the actin. The nucleoli become more mobile and can be seen moving in and out of plane as they sediment. Time reported as min:sec. (MOV 1698 kb)
Actin disruption after 30-min treatment with Latrunculin-A.
Actin is labelled in green with Lifeact::GFP. Lat-A disrupts the actin meshwork and results in small, unconnected filaments that are diffusive. Dark bodies are unlabelled nucleoli that can be seen diffusing and moving in and out of plane as they sediment. Time reported as min:sec. (MOV 2764 kb)
Rights and permissions
About this article
Cite this article
Feric, M., Brangwynne, C. A nuclear F-actin scaffold stabilizes ribonucleoprotein droplets against gravity in large cells. Nat Cell Biol 15, 1253–1259 (2013). https://doi.org/10.1038/ncb2830
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2830
This article is cited by
-
Kinetic control of shape deformations and membrane phase separation inside giant vesicles
Nature Chemistry (2024)
-
Superwettable interface towards biodetection in confined space
Nano Research (2024)
-
Effective simulations of interacting active droplets
Scientific Reports (2023)
-
The secret life of the protein VASP
Nature Physics (2023)
-
Liquid–liquid phase separation within fibrillar networks
Nature Communications (2023)