Abstract
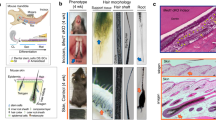
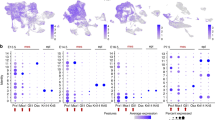
The polycomb group gene Bmi1 is required for maintenance of adult stem cells in many organs1,2. Inactivation of Bmi1 leads to impaired stem cell self-renewal due to deregulated gene expression. One critical target of BMI1 is Ink4a/Arf, which encodes the cell-cycle inhibitors p16Ink4a and p19Arf (ref. 3). However, deletion of Ink4a/Arf only partially rescues Bmi1-null phenotypes4, indicating that other important targets of BMI1 exist. Here, using the continuously growing mouse incisor as a model system, we report that Bmi1 is expressed by incisor stem cells and that deletion of Bmi1 resulted in fewer stem cells, perturbed gene expression and defective enamel production. Transcriptional profiling revealed that Hox expression is normally repressed by BMI1 in the adult, and functional assays demonstrated that BMI1-mediated repression of Hox genes preserves the undifferentiated state of stem cells. As Hox gene upregulation has also been reported in other systems when Bmi1 is inactivated1,2,5,6,7, our findings point to a general mechanism whereby BMI1-mediated repression of Hox genes is required for the maintenance of adult stem cells and for prevention of inappropriate differentiation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Molofsky, A. V. et al. Bmi-1 dependence distinguishes neural stem cell self-renewal from progenitor proliferation. Nature 425, 962–967 (2003).
Park, I. K. et al. Bmi-1 is required for maintenance of adult self-renewing haematopoietic stem cells. Nature 423, 302–305 (2003).
Jacobs, J. J., Kieboom, K., Marino, S., DePinho, R. A. & van Lohuizen, M. The oncogene and Polycomb-group gene bmi-1 regulates cell proliferation and senescence through the ink4a locus. Nature 397, 164–168 (1999).
Molofsky, A. V., He, S., Bydon, M., Morrison, S. J. & Pardal, R. Bmi-1 promotes neural stem cell self-renewal and neural development but not mouse growth and survival by repressing the p16Ink4a and p19Arf senescence pathways. Genes Dev. 19, 1432–1437 (2005).
Zacharek, S. J. et al. Lung stem cell self-renewal relies on BMI1-dependent control of expression at imprinted loci. Cell Stem Cell 9, 272–281 (2011).
Bruggeman, S. W., Hulsman, D. & van Lohuizen, M. Bmi1 deficient neural stem cells have increased integrin dependent adhesion to self-secreted matrix. Biochim. Biophys. Acta 1790, 351–360 (2009).
Fasano, C. A. et al. shRNA knockdown of Bmi-1 reveals a critical role for p21-Rb pathway in NSC self-renewal during development. Cell Stem Cell 1, 87–99 (2007).
Harada, H. et al. Localization of putative stem cells in dental epithelium and their association with Notch and FGF signaling. J. Cell Biol. 147, 105–120 (1999).
Wang, X. P. et al. An integrated gene regulatory network controls stem cell proliferation in teeth. PLoS Biol. 5, 1324–1333 (2007).
Klein, O. D. et al. An FGF signaling loop sustains the generation of differentiated progeny from stem cells in mouse incisors. Development 135, 377–385 (2008).
Smith, C. E. & Warshawsky, H. Cellular renewal in the enamel organ and the odontoblast layer of the rat incisor as followed by radioautography using 3H-thymidine. Anat. Rec. 183, 523–561 (1975).
Seidel, K. et al. Hedgehog signaling regulates the generation of ameloblast progenitors in the continuously growing mouse incisor. Development 137, 3753–3761 (2010).
Juuri, E. et al. Sox2+ stem cells contribute to all epithelial lineages of the tooth via Sfrp5+ progenitors. Dev. Cell 23, 317–328 (2012).
Oguro, H. et al. Poised lineage specification in multipotential hematopoietic stem and progenitor cells by the polycomb protein Bmi1. Cell Stem Cell 6, 279–286 (2010).
Hosen, N. et al. Bmi-1-green fluorescent protein-knock-in mice reveal the dynamic regulation of bmi-1 expression in normal and leukemic hematopoietic cells. Stem Cells 25, 1635–1644 (2007).
Tumbar, T. et al. Defining the epithelial stem cell niche in skin. Science 303, 359–363 (2004).
Joyner, A. L. & Zervas, M. Genetic inducible fate mapping in mouse: establishing genetic lineages and defining genetic neuroanatomy in the nervous system. Dev. Dyn. 235, 2376–2385 (2006).
Sangiorgi, E. & Capecchi, M. R. Bmi1 is expressed in vivo in intestinal stem cells. Nat. Genet. 40, 915–920 (2008).
Muzumdar, M. D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007).
Sangiorgi, E. & Capecchi, M. R. Bmi1 lineage tracing identifies a self-renewing pancreatic acinar cell subpopulation capable of maintaining pancreatic organ homeostasis. Proc. Natl Acad. Sci. USA 106, 7101–7106 (2009).
Li, C. Y. et al. E-cadherin regulates the behavior and fate of epithelial stem cells and their progeny in the mouse incisor. Dev. Biol. 366, 357–366 (2012).
Hwang, W. S. & Tonna, E. A. Autoradiographic analysis of labeling indices and migration rates of cellular component of mouse incisors using tritiated thymidine (H3tdr). J. Dent. Res. 44, 42–53 (1965).
Ramalho-Santos, M., Yoon, S., Matsuzaki, Y., Mulligan, R. C. & Melton, D. A. ‘Stemness’: transcriptional profiling of embryonic and adult stem cells. Science 298, 597–600 (2002).
Rock, J. R. et al. Basal cells as stem cells of the mouse trachea and human airway epithelium. Proc. Natl Acad. Sci. USA 106, 12771–12775 (2009).
Tani, H., Morris, R. J. & Kaur, P. Enrichment for murine keratinocyte stem cells based on cell surface phenotype. Proc. Natl Acad. Sci. USA 97, 10960–10965 (2000).
Xiong, J., Mrozik, K., Gronthos, S. & Bartold, P. M. Epithelial cell rests of malassez contain unique stem cell populations capable of undergoing epithelial-mesenchymal transition. Stem Cells Dev. 21, 2012–2025 (2012).
Simmer, J. P., Richardson, A. S., Smith, C. E., Hu, Y. & Hu, J. C. Expression of kallikrein-related peptidase 4 in dental and non-dental tissues. Eur. J. Oral Sci. 119, 226–233 (2011).
Iwasaki, K. et al. Amelotin–a novel secreted, ameloblast-specific protein. J. Dental Res. 84, 1127–1132 (2005).
Chavez, M. G. et al. Characterization of dental epithelial stem cells from the mouse incisor with two-dimensional and three-dimensional platforms. Tissue Eng. C 19, 15–24 (2013).
Morillo Prado, J. R., Chen, X. & Fuller, M. T. Polycomb group genes Psc and Su(z)2 maintain somatic stem cell identity and activity in Drosophila. PloS ONE 7, e52892 (2012).
Ahn, S. & Joyner, A. L. Dynamic changes in the response of cells to positive hedgehog signaling during mouse limb patterning. Cell 118, 505–516 (2004).
Bai, C. B., Auerbach, W., Lee, J. S., Stephen, D. & Joyner, A. L. Gli2, but not Gli1, is required for initial Shh signaling and ectopic activation of the Shh pathway. Development 129, 4753–4761 (2002).
Diamond, I., Owolabi, T., Marco, M., Lam, C. & Glick, A. Conditional gene expression in the epidermis of transgenic mice using the tetracycline-regulated transactivators tTA and rTA linked to the keratin 5 promoter. J. Invest. Dermatol. 115, 788–794 (2000).
Sousa, V. H., Miyoshi, G., Hjerling-Leffler, J., Karayannis, T. & Fishell, G. Characterization of Nkx6-2-derived neocortical interneuron lineages. Cereb. Cortex 19, i1-10 (2009).
Jung, H. et al. Global control of motor neuron topography mediated by the repressive actions of a single hox gene. Neuron 67, 781–796 (2010).
Rochat, A., Kobayashi, K. & Barrandon, Y. Location of stem cells of human hair follicles by clonal analysis. Cell 76, 1063–1073 (1994).
Blanpain, C., Lowry, W. E., Geoghegan, A., Polak, L. & Fuchs, E. Self-renewal, multipotency, and the existence of two cell populations within an epithelial stem cell niche. Cell 118, 635–648 (2004).
Barrandon, Y. & Green, H. Three clonal types of keratinocyte with different capacities for multiplication. Proc. Natl Acad. Sci. USA 84, 2302–2306 (1987).
Bolstad, B. M., Irizarry, R. A., Astrand, M. & Speed, T. P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003).
Acknowledgements
We thank members of the Klein laboratory for helpful advice, F. Michon for discussion, D-K. Tran and S. Alto for technical assistance, X-P. Wang for help with the culture system, A. Barczak and the UCSF Microarray Core Facilities for help with experiments and analysis, and Irving Weissman for mice. This work was supported by R01-DE021420 (NIH/NIDCR) and a CIRM New Faculty Award II, both to O.D.K.
Author information
Authors and Affiliations
Contributions
B.B., J.K-H.H., N.B.S., H.J., E.S., R-P.H., A.F.G., J.S.D and O.D.K. designed and performed experiments. B.B., J.K-H.H and O.D.K. wrote the manuscript. All authors discussed results, analysed data and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 2415 kb)
Supplementary Table 1
Supplementary Information (XLS 56 kb)
Supplementary Table 2
Supplementary Information (XLS 27 kb)
Supplementary Table 3
Supplementary Information (XLS 80 kb)
Supplementary Table 4
Supplementary Information (XLS 115 kb)
Supplementary Table 5
Supplementary Information (XLS 196 kb)
Supplementary Table 6
Supplementary Information (XLS 35 kb)
Supplementary Table 7
Supplementary Information (XLS 33 kb)
Rights and permissions
About this article
Cite this article
Biehs, B., Hu, JH., Strauli, N. et al. BMI1 represses Ink4a/Arf and Hox genes to regulate stem cells in the rodent incisor. Nat Cell Biol 15, 846–852 (2013). https://doi.org/10.1038/ncb2766
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2766
This article is cited by
-
BMI1 promotes osteosarcoma proliferation and metastasis by repressing the transcription of SIK1
Cancer Cell International (2022)
-
Distinct tooth regeneration systems deploy a conserved battery of genes
EvoDevo (2021)
-
Inverse and reciprocal regulation of p53/p21 and Bmi-1 modulates vasculogenic differentiation of dental pulp stem cells
Cell Death & Disease (2021)
-
LncRNA MALAT1 Functions as a Competing Endogenous RNA to Regulate BMI1 Expression by Sponging miR-200c/miR-203 in the Control of the Differentiation of Pulp Cells
Biochemical Genetics (2021)
-
Loss of PHF6 leads to aberrant development of human neuron-like cells
Scientific Reports (2020)