Abstract
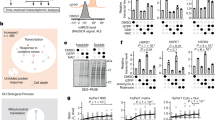
Protein misfolding in the endoplasmic reticulum (ER) leads to cell death through PERK-mediated phosphorylation of eIF2α, although the mechanism is not understood. ChIP-seq and mRNA-seq of activating transcription factor 4 (ATF4) and C/EBP homologous protein (CHOP), key transcription factors downstream of p-eIF2α, demonstrated that they interact to directly induce genes encoding protein synthesis and the unfolded protein response, but not apoptosis. Forced expression of ATF4 and CHOP increased protein synthesis and caused ATP depletion, oxidative stress and cell death. The increased protein synthesis and oxidative stress were necessary signals for cell death. We show that eIF2α-phosphorylation-attenuated protein synthesis, and not Atf4 mRNA translation, promotes cell survival. These results show that transcriptional induction through ATF4 and CHOP increases protein synthesis leading to oxidative stress and cell death. The findings suggest that limiting protein synthesis will be therapeutic for diseases caused by protein misfolding in the ER.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Walter, P. & Ron, D. The unfolded protein response: from stress pathway to homeostatic regulation. Science 334, 1081–1086 (2011).
Hetz, C. The unfolded protein response: controlling cell fate decisions under ER stress and beyond. Nat. Rev. Mol. Cell Biol. 13, 89–102 (2012).
Wang, S. & Kaufman, R. J. The impact of the unfolded protein response on human disease. J. Cell Biol. 197, 857–867 (2012).
Harding, H. P. et al. Regulated translation initiation controls stress-induced gene expression in mammalian cells. Mol. Cell 6, 1099–1108 (2000).
Scheuner, D. et al. Translational control is required for the unfolded protein response and in vivo glucose homeostasis. Mol. Cell 7, 1165–1176 (2001).
Hai, T. W., Liu, F., Coukos, W. J. & Green, M. R. Transcription factor ATF cDNA clones: an extensive family of leucine zipper proteins able to selectively form DNA-binding heterodimers. Genes Dev. 3, 2083–2090 (1989).
Lu, P. D., Harding, H. P. & Ron, D. Translation reinitiation at alternative open reading frames regulates gene expression in an integrated stress response. J. Cell Biol. 167, 27–33 (2004).
Vattem, K. M. & Wek, R. C. Reinitiation involving upstream ORFs regulates ATF4 mRNA translation in mammalian cells. Proc. Natl Acad. Sci. USA 101, 11269–11274 (2004).
Fornace, A. J. Jr, Alamo, I. Jr & Hollander, M. C. DNA damage-inducible transcripts in mammalian cells. Proc. Natl Acad. Sci. USA 85, 8800–8804 (1988).
Ron, D. & Habener, J. F. CHOP, a novel developmentally regulated nuclear protein that dimerizes with transcription factors C/EBP and LAP and functions as a dominant-negative inhibitor of gene transcription. Genes. Dev. 6, 439–453 (1992).
Ma, Y., Brewer, J. W., Diehl, J. A. & Hendershot, L. M. Two distinct stress signalling pathways converge on the CHOP promoter during the mammalian unfolded protein response. J. Mol. Biol. 318, 1351–1365 (2002).
Zinszner, H. et al. CHOP is implicated in programmed cell death in response to impaired function of the endoplasmic reticulum. Genes Dev. 12, 982–995 (1998).
Oyadomari, S. et al. Targeted disruption of the Chop gene delays endoplasmic reticulum stress-mediated diabetes. J. Clin. Invest. 109, 525–532 (2002).
Song, B., Scheuner, D., Ron, D., Pennathur, S. & Kaufman, R. J. Chop deletion reduces oxidative stress, improves β cell function, and promotes cell survival in multiple mouse models of diabetes. J. Clin. Invest. 118, 3378–3389 (2008).
Malhotra, J. D. et al. Antioxidants reduce endoplasmic reticulum stress and improve protein secretion. Proc. Natl Acad. Sci. USA 105, 18525–18530 (2008).
Thorp, E. et al. Reduced apoptosis and plaque necrosis in advanced atherosclerotic lesions of Apoe-/- and Ldlr-/- mice lacking CHOP. Cell Metab. 9, 474–481 (2009).
Tabas, I. & Ron, D. Integrating the mechanisms of apoptosis induced by endoplasmic reticulum stress. Nat. Cell Biol. 13, 184–190 (2011).
Pennuto, M. et al. Ablation of the UPR-mediator CHOP restores motor function and reduces demyelination in Charcot-Marie-Tooth 1B mice. Neuron 57, 393–405 (2008).
McCullough, K. D., Martindale, J. L., Klotz, L. O., Aw, T. Y. & Holbrook, N. J. Gadd153 sensitizes cells to endoplasmic reticulum stress by down-regulating Bcl2 and perturbing the cellular redox state. Mol. Cell Biol. 21, 1249–1259 (2001).
Chikka, M. R., McCabe, D. D., Tyra, H. M. & Rutkowski, D. T. C/EBP Homologous Protein (CHOP) contributes to suppression of metabolic genes during endoplasmic reticulum stress in the liver. J. Biol. Chem. 288, 4405–4415 (2013).
Rutkowski, D. T. et al. Adaptation to ER stress is mediated by differential stabilities of pro-survival and pro-apoptotic mRNAs and proteins. PLoS Biol. 4, e374 (2006).
Wang, X. Z. et al. Identification of novel stress-induced genes downstream of chop. EMBO J. 17, 3619–3630 (1998).
Harding, H. P. et al. An integrated stress response regulates amino acid metabolism and resistance to oxidative stress. Mol. Cell 11, 619–633 (2003).
Sun, X. et al. ATF4 protects against neuronal death in cellular Parkinson’s disease models by maintaining levels of parkin. J. Neurosci. 33, 2398–2407 (2013).
Pike, L. R. et al. Transcriptional up-regulation of ULK1 by ATF4 contributes to cancer cell survival. Biochem. J. 449, 389–400 (2013).
Ord, D., Meerits, K. & Ord, T. TRB3 protects cells against the growth inhibitory and cytotoxic effect of ATF4. Exp. Cell Res. 313, 3556–3567 (2007).
Lange, P. S. et al. ATF4 is an oxidative stress-inducible, prodeath transcription factor in neurons in vitro and in vivo. J. Exp. Med. 205, 1227–1242 (2008).
Galehdar, Z. et al. Neuronal apoptosis induced by endoplasmic reticulum stress is regulated by ATF4-CHOP-mediated induction of the Bcl-2 homology 3-only member PUMA. J. Neurosci. 30, 16938–16948 (2010).
Armstrong, J. L., Flockhart, R., Veal, G. J., Lovat, P. E. & Redfern, C. P. Regulation of endoplasmic reticulum stress-induced cell death by ATF4 in neuroectodermal tumour cells. J. Biol. Chem. 285, 6091–6100 (2010).
Qing, G. et al. ATF4 regulates MYC-mediated neuroblastoma cell death on glutamine deprivation. Cancer Cell 22, 631–644 (2012).
Ubeda, M. et al. Stress-induced binding of the transcriptional factor CHOP to a novel DNA control element. Mol. Cell Biol. 16, 1479–1489 (1996).
Chen, B. P., Wolfgang, C. D. & Hai, T. Analysis of ATF3, a transcription factor induced by physiological stresses and modulated by gadd153/Chop10. Mol. Cell Biol. 16, 1157–1168 (1996).
Marciniak, S. J. et al. CHOP induces death by promoting protein synthesis and oxidation in the stressed endoplasmic reticulum. Gen. Dev. 18, 3066–3077 (2004).
Ma, Y. & Hendershot, L. M. Delineation of a negative feedback regulatory loop that controls protein translation during endoplasmic reticulum stress. J. Biol. Chem. 278, 34864–34873 (2003).
Ohoka, N., Yoshii, S., Hattori, T., Onozaki, K. & Hayashi, H. TRB3, a novel ER stress-inducible gene, is induced via ATF4-CHOP pathway and is involved in cell death. EMBO J. 24, 1243–1255 (2005).
Gill, G. & Ptashne, M. Negative effect of the transcriptional activator GAL4. Nature 334, 721–724 (1988).
Valouev, A. et al. Genome-wide analysis of transcription factor binding sites based on ChIP-seq data. Nat. Methods 5, 829–834 (2008).
Su, N. & Kilberg, M. S. C/EBP Homology Protein (CHOP) Interacts with Activating Transcription Factor 4 (ATF4) and negatively regulates the stress-dependent induction of the asparagine synthetase gene. J. Biol. Chem. 283, 35106–35117 (2008).
Bromati, C. R. et al. UPR induces transient burst of apoptosis in islets of early lactating rats through reduced AKT phosphorylation via ATF4/CHOP stimulation of TRB3 expression. Am. J. Physiol. Regul. Integr. Comp. Physiol. 300, R92–R100 (2011).
Novoa, I., Zeng, H., Harding, H. P. & Ron, D. Feedback inhibition of the unfolded protein response by GADD34-mediated dephosphorylation of eIF2α. J. Cell Biol. 153, 1011–1022 (2001).
Kimball, S. R., Farrell, P. A. & Jefferson, L. S. Invited review: Role of insulin in translational control of protein synthesis in skeletal muscle by amino acids or exercise. J. Appl. Physiol. 93, 1168–1180 (2002).
Anderson, L. L., Mao, X., Scott, B. A. & Crowder, C. M. Survival from hypoxia in C. elegans by inactivation of aminoacyl-tRNA synthetases. Science 323, 630–633 (2009).
Back, S. H. et al. Translation attenuation through eIF2α phosphorylation prevents oxidative stress and maintains the differentiated state in β cells. Cell Metab. 10, 13–26 (2009).
Boyce, M. et al. A selective inhibitor of eIF2α dephosphorylation protects cells from ER stress. Science 307, 935–939 (2005).
Tsaytler, P., Harding, H. P., Ron, D. & Bertolotti, A. Selective inhibition of aregulatory subunit of protein phosphatase 1 restores proteostasis. Science 332, 91–94 (2011).
Yamaguchi, S. et al. ATF4-Mediated Induction of 4E-BP1 contributes to pancreatic beta cell survival under endoplasmic reticulum stress. Cell Metab. 7, 269–276 (2008).
Oliver, E. R., Saunders, T. L., Tarle, S. A. & Glaser, T. Ribosomal protein L24defect in belly spot and tail (Bst), a mouse Minute. Development 131, 3907–3920 (2004).
Lu, P. D. et al. Cytoprotection by pre-emptive conditional phosphorylation of translation initiation factor 2. EMBO J. 23, 169–179 (2004).
Enyedi, B., Varnai, P. & Geiszt, M. Redox state of the endoplasmic reticulum is controlled by Ero1L- α and intraluminal calcium. Antioxid. Redox Signal. 13, 721–729 (2010).
Yuan, C. L. et al. Preserved protein synthesis in the heart in response to acute fasting and chronic food restriction despite reductions in liver and skeletal muscle. Am. J. Physiol. Endocrinol. Metabol. 295, E216–E222 (2008).
Puthalakath, H. et al. ER stress triggers apoptosis by activating BH3-only protein Bim. Cell 129, 1337–1349 (2007).
Ghosh, A. P., Klocke, B. J., Ballestas, M. E. & Roth, K. A. CHOP potentially co-operates with FOXO3a in neuronal cells to regulate PUMA and BIM expression in response to ER stress. PloS One 7, e39586 (2012).
Li, G. et al. Role of ERO1- α-mediated stimulation of inositol 1,4,5-triphosphate receptor activity in endoplasmic reticulum stress-induced apoptosis. J. Cell Biol. 186, 783–792 (2009).
Huang, C. C. et al. A bifunctional intronic element regulates the expression of the arginine/lysine transporter Cat-1 via mechanisms involving the purine-rich element binding protein A (Pur α). J. Biol. Chem. 284, 32312–32320 (2009).
Kilberg, M. S., Balasubramanian, M., Fu, L. & Shan, J. The transcription factor network associated with the amino acid response in mammalian cells. Adv. Nutr. 3, 295–306 (2012).
Harding, H. P., Zyryanova, A. F. & Ron, D. Uncoupling proteostasis and development in vitro with a small molecule inhibitor of the pancreatic endoplasmic reticulum kinase, PERK. J. Biol. Chem. 287, 44338–44344 (2012).
Palam, L. R., Baird, T. D. & Wek, R. C. Phosphorylation of eIF2 facilitates ribosomal bypass of an inhibitory upstream ORF to enhance CHOP translation. J. Biol. Chem. 286, 10939–10949 (2011).
Johnson, D. S., Mortazavi, A., Myers, R. M. & Wold, B. Genome-Wide Mapping of in vivo Protein-DNA Interactions. Science 316, 1497–1502 (2007).
Wilbanks, E. G. & Facciotti, M. T. Evaluation of algorithm performance in ChIP-seq peak detection. PLoS One 5, e11471 (2010).
Sartor, M. et al. Intensity-based hierarchical Bayes method improves testing for differentially expressed genes in microarray experiments. BMC Bioinformatics 7, 538 (2006).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Sartor, M. A. et al. ConceptGen: a gene set enrichment and gene set relation mapping tool. Bioinformatics 26, 456–463 (2010).
Bailey, T., Boden, M., Whitington, T. & Machanick, P. The value of position-specific priors in motif discovery using MEME. BMC Bioinformatics 11, 179 (2010).
Hettmann, T., Barton, K. & Leiden, J. M. Microphthalmia due to p53-mediated apoptosis of anterior lens epithelial cells in mice lacking the CREB-2 transcription factor. Dev. Biol. 222, 110–123 (2000).
Acknowledgements
We thank the University of Michigan Sequencing Core for next-generation sequencing, D. Ron (University of Cambridge, UK) for recombinant retroviral vectors expressing GADD34 derivatives and GFP and Ratan (Cornell University, USA) for adenovirus expressing ATF4 and ATF4ΔRK. We are grateful to members of the R.J.K. laboratory for assistance, advice and stimulating discussions. This work was supported by a University of Michigan CCMB Pilot Grant (J.H.), NIH grants (HL057346, DK042394, DK088227, HL052173, DK093074 (R.J.K.)), (DK092062, DK094729 (M.S.K.)), (DK60596, DK53307 (M.H.)), and the National Research Foundation of Korea (NRF) grants 2010-0001199, 2011-0011433, 2012M3A9C3048686 (S.H.B.).
Author information
Authors and Affiliations
Contributions
J.H., S.H.B. and R.J.K. designed all experiments and performed most of them. J.H., Y-H.L and M.A.S. performed bioinformatic analysis and contributed to preparation of figures and tables. R.G. performed western blot and cell viability assays using Eif2 αA/A and Atf4−/− MEFs and GADD34 overexpression experiments. J.S. and M.S.K. generated and characterized the ATF4 antibody and performed co-immunoprecipitation and sequential ChIP experiments. C.L.Y., D.K. and M.H. measured in vivo protein synthesis using 2H2O. S.W. analysed protein synthesis and western blots. J.H., S.H.B. and R.J.K. prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1038 kb)
Supplementary Table 1
Supplementary Information (XLSX 57 kb)
Supplementary Table 2
Supplementary Information (XLSX 10 kb)
Supplementary Table 3
Supplementary Information (XLSX 115 kb)
Supplementary Table 4
Supplementary Information (XLSX 156 kb)
Supplementary Table 5
Supplementary Information (XLSX 14 kb)
Supplementary Table 6
Supplementary Information (XLSX 10 kb)
Supplementary Table 7
Supplementary Information (XLSX 14 kb)
Supplementary Information
Supplementary Information (PDF 747 kb)
Rights and permissions
About this article
Cite this article
Han, J., Back, S., Hur, J. et al. ER-stress-induced transcriptional regulation increases protein synthesis leading to cell death. Nat Cell Biol 15, 481–490 (2013). https://doi.org/10.1038/ncb2738
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2738
This article is cited by
-
A beneficial adaptive role for CHOP in driving cell fate selection during ER stress
EMBO Reports (2024)
-
Targeting paraptosis in cancer: opportunities and challenges
Cancer Gene Therapy (2024)
-
Glutamate is effective in decreasing opacity formed in galactose-induced cataract model
Scientific Reports (2024)
-
Mitochondrial dysfunction abrogates dietary lipid processing in enterocytes
Nature (2024)
-
UBXN1 maintains ER proteostasis and represses UPR activation by modulating translation
EMBO Reports (2024)