Abstract
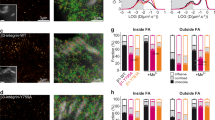
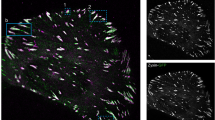
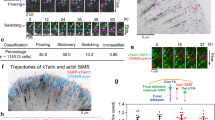
Integrins in focal adhesions (FAs) mediate adhesion and force transmission to extracellular matrices essential for cell motility, proliferation and differentiation. Different fibronectin-binding integrins, simultaneously present in FAs, perform distinct functions. Yet, how integrin dynamics control biochemical and biomechanical processes in FAs is still elusive. Using single-protein tracking and super-resolution imaging we revealed the dynamic nano-organizations of integrins and talin inside FAs. Integrins reside in FAs through free-diffusion and immobilization cycles. Integrin activation promotes immobilization, stabilized in FAs by simultaneous connection to fibronectin and actin-binding proteins. Talin is recruited in FAs directly from the cytosol without membrane free-diffusion, restricting integrin immobilization to FAs. Immobilized β3-integrins are enriched and stationary within FAs, whereas immobilized β1-integrins are less enriched and exhibit rearward movements. Talin is enriched and mainly stationary, but also exhibited rearward movements in FAs, consistent with stable connections with both β-integrins. Thus, differential transmission of actin motion to fibronectin occurs through specific integrins within FAs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Geiger, B., Spatz, J. P. & Bershadsky, A. D. Environmental sensing through focal adhesions. Nat. Rev. Mol. Cell Biol. 10, 21–33 (2009).
Giannone, G., Mege, R. M. & Thoumine, O. Multi-level molecular clutches in motile cell processes. Trends Cell Biol. 19, 475–486 (2009).
Parsons, J. T., Horwitz, A. R. & Schwartz, M. A. Cell adhesion: integrating cytoskeletal dynamics and cellular tension. Nat. Rev. Mol. Cell Biol. 11, 633–643 (2010).
Zaidel-Bar, R., Itzkovitz, S., Ma’ayan, A., Iyengar, R. & Geiger, B. Functional atlas of the integrin adhesome. Nat. Cell Biol. 9, 858–867 (2007).
Sheetz, M. P., Felsenfeld, D., Galbraith, C. G. & Choquet, D. Cell migration as a five-step cycle. Biochem. Soc. Symp. 65, 233–243 (1999).
Giannone, G. et al. Lamellipodial actin mechanically links myosin activity with adhesion-site formation. Cell 128, 561–575 (2007).
Tkachenko, E. et al. Protein kinase A governs a RhoA-RhoGDI protrusion-retraction pacemaker in migrating cells. Nat. Cell Biol. 13, 660–667 (2011).
Pollard, T. D. & Borisy, G. G. Cellular motility driven by assembly and disassembly of actin filaments. Cell 112, 453–465 (2003).
Kanchanawong, P. et al. Nanoscale architecture of integrin-based cell adhesions. Nature 468, 580–584 (2010).
Zamir, E. et al. Dynamics and segregation of cell-matrix adhesions in cultured fibroblasts. Nat. Cell Biol. 2, 191–196 (2000).
Danen, E. H., Sonneveld, P., Brakebusch, C., Fassler, R. & Sonnenberg, A. The fibronectin-binding integrins α5β1 and αvβ3 differentially modulate RhoA-GTP loading, organization of cell matrix adhesions, and fibronectin fibrillogenesis. J. Cell Biol. 159, 1071–1086 (2002).
Roca-Cusachs, P., Gauthier, N. C., Del Rio, A. & Sheetz, M. P. Clustering of α(5)β(1) integrins determines adhesion strength whereas α(v)β(3) and talin enable mechanotransduction. Proc. Natl Acad. Sci. USA 106, 16245–16250 (2009).
Choquet, D., Felsenfeld, D. P. & Sheetz, M.P. Extracellular matrix rigidity causes strengthening of integrin-cytoskeleton linkages. Cell 88, 39–48 (1997).
Jiang, G., Giannone, G., Critchley, D. R., Fukumoto, E. & Sheetz, M. P. Two-piconewton slip bond between fibronectin and the cytoskeleton depends on talin. Nature 424, 334–337 (2003).
Giannone, G., Jiang, G., Sutton, D. H., Critchley, D. R. & Sheetz, M. P. Talin1 is critical for force-dependent reinforcement of initial integrin-cytoskeleton bonds but not tyrosine kinase activation. J. Cell Biol. 163, 409–419 (2003).
Tadokoro, S. et al. Talin binding to integrin β tails: a final common step in integrin activation. Science 302, 103–106 (2003).
Moser, M., Legate, K. R., Zent, R. & Fassler, R. The tail of integrins, talin, and kindlins. Science 324, 895–899 (2009).
Shattil, S. J., Kim, C. & Ginsberg, M. H. The final steps of integrin activation: the end game. Nat. Rev. Mol. Cell Biol. 11, 288–300 (2010).
Manley, S. et al. High-density mapping of single-molecule trajectories with photoactivated localization microscopy. Nature Meth. 5, 155 (2008).
Luo, B. H., Springer, T. A. & Takagi, J. Stabilizing the open conformation of the integrin headpiece with a glycan wedge increases affinity for ligand. Proc. Natl Acad. Sci. USA 100, 2403–2408 (2003).
Cluzel, C. et al. The mechanisms and dynamics of (α)v(β)3 integrin clustering in living cells. J. Cell Biol. 171, 383–392 (2005).
O’Toole, T. E., Ylanne, J. & Culley, B. M. Regulation of integrin affinity states through an NPXY motif in the β subunit cytoplasmic domain. J. Biol. Chem. 270, 8553–8558 (1995).
Loftus, J. C. et al. A β3 integrin mutation abolishes ligand binding and alters divalent cation-dependent conformation. Science 249, 915–918 (1990).
McCleverty, C. J., Lin, D. C. & Liddington, R. C. Structure of the PTB domain of tensin1 and a model for its recruitment to fibrillar adhesions. Prot. Sci. 16, 1223–1229 (2007).
Legate, K. R. & Fassler, R. Mechanisms that regulate adaptor binding to β-integrin cytoplasmic tails. J. Cell Sci. 122, 187–198 (2009).
Humphries, J. D. et al. Vinculin controls focal adhesion formation by direct interactions with talin and actin. J. Cell Biol. 179, 1043–1057 (2007).
Shroff, H., Galbraith, C. G., Galbraith, J. A. & Betzig, E. Live-cell photoactivated localization microscopy of nanoscale adhesion dynamics. Nature Meth. 5, 417–423 (2008).
Lata, S., Gavutis, M., Tampe, R. & Piehler, J. Specific and stable fluorescence labeling of histidine-tagged proteins for dissecting multi-protein complex formation. J. Am. Chem. Soc. 128, 2365–2372 (2006).
Giannone, G. et al. Dynamic superresolution imaging of endogenous proteins on living cells at ultra-high density. Biophys. J. 99, 1303–1310 (2010).
Brown, C. M. et al. Probing the integrin-actin linkage using high-resolution protein velocity mapping. J. Cell Sci. 119, 5204–5214 (2006).
Hu, K., Ji, L., Applegate, K. T., Danuser, G. & Waterman-Storer, C. M. Differential transmission of actin motion within focal adhesions. Science 315, 111–115 (2007).
Kiema, T. et al. The molecular basis of filamin binding to integrins and competition with talin. Mol. Cell 21, 337–347 (2006).
Wegener, K. L. et al. Structural basis of integrin activation by talin. Cell 128, 171–182 (2007).
Ye, F. et al. Recreation of the terminal events in physiological integrin activation. J. Cell Biol. 188, 157–173 (2010).
Zhang, X. et al. Talin depletion reveals independence of initial cell spreading from integrin activation and traction. Nat. Cell Biol. 10, 1062–1068 (2008).
Goksoy, E. et al. Structural basis for the autoinhibition of talin in regulating integrin activation. Mol. Cell 31, 124–133 (2008).
Banno, A. et al. Subcellular localization of talin is regulated by inter-domain interactions. J. Biol. Chem. 287, 13799–13812 (2012).
Martel, V. et al. Conformation, localization, and integrin binding of talin dependon its interaction with phosphoinositides. J. Biol. Chem. 276, 21217–21227 (2001).
Saltel, F. et al. New PI(4,5)P2- and membrane proximal integrin-bindingmotifs in the talin head control β3-integrin clustering. J. Cell Biol. 187, 715–731 (2009).
Pankov, R. et al. Integrin dynamics and matrix assembly: tensin-dependent translocation of α(5)β(1) integrins promotes early fibronectin fibrillogenesis. J. Cell Biol. 148, 1075–1090 (2000).
Humphries, J. D., Byron, A. & Humphries, M. J. Integrin ligands at a glance. J. Cell Sci. 119, 3901–3903 (2006).
Huveneers, S., Truong, H., Fassler, R., Sonnenberg, A. & Danen, E. H. Binding of soluble fibronectin to integrin α5β1—link to focal adhesion redistribution and contractile shape. J. Cell Sci. 121, 2452–2462 (2008).
Anthis, N. J., Wegener, K. L., Critchley, D. R. & Campbell, I. D. Structural diversity in integrin/talin interactions. Structure 18, 1654–1666 (2010).
Friedland, J. C., Lee, M. H. & Boettiger, D. Mechanically activated integrin switch controls α5β1 function. Science 323, 642–644 (2009).
Westphal, V. et al. Video-rate far-field optical nanoscopy dissects synaptic vesicle movement. Science 320, 246–249 (2008).
Arias-Salgado, E. G. et al. Src kinase activation by direct interaction with the integrin β cytoplasmic domain. Proc. Natl Acad. Sci. USA 100, 13298–13302 (2003).
Millon-Fremillon, A. et al. Cell adaptive response to extracellular matrix density is controlled by ICAP-1-dependent β1-integrin affinity. J. Cell Biol. 180, 427–441 (2008).
Worth, D. C. et al. αvβ3 integrin spatially regulates VASP and RIAM to control adhesion dynamics and migration. J. Cell Biol. 189, 369–383 (2010).
Rantala, J. K. et al. SHARPIN is an endogenous inhibitor of [β]1-integrin activation. Nat. Cell Biol. 13, 1315–1324 (2011).
Tanentzapf, G. & Brown, N. H. An interaction between integrin and the talin FERM domain mediates integrin activation but not linkage to the cytoskeleton. Nat. Cell Biol. 8, 601–606 (2006).
Himmel, M. et al. Control of high affinity interactions in the talin C terminus: how talin domains coordinate protein dynamics in cell adhesions. J. Biol. Chem. 284, 13832–13842 (2009).
Wang, P., Ballestrem, C. & Streuli, C. H. The C terminus of talin links integrins to cell cycle progression. J. Cell Biol. 195, 499–513 (2011).
Galbraith, C. G., Yamada, K. M. & Galbraith, J. A. Polymerizing actin fibers position integrins primed to probe for adhesion sites. Science 315, 992–995 (2007).
Margadant, F. et al. Mechanotransduction in vivo by repeated talin stretch-relaxation events depends upon vinculin. PLoS Biol. 9, e1001223 (2011).
Del Rio, A. et al. Stretching single talin rod molecules activates vinculin binding. Science 323, 638–641 (2009).
Kong, F., Garcia, A. J., Mould, A. P., Humphries, M. J. & Zhu, C. Demonstration of catch bonds between an integrin and its ligand. J. Cell Biol. 185, 1275–1284 (2009).
Smith, M. L. et al. Force-induced unfolding of fibronectin in the extracellular matrix of living cells. PLoS Biol. 5, e268 (2007).
Rossier, O. M. et al. Force generated by actomyosin contraction builds bridges between adhesive contacts. EMBO J. 29, 1055–1068 (2010).
Parsons, M., Messent, A. J., Humphries, J. D., Deakin, N. O. & Humphries, M. J. Quantification of integrin receptor agonism by fluorescence lifetime imaging. J. Cell Sci. 121, 265–271 (2008).
Plançon, S., Morel-Kopp, M. C., Schaffner-Reckinger, E., Chen, P. & Kieffer, N. Green fluorescent protein (GFP) tagged to the cytoplasmic tail of αIIb or β3 allows the expression of a fully functional integrin αIIb(β3): effect of β3GFP on αIIb(β3) ligand binding. Biochem. J. 357, 529–536 (2001).
Chen, I., Howarth, M., Lin, W. & Ting, A. Y. Site-specific labelling of cell surface proteins with biophysical probes using biotin ligase. Nature Meth. 2, 99–104 (2005).
Rottner, K., Behrendt, B., Small, J. V. & Wehland, J. VASP dynamics during lamellipodia protrusion. Nat. Cell Biol. 1, 321–322 (1999).
Racine, V. et al. Multiple-target tracking of 3D fluorescent objects based on simulated annealing. IEEE Int. Symp. Biomed. Imag. 1020–1023 (2006).
Racine, V. et al. Visualization and quantification of vesicle trafficking on a three-dimensional cytoskeleton network in living cells. J. Microsci. 225, 214–228 (2007).
Izeddin, I. et al. Wavelet analysis for single molecule localization microscopy. Opt. Exp. 20, 2081–2095 (2012).
Cheezum, M. K., Walker, W. F. & Guilford, W. H. Quantitative comparison of algorithms for tracking single fluorescent particles. Biophys. J. 81, 2378–2388 (2001).
Tardin, C., Cognet, L., Bats, C., Lounis, B. & Choquet, D. Direct imaging of lateral movements of AMPA receptors inside synapses. Embo J. 22, 4656–4665 (2003).
Annibale, P., Scarselli, M., Kodiyan, A. & Radenovic, A. Photoactivatable fluorescent protein mEos2 displays repeated photoactivation after a long-lived dark state in the red photoconverted form. J. Phys. Chem. Lett. 1, 1506–1510 (2010).
Grunwald, C. et al. Quantum-yield-optimized fluorophores for site-specific labelling and super-resolution imaging. J. Am. Chem. Soc. 133, 8090–8093 (2011).
Groc, L. et al. Surface trafficking of neurotransmitter receptor: comparison between single-molecule/quantum dot strategies. J. Neurosci. 27, 12433–12437 (2007).
Acknowledgements
We thank C. Breillat, A. Frouin, D. Bouchet and P. Gonzales for technical assistance; M. P. Sheetz, O. Thoumine, J. Petersen, B. Fourcade, M. Block, L. Duchesne and D.G. Fernig for helpful discussions; M. Humphries, N. Kieffer, J. Wehland, A. Gautreau and P. Kanchanawong for the gift of reagents; P. Legros and C. Poujol (Bordeaux Imaging Center) for STED imaging. We acknowledge financial support from the French Ministry of Research and CNRS, ANR grant Nanomotility (G.G., B.L., O.R.), Fondation ARC pour la Recherche sur le Cancer (O.R.), Conseil Régional Aquitaine, Fondation pour la Recherche Médicale, the ERC Program numbers 232942 Nano-Dyn-Syn (D.C., B.L.) and 235552 Glutraf (D.N.), the Human Frontiers Science Programme (B.L.) and The Ligue National contre le Cancer—équipe labellisée 2010 (C.A-R., O.D.). The research was conducted in the scope of the International Associated Laboratory LIA CAFS.
Author information
Authors and Affiliations
Contributions
O.R. and G.G. conceptualized and performed the sptPALM and STED experiments. V.O., C.L., L.C. and B.L. conceptualized and developed the single-protein-tracking set-up used for ATTO647N. G.G. performed single-protein-tracking experiments using ATTO647N. B.T. designed and generated new protein constructs. V.G. and R.T. designed and synthesized the TrisNTA–ATTO647N. D.N. and J-B.S. developed the sptPALM set-up. J-B.S. developed the analytical tools for sptPALM. V.O., C.L. and L.C. developed the analytical tools for single-protein tracking using ATTO647N. O.D. and C.A-R. contributed valuable scientific advice and developed the β1-integrin–mEOS2 and chimaeric integrin constructs. O.D. performed FACS experiments. L.C., D.C., B.L. and G.G. coordinated the study. O.R. and G.G. wrote the manuscript and Supplementary Information. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1989 kb)
Supplementary Table 1
Supplementary Information (XLSX 17 kb)
Supplementary Table 2
Supplementary Information (XLSX 13 kb)
Supplementary Table 3
Supplementary Information (XLSX 11 kb)
Supplementary Table 4
Supplementary Information (XLSX 11 kb)
Supplementary Table 5
Supplementary Information (XLSX 11 kb)
Supplementary Movie 1
Supplementary Information (AVI 886 kb)
Rights and permissions
About this article
Cite this article
Rossier, O., Octeau, V., Sibarita, JB. et al. Integrins β1 and β3 exhibit distinct dynamic nanoscale organizations inside focal adhesions. Nat Cell Biol 14, 1057–1067 (2012). https://doi.org/10.1038/ncb2588
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2588
This article is cited by
-
Single-molecule imaging of Tau reveals how phosphorylation affects its movement and confinement in living cells
Molecular Brain (2024)
-
Organization, dynamics and mechanoregulation of integrin-mediated cell–ECM adhesions
Nature Reviews Molecular Cell Biology (2023)
-
Biased activation of the receptor tyrosine kinase HER2
Cellular and Molecular Life Sciences (2023)
-
Monitoring cell membrane recycling dynamics of proteins using whole-cell fluorescence recovery after photobleaching of pH-sensitive genetic tags
Nature Protocols (2022)
-
Rear traction forces drive adherent tissue migration in vivo
Nature Cell Biology (2022)