Abstract
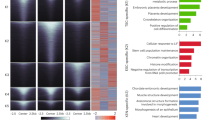
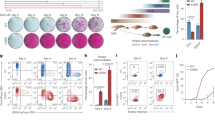
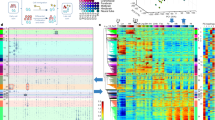
It is becoming clear that interconnected functional gene networks, rather than individual genes, govern stem cell self-renewal and differentiation. To identify epigenetic factors that impact on human epidermal stem cells we performed siRNA-based genetic screens for 332 chromatin modifiers. We developed a Bayesian mixture model to predict putative functional interactions between epigenetic modifiers that regulate differentiation. We discovered a network of genetic interactions involving EZH2, UHRF1 (both known to regulate epidermal self-renewal), ING5 (a MORF complex component), BPTF and SMARCA5 (NURF complex components). Genome-wide localization and global mRNA expression analysis revealed that these factors impact two distinct but functionally related gene sets, including integrin extracellular matrix receptors that mediate anchorage of epidermal stem cells to their niche. Using a competitive epidermal reconstitution assay we confirmed that ING5, BPTF, SMARCA5, EZH2 and UHRF1 control differentiation under physiological conditions. Thus, regulation of distinct gene expression programs through the interplay between diverse epigenetic strategies protects epidermal stem cells from differentiation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Accession codes
References
Barabasi, A. L. & Oltvai, Z. N. Network biology: understanding the cell’s functional organization. Nat. Rev. Genet. 5, 101–113 (2004).
Boutros, M. & Ahringer, J. The art and design of genetic screens: RNA interference. Nat. Rev. Genet. 9, 554–566 (2008).
Mani, R., St Onge, R. P., Hartman, J. L. t., Giaever, G. & Roth, F. P. Defining genetic interaction. Proc. Natl Acad. Sci. USA 105, 3461–3466 (2008).
Perez-Perez, J. M., Candela, H. & Micol, J. L. Understanding synergy in genetic interactions. Trends Genet. 25, 368–376 (2009).
Blanpain, C. & Fuchs, E. Epidermal stem cells of the skin. Annu. Rev. Cell Dev. Biol. 22, 339–373 (2006).
Watt, F. M. Role of integrins in regulating epidermal adhesion, growth and differentiation. EMBO J. 21, 3919–3926 (2002).
Green, H. The birth of therapy with cultured cells. Bioessays 30, 897–903 (2008).
Pellegrini, G. et al. p63 identifies keratinocyte stem cells. Proc. Natl Acad. Sci. USA 98, 3156–3161 (2001).
Koster, M. I. & Roop, D. R. The role of p63 in development and differentiation of the epidermis. J. Dermatol. Sci. 34, 3–9 (2004).
Eckert, R. L., Crish, J. F., Banks, E. B. & Welter, J. F. The epidermis: genes on–genes off. J. Invest. Dermatol. 109, 501–509 (1997).
Driskell, I. et al. The histone methyltransferase Setd8 acts in concert with c-Myc and is required to maintain skin. EMBO J. 31, 616–629 (2012).
Ezhkova, E. et al. EZH1 and EZH2 cogovern histone H3K27 trimethylation and are essential for hair follicle homeostasis and wound repair. Genes Dev. 25, 485–498 (2011).
Ezhkova, E. et al. Ezh2 orchestrates gene expression for the stepwise differentiation of tissue-specific stem cells. Cell 136, 1122–1135 (2009).
Luis, N. M. et al. Regulation of human epidermal stem cell proliferation and senescence requires polycomb- dependent and -independent functions of Cbx4. Cell Stem. Cell 9, 233–246 (2011).
Mejetta, S. et al. Jarid2 regulates mouse epidermal stem cell activation and differentiation. EMBO J. 30, 3635–3646 (2011).
Sen, G. L., Reuter, J. A., Webster, D. E., Zhu, L. & Khavari, P. A. DNMT1 maintains progenitor function in self-renewing somatic tissue. Nature 463, 563–567 (2010).
Sen, G. L., Webster, D. E., Barragan, D. I., Chang, H. Y. & Khavari, P. A. Control of differentiation in a self-renewing mammalian tissue by the histone demethylase JMJD3. Genes Dev. 22, 1865–1870 (2008).
Connelly, J. T. et al. Actin and serum response factor transduce physical cues from the microenvironment to regulate epidermal stem cell fate decisions. Nat. Cell Biol. 12, 711–718 (2010).
Gosselet, F. P., Magnaldo, T., Culerrier, R. M., Sarasin, A. & Ehrhart, J. C. BMP2and BMP6 control p57(Kip2) expression and cell growth arrest/terminal differentiation in normal primary human epidermal keratinocytes. Cell Signal 19, 731–739 (2007).
Kolev, V. et al. EGFR signalling as a negative regulator of Notch1 gene transcription and function in proliferating keratinocytes and cancer. Nat. Cell Biol. 10, 902–911 (2008).
Kuramoto, N. et al. Development of ichthyosiform skin compensates for defective permeability barrier function in mice lacking transglutaminase 1. J. Clin. Invest. 109, 243–250 (2002).
Matsuki, M. et al. Defective stratum corneum and early neonatal death in mice lacking the gene for transglutaminase 1 (keratinocyte transglutaminase). Proc. Natl Acad. Sci. USA 95, 1044–1049 (1998).
Vermeulen, M. et al. Quantitative interaction proteomics and genome-wide profiling of epigenetic histone marks and their readers. Cell 142, 967–980 (2010).
Doyon, Y. et al. ING tumor suppressor proteins are critical regulators of chromatin acetylation required for genome expression and perpetuation. Mol. Cell 21, 51–64 (2006).
Luc, P. V. & Tempst, P. PINdb: a database of nuclear protein complexes from human and yeast. Bioinformatics 20, 1413–1415 (2004).
Rahman, S. et al. The Brd4 extraterminal domain confers transcription activation independent of pTEFb by recruiting multiple proteins, including NSD3. Mol. Cell Biol. 31, 2641–2652 (2011).
Indra, A. K. et al. Temporally controlled targeted somatic mutagenesis in embryonic surface ectoderm and fetal epidermal keratinocytes unveils two distinct developmental functions of BRG1 in limb morphogenesis and skin barrier formation. Development 132, 4533–4544 (2005).
Kashiwagi, M., Morgan, B. A. & Georgopoulos, K. The chromatin remodeler Mi- 2β is required for establishment of the basal epidermis and normal differentiation of its progeny. Development 134, 1571–1582 (2007).
Costanzo, M. et al. The genetic landscape of a cell. Science 327, 425–431 (2010).
Wang, X., Castro, M. A., Mulder, K. W. & Markowetz, F. Posterior association networks and functional modules inferred from rich phenotypes of gene perturbation. PLoS Comput. Biol. 8, e1002566 (2012).
Horn, T. et al. Mapping of signaling networks through synthetic genetic interaction analysis by RNAi. Nat. Methods 8, 341–346 (2011).
Wysocka, J. et al. A PHD finger of NURF couples histone H3 lysine 4 trimethylation with chromatin remodelling. Nature 442, 86–90 (2006).
LeRoy, G., Loyola, A., Lane, W. S. & Reinberg, D. Purification and characterization of a human factor that assembles and remodels chromatin. J. Biol. Chem. 275, 14787–14790 (2000).
Bernstein, B. E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 (2006).
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011).
Champagne, K. S. et al. The crystal structure of the ING5 PHD finger in complex with an H3K4me3 histone peptide. Proteins 72, 1371–1376 (2008).
Kouwenhoven, E. N. et al. Genome-wide profiling of p63 DNA-binding sites identifies an element that regulates gene expression during limb development in the 7q21 SHFM1 locus. PLoS Genet. 6, e1001065 (2010).
Li, A. G., Koster, M. I. & Wang, X. J. Roles of TGFβ signaling in epidermal/appendage development. Cytokine Growth Factor Rev. 14, 99–111 (2003).
Ficz, G. et al. Dynamic regulation of 5-hydroxymethylcytosine in mouse ES cells and during differentiation. Nature 473, 398–402 (2011).
Williams, K. et al. TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity. Nature 473, 343–348 (2011).
Truong, A. B., Kretz, M., Ridky, T. W., Kimmel, R. & Khavari, P. A. p63 regulates proliferation and differentiation of developmentally mature keratinocytes. Genes Dev. 20, 3185–3197 (2006).
LeBoeuf, M. et al. Hdac1 and Hdac2 act redundantly to control p63 and p53 functions in epidermal progenitor cells. Dev. Cell 19, 807–818 (2010).
Gandarillas, A. & Watt, F. M. c-Myc promotes differentiation of human epidermal stem cells. Genes Dev. 11, 2869–2882 (1997).
Birmingham, A. et al. Statistical methods for analysis of high-throughput RNA interference screens. Nat. Methods 6, 569–575 (2009).
Reich, M. et al. GenePattern 2.0. Nat. Genet 38, 500–501 (2006).
Pavlidis, P. & Noble, W. S. Matrix2png: a utility for visualizing matrix data. Bioinformatics 19, 295–296 (2003).
Backes, C. et al. GeneTrail–advanced gene set enrichment analysis. Nucleic Acids Res. 35, W186–W192 (2007).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Dempster, A. P., Laird, N. M. & Rubin, D. B. Maximum likelihood from incomplete data via the EM algorithm. J. R. Statist. Soc. B 39, 1–38 (1977).
Jeffreys, H. Theory of Probability 3rd edn, (Oxford Univ. Press, 1998).
Suzuki, R. & Shimodaira, H. Pvclust: an R package for assessing the uncertainty in hierarchical clustering. Bioinformatics 22, 1540–1542 (2006).
Schmidt, D. et al. ChIP-seq: using high-throughput sequencing to discover protein-DNA interactions. Methods 48, 240–248 (2009).
Acknowledgements
We thank A. Brenkman, J. Carroll, N. Rosenfeld, M. Narita, D. Odom and Watt laboratory members for critical discussion and comments. This work was supported by Cancer Research UK (CRUK), Hutchinson Wampoa, University of Cambridge (F.M.W.) and a Marie Curie Fellowship (PIEF-GA-2008-220642) to K.W.M. We are indebted to the core facilities of the CRUK Cambridge Research Institute for excellent technical assistance.
Author information
Authors and Affiliations
Contributions
K.W.M. conceived the study, designed, performed and analysed experiments and performed bioinformatic analysis. X.W. developed the Bayesian mixture model and network analysis. C.E. and G.S. contributed to the skin reconstitution assays. Y.I. performed bisulphite sequencing analysis. R.F.S. performed bioinformatic analysis. J.G. performed mRNA co-expression analysis. G.D. analysed data and contributed to experimental design. S.U-L. performed 5hmeDNA dot-blot analysis. P.P. performed mRNA co-expression analysis. A.M. oversaw the work of Y.I. and S.U-L. F.M. analysed data, performed bioinformatic analysis and contributed computational tools. F.M.W. designed experiments, analysed the data and oversaw the study. K.W.M and F.M.W. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 554 kb)
Supplementary Table 1
Supplementary Information (XLS 371 kb)
Supplementary Table 2
Supplementary Information (XLS 2342 kb)
Supplementary Table 3
Supplementary Information (XLS 32 kb)
Supplementary Dataset 1
Supplementary Information (XLS 136 kb)
Rights and permissions
About this article
Cite this article
Mulder, K., Wang, X., Escriu, C. et al. Diverse epigenetic strategies interact to control epidermal differentiation. Nat Cell Biol 14, 753–763 (2012). https://doi.org/10.1038/ncb2520
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2520
This article is cited by
-
5-hydroxymethylcytosine and gene activity in mouse intestinal differentiation
Scientific Reports (2020)
-
Epidermal stem cells in wound healing and their clinical applications
Stem Cell Research & Therapy (2019)
-
TBL1 is required for the mesenchymal phenotype of transformed breast cancer cells
Cell Death & Disease (2019)
-
Delta-like 1-mediated cis-inhibition of Jagged1/2 signalling inhibits differentiation of human epidermal cells in culture
Scientific Reports (2019)
-
Comprehensive Proteomic Analysis Reveals Intermediate Stage of Non-Lesional Psoriatic Skin and Points out the Importance of Proteins Outside this Trend
Scientific Reports (2019)