Abstract
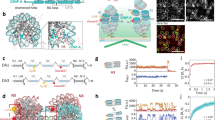
The centromere-specific histone H3 variant CENH3 (also known as CENP-A) is considered to be an epigenetic mark for establishment and propagation of centromere identity. Pulse induction of CENH3 (Drosophila CID) in Schneider S2 cells leads to its incorporation into non-centromeric regions and generates CID islands that resist clearing from chromosome arms for multiple cell generations. We demonstrate that CID islands represent functional ectopic kinetochores, which are non-randomly distributed on the chromosome and show a preferential localization near telomeres and pericentric heterochromatin in transcriptionally silent, intergenic chromatin domains. Although overexpression of heterochromatin protein 1 (HP1) or increasing histone acetylation interferes with CID island formation on a global scale, induction of a locally defined region of synthetic heterochromatin by targeting HP1–LacI fusions to stably integrated Lac operator arrays produces a proximal hotspot for CID deposition. These data indicate that the characteristics of regions bordering heterochromatin promote de novo kinetochore assembly and thereby contribute to centromere identity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Przewloka, M. R. & Glover, D. M. The kinetochore and the centromere: a working long distance relationship. Annu. Rev. Genet. 43, 439–465 (2009).
Welburn, J. P. & Cheeseman, I. M. Toward a molecular structure of the eukaryotic kinetochore. Dev. Cell 15, 645–655 (2008).
Santaguida, S. & Musacchio, A. The life and miracles of kinetochores. EMBO J. 28, 2511–2531 (2009).
Vagnarelli, P., Ribeiro, S. A. & Earnshaw, W. C. Centromeres: old tales and new tools. FEBS Lett. 582, 1950–1959 (2008).
Warburton, P. E. Chromosomal dynamics of human neocentromere formation. Chromosom. Res. 12, 617–626 (2004).
Marshall, O. J., Chueh, A. C., Wong, L. H. & Choo, K. H. Neocentromeres: new insights into centromere structure, disease development, and karyotype evolution. Am. J. Hum. Genet. 82, 261–282 (2008).
Allshire, R. C. & Karpen, G. H. Epigenetic regulation of centromeric chromatin: old dogs, new tricks? Nat. Rev. Genet. 9, 923–937 (2008).
Schuh, M., Lehner, C. F. & Heidmann, S. Incorporation of Drosophila CID/CENP-A and CENP-C into centromeres during early embryonic anaphase. Curr. Biol. 17, 237–243 (2007).
Jansen, L. E., Black, B. E., Foltz, D. R. & Cleveland, D. W. Propagation of centromeric chromatin requires exit from mitosis. J. Cell Biol. 176, 795–805 (2007).
Capozzi, O. et al. Evolutionary and clinical neocentromeres: two faces of the same coin? Chromosoma 117, 339–344 (2008).
Ventura, M. et al. Evolutionary formation of new centromeres in macaque. Science 316, 243–246 (2007).
Williams, B. C., Murphy, T. D., Goldberg, M. L. & Karpen, G. H. Neocentromere activity of structurally acentric mini-chromosomes in Drosophila . Nat. Genet. 18, 30–37 (1998).
Maggert, K. A. & Karpen, G. H. The activation of a neocentromere in Drosophila requires proximity to an endogenous centromere. Genetics 158, 1615–1628 (2001).
Platero, J. S., Ahmad, K. & Henikoff, S. A distal heterochromatic block displays centromeric activity when detached from a natural centromere. Mol. Cell. 4, 995–1004 (1999).
Nasuda, S., Hudakova, S., Schubert, I., Houben, A. & Endo, T. R. Stable barley chromosomes without centromeric repeats. Proc. Natl Acad. Sci. USA 102, 9842–9847 (2005).
Ishii, K. et al. Heterochromatin integrity affects chromosome reorganization after centromere dysfunction. Science 321, 1088–1091 (2008).
Ketel, C. et al. Neocentromeres form efficiently at multiple possible loci in Candida albicans . PLoS Genet. 5, e1000400 (2009).
Marshall, O. J. & Choo, K. H. Neocentromeres come of age. PLoS Genet. 5, e1000370 (2009).
Folco, H. D., Pidoux, A. L., Urano, T. & Allshire, R. C. Heterochromatin and RNAi are required to establish CENP-A chromatin at centromeres. Science 319, 94–97 (2008).
Durand-Dubief, M. & Ekwall, K. Heterochromatin tells CENP-A where to go. Bioessays 30, 526–529 (2008).
Alonso, A., Hasson, D., Cheung, F. & Warburton, P. E. A paucity of heterochromatin at functional human neocentromeres. Epigenetics Chromatin 3, 6 (2010).
Saffery, R. et al. Transcription within a functional human centromere. Mol. Cell. 12, 509–516 (2003).
Tomonaga, T. et al. Overexpression and mistargeting of centromere protein-A in human primary colorectal cancer. Cancer Res. 63, 3511–3516 (2003).
Ahmad, K. & Henikoff, S. Histone H3 variants specify modes of chromatin assembly. Proc. Natl Acad. Sci. USA 99 (Suppl 4), 16477–16484 (2002).
Heun, P. et al. Mislocalization of the Drosophila centromere-specific histone CID promotes formation of functional ectopic kinetochores. Dev. Cell. 10, 303–315 (2006).
Collins, K. A., Furuyama, S. & Biggins, S. Proteolysis contributes to the exclusive centromere localization of the yeast Cse4/CENP-A histone H3 variant. Curr. Biol. 14, 1968–1972 (2004).
Moreno-Moreno, O., Torras-Llort, M. & Azorin, F. Proteolysis restricts localization of CID, the centromere-specific histone H3 variant of Drosophila, to centromeres. Nucleic Acids Res. 34, 6247–6255 (2006).
Heeger, S. et al. Genetic interactions of separase regulatory subunits reveal the diverged Drosophila Cenp-C homolog. Genes Dev. 19, 2041–2053 (2005).
Basto, R., Gomes, R. & Karess, R. E. Rough deal and Zw10 are required for the metaphase checkpoint in Drosophila . Nat. Cell Biol. 2, 939–943 (2000).
Logarinho, E. & Sunkel, C. E. The Drosophila POLO kinase localises to multiple compartments of the mitotic apparatus and is required for the phosphorylation of MPM2 reactive epitopes. J. Cell Sci. 111 (Pt 19), 2897–2909 (1998).
Maia, A. F., Lopes, C. S. & Sunkel, C. E. BubR1 and CENP-E have antagonistic effects on the stability of microtubule-kinetochore attachments in Drosophila S2 cell mitosis. Cell Cycle 6, 1367–1378 (2007).
Zinchuk, V. & Zinchuk, O. Quantitative colocalization analysis of confocal fluorescence microscopy images. Curr. Protoc. Cell. Biol. Chapter 4, Unit 4 19 (2008).
Celniker, S. E. et al. Unlocking the secrets of the genome. Nature 459, 927–930 (2009).
Kharchenko, P. V. et al. Comprehensive analysis of the chromatin landscape in Drosophila melanogaster . Nature (2010).
Roy, S. et al. Identification of functional elements and regulatory circuits by Drosophila modENCODE. Science 330, 1787–1797 (2010).
Alonso, A. et al. Co-localization of CENP-C and CENP-H to discontinuous domains of CENP-A chromatin at human neocentromeres. Genome Biol. 8, R148 (2007).
Ciferri, C. et al. Implications for kinetochore–microtubule attachment from the structure of an engineered Ndc80 complex. Cell 133, 427–439 (2008).
Wei, R. R., Al-Bassam, J. & Harrison, S. C. The Ndc80/HEC1 complex is a contact point for kinetochore–microtubule attachment. Nat. Struct. Mol. Biol. 14, 54–59 (2007).
Cheeseman, I. M., Chappie, J. S., Wilson-Kubalek, E. M. & Desai, A. The conserved KMN network constitutes the core microtubule-binding site of the kinetochore. Cell 127, 983–997 (2006).
Lam, A. L., Boivin, C. D., Bonney, C. F., Rudd, M. K. & Sullivan, B. A. Human centromeric chromatin is a dynamic chromosomal domain that can spread over noncentromeric DNA. Proc. Natl Acad. Sci. USA 103, 4186–4191 (2006).
Bergmann, J. H. et al. Epigenetic engineering shows H3K4me2 is required for HJURP targeting and CENP-A assembly on a synthetic human kinetochore. EMBO J. 30, 328–340 (2011).
Kagansky, A. et al. Synthetic heterochromatin bypasses RNAi and centromeric repeats to establish functional centromeres. Science 324, 1716–1719 (2009).
Verschure, P. J. et al. In vivo HP1 targeting causes large-scale chromatin condensation and enhanced histone lysine methylation. Mol. Cell. Biol. 25, 4552–4564 (2005).
Danzer, J. R. & Wallrath, L. L. Mechanisms of HP1-mediated gene silencing in Drosophila . Development 131, 3571–3580 (2004).
Blower, M. D. & Karpen, G. H. The role of Drosophila CID in kinetochore formation, cell-cycle progression and heterochromatin interactions. Nat. Cell Biol. 3, 730–739 (2001).
Frydrychova, R. C., Mason, J. M. & Archer, T. K. HP1 is distributed within distinct chromatin domains at Drosophila telomeres. Genetics 180, 121–131 (2008).
Riddle, N. C. et al. Plasticity in patterns of histone modifications and chromosomal proteins in Drosophila heterochromatin. Genome Res. 21, 147–163 (2011).
Ebert, A., Lein, S., Schotta, G. & Reuter, G. Histone modification and the control of heterochromatic gene silencing in Drosophila . Chromosom. Res. 14, 377–392 (2006).
Villasante, A., Abad, J. P. & Mendez-Lago, M. Centromeres were derived from telomeres during the evolution of the eukaryotic chromosome. Proc. Natl Acad. Sci. USA 104, 10542–10547 (2007).
Losada, A., Agudo, M., Abad, J. P. & Villasante, A. HeT-A telomere-specific retrotransposons in the centric heterochromatin of Drosophila melanogaster chromosome 3. Mol. Gen. Genet. 262, 618–622 (1999).
Mendez-Lago, M. et al. Novel sequencing strategy for repetitive DNA in a Drosophila BAC clone reveals that the centromeric region of the Y chromosome evolved from a telomere. Nucleic Acids Res. 37, 2264–2273 (2009).
Nakano, M. et al. Inactivation of a human kinetochore by specific targeting of chromatin modifiers. Dev. Cell 14, 507–522 (2008).
Cardinale, S. et al. Hierarchical inactivation of a synthetic human kinetochore by a chromatin modifier. Mol. Biol. Cell. 20, 4194–4204 (2009).
Sullivan, B. A. & Karpen, G. H. Centromeric chromatin exhibits a histone modification pattern that is distinct from both euchromatin and heterochromatin. Nat. Struct. Mol. Biol. 11, 1076–1083 (2004).
Chueh, A. C., Northrop, E. L., Brettingham-Moore, K. H., Choo, K. H. & Wong, L. H. LINE retrotransposon RNA is an essential structural and functional epigenetic component of a core neocentromeric chromatin. PLoS Genet. 5, e1000354 (2009).
Wong, N. C. et al. Permissive transcriptional activity at the centromere through pockets of DNA hypomethylation. PLoS Genet. 2, e17 (2006).
Obuse, C. et al. A conserved Mis12 centromere complex is linked to heterochromatic HP1 and outer kinetochore protein Zwint-1. Nat. Cell Biol. 6, 1135–1141 (2004).
Cheeseman, I. M. & Desai, A. Molecular architecture of the kinetochore–microtubule interface. Nat. Rev. Mol. Cell. Biol. 9, 33–46 (2008).
de Wit, E., Greil, F. & van Steensel, B. Genome-wide HP1 binding in Drosophila: developmental plasticity and genomic targeting signals. Genome Res. 15, 1265–1273 (2005).
Straight, A. F., Belmont, A. S., Robinett, C. C. & Murray, A. W. GFP tagging of budding yeast chromosomes reveals that protein–protein interactions can mediate sister chromatid cohesion. Curr. Biol. 6, 1599–1608 (1996).
Pereira, A. J., Matos, I., Lince-Faria, M. & Maiato, H. Dissecting mitosis with laser microsurgery and RNAi in Drosophila cells. Methods Mol. Biol. 545, 145–164 (2009).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Jurka, J. Repbase update: a database and an electronic journal of repetitive elements. Trends Genet. 16, 418–420 (2000).
Audic, S. & Claverie, J. M. The significance of digital gene expression profiles. Genome Res. 7, 986–995 (1997).
Acknowledgements
We thank the following people for antibodies: T. Maresca and E. Salmon, R. Karess, C. Sunkel, S. Heidmann, C. Lehner and G. H. Karpen and R. Tsien for the mRFP construct. We would also like to thank B. Mellone, P. Pasero, B. Duncker and the laboratory members, particularly M-J. Mendiburo and J. Padeken for reagents, discussions and critical reading of the manuscript, and H-J. Schwarz for general assistance. Work in the laboratory of H.M. is funded by grants PTDC/SAU-GMG/099704/2008 and PTDC/SAU-ONC/112917/2009 from Fundação para a Ciência e a Tecnologia of Portugal (COMPETE-FEDER), the Human Frontier Research Program and the Seventh Framework Programme grant PRECISE from the European Research Council. Funding for this research project of the laboratory of P.H. was provided by the Max Planck Society, the Human Frontier Science Program (CDA) and by the Excellence Initiative of the German Federal and State Governments (EXC 294).
Author information
Authors and Affiliations
Contributions
A.M.O. carried out the cytological characterization of CID islands. A.J.P., A.M.O. and H.M. carried out laser-microsurgery experiments. D.v.E. and S.S. conducted the ChIP-seq and qPCR. T.M. and S.D. carried out bioinformatic analysis of ChIP-seq. A.M.O. and P.H. did the project planning and data analysis. P.H. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1749 kb)
Rights and permissions
About this article
Cite this article
Olszak, A., van Essen, D., Pereira, A. et al. Heterochromatin boundaries are hotspots for de novo kinetochore formation. Nat Cell Biol 13, 799–808 (2011). https://doi.org/10.1038/ncb2272
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2272
This article is cited by
-
Incorporation of CENP-A/CID into centromeres during early Drosophila embryogenesis does not require RNA polymerase II–mediated transcription
Chromosoma (2022)
-
Genetic screening identifies a SUMO protease dynamically maintaining centromeric chromatin
Nature Communications (2020)
-
Genetic and epigenetic effects on centromere establishment
Chromosoma (2020)
-
The dark side of centromeres: types, causes and consequences of structural abnormalities implicating centromeric DNA
Nature Communications (2018)
-
Segmental Duplication of Chromosome 11 and its Implications for Cell Division and Genome-wide Expression in Rice
Scientific Reports (2017)