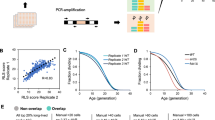
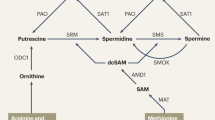
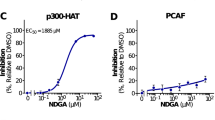
Abstract
Ageing results from complex genetically and epigenetically programmed processes that are elicited in part by noxious or stressful events that cause programmed cell death. Here, we report that administration of spermidine, a natural polyamine whose intracellular concentration declines during human ageing, markedly extended the lifespan of yeast, flies and worms, and human immune cells. In addition, spermidine administration potently inhibited oxidative stress in ageing mice. In ageing yeast, spermidine treatment triggered epigenetic deacetylation of histone H3 through inhibition of histone acetyltransferases (HAT), suppressing oxidative stress and necrosis. Conversely, depletion of endogenous polyamines led to hyperacetylation, generation of reactive oxygen species, early necrotic death and decreased lifespan. The altered acetylation status of the chromatin led to significant upregulation of various autophagy-related transcripts, triggering autophagy in yeast, flies, worms and human cells. Finally, we found that enhanced autophagy is crucial for polyamine-induced suppression of necrosis and enhanced longevity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Fabrizio, P. et al. Superoxide is a mediator of an altruistic aging program in Saccharomyces cerevisiae. J. Cell Biol. 166, 1055–1067 (2004).
Herker, E. et al. Chronological aging leads to apoptosis in yeast. J. Cell Biol. 164, 501–507 (2004).
Vicencio, J. M. et al. Senescence, apoptosis or autophagy? When a damaged cell must decide its path—a mini-review. Gerontology 54, 92–99 (2008).
Golstein, P. & Kroemer, G. Cell death by necrosis: towards a molecular definition. Trends Biochem. Sci. 32, 37–43 (2007).
Zong, W. X. & Thompson, C. B. Necrotic death as a cell fate. Genes Dev. 20, 1–15 (2006).
Imai, S., Armstrong, C. M., Kaeberlein, M. & Guarente, L. Transcriptional silencing and longevity protein Sir2 is an NAD-dependent histone deacetylase. Nature 403, 795–800 (2000).
Lin, S. J., Defossez, P. A. & Guarente, L. Requirement of NAD and SIR2 for life-span extension by calorie restriction in Saccharomyces cerevisiae. Science 289, 2126–2128 (2000).
Lamming, D. W. et al. HST2 mediates SIR2-independent life-span extension by calorie restriction. Science 309, 1861–1864 (2005).
Jenuwein, T. & Allis, C. D. Translating the histone code. Science 293, 1074–1080 (2001).
Kaeberlein, M., Burtner, C. R. & Kennedy, B. K. Recent developments in yeast aging. PLoS Genet. 3, e84 (2007).
Longo, V. D. & Kennedy, B. K. Sirtuins in aging and age-related disease. Cell 126, 257–268 (2006).
Dang, W. et al. Histone H4 lysine 16 acetylation regulates cellular lifespan. Nature 459, 802–807 (2009).
Longo, V. D. & Finch, C. E. Evolutionary medicine: from dwarf model systems to healthy centenarians? Science 299, 1342–1346 (2003).
Powers, R. W., 3rd, Kaeberlein, M., Caldwell, S. D., Kennedy, B. K. & Fields, S. Extension of chronological life span in yeast by decreased TOR pathway signaling. Genes Dev. 20, 174–184 (2006).
Bonawitz, N. D., Chatenay-Lapointe, M., Pan, Y. & Shadel, G. S. Reduced TOR signaling extends chronological life span via increased respiration and upregulation of mitochondrial gene expression. Cell Metab. 5, 265–277 (2007).
Harrison, D. E. et al. Rapamycin fed late in life extends lifespan in genetically heterogeneous mice. Nature 460, 392–395 (2009).
Galluzzi, L. et al. To die or not to die: that is the autophagic question. Curr. Mol. Med. 8, 78–91 (2008).
Jia, K. & Levine, B. Autophagy is required for dietary restriction-mediated life span extension in C. elegans. Autophagy 3, 597–599 (2007).
Scalabrino, G. & Ferioli, M. E. Polyamines in mammalian ageing: an oncological problem, too? A review. Mech. Ageing Dev. 26, 149–164 (1984).
Weinberger, M. et al. Apoptosis in budding yeast caused by defects in initiation of DNA replication. J. Cell Sci. 118, 3543–3553 (2005).
Allen, C. et al. Isolation of quiescent and nonquiescent cells from yeast stationary-phase cultures. J. Cell Biol. 174, 89–100 (2006).
Laun, P. et al. Aged mother cells of Saccharomyces cerevisiae show markers of oxidative stress and apoptosis. Mol. Microbiol. 39, 1166–1173 (2001).
Fabrizio, P., Pozza, F., Pletcher, S. D., Gendron, C. M. & Longo, V. D. Regulation of longevity and stress resistance by Sch9 in yeast. Science 292, 288–290 (2001).
Harman, D. Aging: a theory based on free radical and radiation chemistry. J. Gerontol. 11, 298–300 (1956).
Schraml, E. et al. Norepinephrine treatment and aging lead to systemic and intracellular oxidative stress in rats. Exp. Gerontol. 42, 1072–1078 (2007).
Artal-Sanz, M., Samara, C., Syntichaki, P. & Tavernarakis, N. Lysosomal biogenesis and function is critical for necrotic cell death in Caenorhabditis elegans. J. Cell Biol. 173, 231–239 (2006).
Syntichaki, P., Samara, C. & Tavernarakis, N. The vacuolar H+-ATPase mediates intracellular acidification required for neurodegeneration in C. elegans. Curr. Biol. 15, 1249–1254 (2005).
Burtner, C. R., Murakami, C. J., Kennedy, B. K. & Kaeberlein, M. A molecular mechanism of chronological aging in yeast. Cell Cycle 8, 1256–1270 (2009).
Fabrizio, P. et al. Sir2 blocks extreme life-span extension. Cell 123, 655–667 (2005).
Burhans, W. C. & Weinberger, M. Acetic acid effects on aging in budding yeast: Are they relevant to aging in higher eukaryotes? Cell Cycle 8 (2009).
Scaffidi, P., Misteli, T. & Bianchi, M. E. Release of chromatin protein HMGB1 by necrotic cells triggers inflammation. Nature 418, 191–195 (2002).
Weinberger, M. et al. DNA replication stress is a determinant of chronological lifespan in budding yeast. PLoS ONE 2, e748 (2007).
Duncan, E. M. et al. Cathepsin L proteolytically processes histone H3 during mouse embryonic stem cell differentiation. Cell 135, 284–294 (2008).
Howe, L. et al. Histone H3 specific acetyltransferases are essential for cell cycle progression. Genes Dev. 15, 3144–3154 (2001).
Winkler, G. S., Kristjuhan, A., Erdjument-Bromage, H., Tempst, P. & Svejstrup, J. Q. Elongator is a histone H3 and H4 acetyltransferase important for normal histone acetylation levels in vivo. Proc. Natl Acad. Sci. USA 99, 3517–3522 (2002).
Hobbs, C. A., Paul, B. A. & Gilmour, S. K. Elevated levels of polyamines alter chromatin in murine skin and tumors without global changes in nucleosome acetylation. Exp. Cell Res. 290, 427–436 (2003).
Hobbs, C. A., Paul, B. A. & Gilmour, S. K. Deregulation of polyamine biosynthesis alters intrinsic histone acetyltransferase and deacetylase activities in murine skin and tumors. Cancer Res. 62, 67–74 (2002).
Liu, B., Sutton, A. & Sternglanz, R. A yeast polyamine acetyltransferase. J. Biol. Chem. 280, 16659–16664 (2005).
Juhasz, G., Erdi, B., Sass, M. & Neufeld, T. P. Atg7-dependent autophagy promotes neuronal health, stress tolerance, and longevity but is dispensable for metamorphosis in Drosophila. Genes Dev. 21, 3061–3066 (2007).
Tavernarakis, N., Pasparaki, A., Tasdemir, E., Maiuri, M. C. & Kroemer, G. The effects of p53 on whole organism longevity are mediated by autophagy. Autophagy 4, 870–873 (2008).
Melendez, A. et al. Autophagy genes are essential for dauer development and life-span extension in C. elegans. Science 301, 1387–1391 (2003).
Orvedahl, A. & Levine, B. Eating the enemy within: autophagy in infectious diseases. Cell Death Differ. 16, 57-69 (2009).
Hoyer-Hansen, M. & Jaattela, M. Autophagy: an emerging target for cancer therapy. Autophagy 4, 574–580 (2008).
Lambert, L. A. et al. Autophagy: a novel mechanism of synergistic cytotoxicity between doxorubicin and roscovitine in a sarcoma model. Cancer Res. 68, 7966–7974 (2008).
Lefranc, F., Facchini, V. & Kiss, R. Proautophagic drugs: a novel means to combat apoptosis-resistant cancers, with a special emphasis on glioblastomas. Oncologist 12, 1395–1403 (2007).
Franceschi, C. et al. Inflammaging and anti-inflammaging: a systemic perspective on aging and longevity emerged from studies in humans. Mech. Ageing Dev. 128, 92–105 (2007).
Levine, B. & Kroemer, G. Autophagy in the pathogenesis of disease. Cell 132, 27–42 (2008).
Balasundaram, D., Tabor, C. W. & Tabor, H. Spermidine or spermine is essential for the aerobic growth of Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 88, 5872–5876 (1991).
Gueldener, U., Heinisch, J., Koehler, G. J., Voss, D. & Hegemann, J. H. A second set of loxP marker cassettes for Cre-mediated multiple gene knockouts in budding yeast. Nucleic Acids Res. 30, e23 (2002).
Buttner, S. et al. Endonuclease G regulates budding yeast life and death. Mol. Cell 25, 233–246 (2007).
Lotze, M. T. & Tracey, K. J. High-mobility group Box 1 protein (HMGB1): nuclear weapon in the immune arsenal. Nature Rev. Immunol. 5, 331–342 (2005).
Kirisako, T. et al. Formation process of autophagosome is traced with Apg8/Aut7p in yeast. J. Cell Biol. 147, 435–446 (1999).
Noda, T., Matsuura, A., Wada, Y. & Ohsumi, Y. Novel system for monitoring autophagy in the yeast Saccharomyces cerevisiae. Biochem. Biophys. Res. Commun. 210, 126–132 (1995).
Vermes, I., Haanen, C., Steffens-Nakken, H. & Reutelingsperger, C. A novel assay for apoptosis. Flow cytometric detection of phosphatidylserine expression on early apoptotic cells using fluorescein labelled annexin V. J. Immunol. Methods 184, 39–51 (1995).
Criollo, A. et al. Regulation of autophagy by the inositol trisphosphate receptor. Cell Death Differ. 14, 1029–1039 (2007).
Ellman, G. L. Tissue sulfhydryl groups. Arch. Biochem. Biophys. 82, 70–77 (1959).
Riener, C. K., Kada, G. & Gruber, H. J. Quick measurement of protein sulfhydryls with Ellman's reagent and with 4, 4′-dithiodipyridine. Anal. Bioanal. Chem. 373, 266–276 (2002).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Samara, C., Syntichaki, P. & Tavernarakis, N. Autophagy is required for necrotic cell death in Caenorhabditis elegans. Cell Death Differ. 15, 105–112 (2008).
Minocha, R., Shortle, W. C., Long, S. L. & Minocha, S. C. A rapid and reliable procedure for extraction of cellular polyamines and inorganic ions from plant tissues. J. Plant Growth Regul. 13, 187–193 (1994).
Gianotti, V. et al. A new hydrophilic interaction liquid chromatography tandem mass spectrometry method for the simultaneous determination of seven biogenic amines in cheese. J. Chromatogr. A 1185, 296–300 (2008).
Fahrenkrog, B., Hurt, E. C., Aebi, U. & Pante, N. Molecular architecture of the yeast nuclear pore complex: localization of Nsp1p subcomplexes. J. Cell Biol. 143, 577–588 (1998).
Wu, Z., Irizarry, R. A., Gentleman, R., Martinez-Murillo, F. & Spencer, F. A model-based background adjustment for oligonucleotide expression arrays. J. Am. Stat. Assoc. 99, 909–918 (2004).
Rainer, J., Sanchez-Cabo, F., Stocker, G., Sturn, A. & Trajanoski, Z. CARMAweb: comprehensive R- and bioconductor-based web service for microarray data analysis. Nucleic Acids Res. 34, W498–W503 (2006).
Saeed, A. I. et al. TM4: a free, open-source system for microarray data management and analysis. Biotechniques 34, 374–378 (2003).
Rozen, S. & Skaletsky, H. Primer3 on the WWW for general users and for biologist programmers. Methods Mol. Biol. 132, 365–386 (2000).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-δ δ C(T)) method. Methods 25, 402–408 (2001).
Strahl-Bolsinger, S., Hecht, A., Luo, K. & Grunstein, M. SIR2 and SIR4 interactions differ in core and extended telomeric heterochromatin in yeast. Genes Dev. 11, 83–93 (1997).
Evans, E., Sugawara, N., Haber, J. E. & Alani, E. The Saccharomyces cerevisiae Msh2 mismatch repair protein localizes to recombination intermediates in vivo. Mol. Cell 5, 789–799 (2000).
Acknowledgements
We thank Ulrike Potocnik, Silvia Dichtinger, Arno Absenger and Elfgard Heintz for assistance. We are grateful to the Austrian Science Fund FWF (Austria) for grant S-9304-B05 (to F.M., S.B. and D.C.-G.), grant S-9303-B05 (to K.-U.F. and H.K.), grant LIPOTOX (to F.M. and S.B.), and grant S-9301-B05 (to B.G.-L.) and to the European Commission for project TransDeath (to K.-U.F. and C.R.), project APOSYS (to F.M., T.E. and G.K.) and project Lifespan (FP6 036894 to B.G.-L.). We are grateful to the Austrian Science Fund FWF (Vienna, Austria) for grants S9302-B05 (to M.B.) and to the European Commission (Brussels, Europe) for project MIMAGE (contract no. 512020; to M.B.).
Author information
Authors and Affiliations
Contributions
T.E., G.K. and F.M. designed and organized this study and wrote the manuscript. T.E. performed the largest part of the yeast and mouse experiments and contributed significantly to the fly and PBMC cell culture data. H.K., A.S., S.B., C.R., D.C-G., J.R., S.S., H.F., L.A. and B.A. contributed to the yeast experiments. C.M. and F.S. performed polyamine measurements by MassSpec. L.D., R.H. and N.M. performed the fly experiments. E.S. contributed to mouse data. A.C. performed HELA cell culture experiments. E.M. and N.T. contributed the worm data. D.W. and B.G-L. performed PBMC experiments. P.L., G.H. and M.B. determined yeast replicative life spans. N.M., E.H., K.-U.F., S.B., C.R. and D.C-G. contributed with decisive discussions and helped to design experiments.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 2045 kb)
Rights and permissions
About this article
Cite this article
Eisenberg, T., Knauer, H., Schauer, A. et al. Induction of autophagy by spermidine promotes longevity. Nat Cell Biol 11, 1305–1314 (2009). https://doi.org/10.1038/ncb1975
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1975
This article is cited by
-
Toward nutrition improving outcome of critically ill patients: How to interpret recent feeding RCTs?
Critical Care (2023)
-
The effect of spermine on Tetranychus urticae-Cucumis sativus interaction
BMC Plant Biology (2023)
-
Polyamines: their significance for maintaining health and contributing to diseases
Cell Communication and Signaling (2023)
-
Spermidine promotes fertility in aged female mice
Nature Aging (2023)
-
Spermidine improves angiogenic capacity of senescent endothelial cells, and enhances ischemia-induced neovascularization in aged mice
Scientific Reports (2023)