Abstract
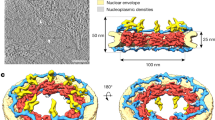
Nucleocytoplasmic transport occurs through nuclear pore complexes (NPCs) embedded in the nuclear envelope1. Here, we discovered an unexpected role for yeast dynein light chain (Dyn2)2 in the NPC. Dyn2 is a previously undescribed nucleoporin that functions as molecular glue to dimerize and stabilize the Nup82–Nsp1–Nup159 complex, a module of the cytoplasmic pore filaments3. Biochemical analyses showed that Dyn2 binds to a linear motif (termed DIDNup159) inserted between the Phe-Gly repeat and coiled-coil domain of Nup159. Electron microscopy revealed that the reconstituted Dyn2–DIDNup159 complex forms a rigid rod-like structure, in which five Dyn2 homodimers align like 'pearls on a string' between two extented DIDNup159 strands. These findings imply that the rigid 20 nm long Dyn2–DIDNup159 filament projects the Nup159 Phe-Gly repeats from the Nup82 module. Thus, it is possible that dynein light chain plays a role in organizing natively unfolded Phe-Gly repeats within the NPC scaffold to facilitate nucleocytoplasmic transport.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Hetzer, M. W., Walther, T. C. & Mattaj, I. W. Pushing the envelope: structure, function, and dynamics of the nuclear periphery. Annu. Rev. Cell. Dev. Biol. 21, 347–380 (2005).
Dick, T., Surana, U. & Chia, W. Molecular and genetic characterization of SLC1, a putative Saccharomyces cerevisiae homolog of the metazoan cytoplasmic dynein light chain 1. Mol. Gen. Genet. 251, 38–43 (1996).
Cole, C. N. & Scarcelli, J. J. Transport of messenger RNA from the nucleus to the cytoplasm. Curr. Opin. Cell Biol. 18, 299–306 (2006).
Schwartz, T. U. Modularity within the architecture of the nuclear pore complex. Curr. Opin. Struct. Biol. 15, 221–226 (2005).
Beck, M. et al. Nuclear pore complex structure and dynamics revealed by cryoelectron tomography. Science 306, 1387–1390 (2004).
Rout, M. P. et al. The yeast nuclear pore complex: composition, architecture, and transport mechanism. J. Cell Biol. 148, 635–651 (2000).
Hurwitz, M. E., Strambio-de-Castillia, C. & Blobel, G. Two yeast nuclear pore complex proteins involved in mRNA export form a cytoplasmically oriented subcomplex. Proc. Natl Acad. Sci. USA 95, 11241–11245 (1998).
Belgareh, N. et al. Functional characterization of a Nup159p-containing nuclear pore subcomplex. Mol. Biol. Cell. 9, 3475–3492 (1998).
Weirich, C. S., Erzberger, J. P., Berger, J. M. & Weis, K. The N-terminal domain of Nup159 forms a β-propeller that functions in mRNA export by tethering the helicase Dbp5 to the nuclear pore. Mol. Cell 16, 749–760 (2004).
Denning, D. P., Patel, S. S., Uversky, V., Fink, A. L. & Rexach, M. Disorder in the nuclear pore complex: the FG repeat regions of nucleoporins are natively unfolded. Proc. Natl Acad. Sci. USA 100, 2450–2455 (2003).
Vallee, R. B., Williams, J. C., Varma, D. & Barnhart, L. E. Dynein: An ancient motor protein involved in multiple modes of transport. J. Neurobiol. 58, 189–200 (2004).
Fan, J. S. et al. Protein inhibitor of neuronal nitric-oxide synthase, PIN, binds to a 17-amino acid residue fragment of the enzyme. J. Biol. Chem. 273, 33472–33481 (1998).
Navarro-Lerida, I. et al. Proteomic identification of brain proteins that interact with dynein light chain LC8. Proteomics 4, 339–346 (2004).
Liang, J., Jaffrey, S. R., Guo, W., Snyder, S. H. & Clardy, J. Structure of the PIN/LC8 dimer with a bound peptide. Nature Struct. Biol. 6, 735–740 (1999).
Fan, J., Zhang, Q., Tochio, H., Li, M. & Zhang, M. Structural basis of diverse sequence-dependent target recognition by the 8 kDa dynein light chain. J. Mol. Biol. 306, 97–108 (2001).
Hodge, C. A., Colot, H. V., Stafford, P. & Cole, C. N. Rat8p/Dbp5p is a shuttling transport factor that interacts with Rat7p/Nup159p and Gle1p and suppresses the mRNA export defect of xpo1-1 cells. EMBO J. 18, 5778–5788 (1999).
Komori, M. et al. The Hansenula polymorpha PEX14 gene encodes a novel peroxisomal membrane protein essential for peroxisome biogenesis. EMBO J. 16, 44–53 (1997).
Doye, V., Wepf, R. & Hurt, E. C. A novel nuclear pore protein Nup133p with distinct roles in poly (A)+ RNA transport and nuclear pore distribution. EMBO J. 13, 6062–6075 (1994).
Sheeman, B. et al. Determinants of S. cerevisiae dynein localization and activation: implications for the mechanism of spindle positioning. Curr. Biol. 13, 364–372 (2003).
Nyarko, A. et al. Ionization of His 55 at the dimer interface of dynein light-chain LC8 is coupled to dimer dissociation. Biochemistry 44, 14248–14255 (2005).
Bailer, S. M., Balduf, C. & Hurt, E. C. The Nsp1p carboxy-terminal domain is organized in functionally distinct coiled-coil regions required for assembly of nucleoporin subcomplexes and nucleocytoplasmic transport. Mol. Cell Biol. 21, 7944–7955 (2001).
Grandi, P. et al. A novel nuclear pore protein Nup82p which specifically binds to a fraction of Nsp1p. J. Cell Biol. 130, 1263–1273 (1995).
Gorsch, L. C., Dockendorff, T. C. & Cole, C. N. A conditional allele of the novel repeat-containing yeast nucleoporin RAT7/NUP159 causes both rapid cessation of mRNA export and reversible clustering of nuclear pore complexes. J. Cell Biol. 129, 939–955 (1995).
Miki, F. et al. The 14-kDa dynein light chain-family protein Dlc1 is required for regular oscillatory nuclear movement and efficient recombination during meiotic prophase in fission yeast. Mol. Biol. Cell. 13, 930–946 (2002).
Stewart, M. Molecular mechanism of the nuclear protein import cycle. Nature Rev. Mol. Cell. Biol. 8, 195–208 (2007).
Bernad, R., van der Velde, H., Fornerod, M. & Pickersgill, H. Nup358/RanBP2 attaches to the nuclear pore complex via association with Nup88 and Nup214/CAN and plays a supporting role in CRM1-mediated nuclear protein export. Mol. Cell Biol. 24, 2373–2384 (2004).
Walther, T. C. et al. The cytoplasmic filaments of the nuclear pore complex are dispensable for selective nuclear protein import. J. Cell Biol. 158, 63–77 (2002).
Delphin, C., Guan, T., Melchior, F. & Gerace, L. RanGTP targets p97 to RanBP2, a filamentous protein localized at the cytoplasmic periphery of the nuclear pore complex. Mol. Biol. Cell 8, 2379–2390 (1997).
Puig, O. et al. New constructs and strategies for efficient PCR-based gene manipulations in yeast. Yeast 14, 1139–1146 (1998).
Baßler, J. et al. Identification of a 60S pre-ribosomal particle that is closely linked to nuclear export. Mol. Cell 8, 517–529 (2001).
Lutzmann, M., Kunze, R., Buerer, A., Aebi, U. & Hurt, E. Modular self-assembly of a Y-shaped multiprotein complex from seven nucleoporins. EMBO J. 21, 387–397 (2002).
Lutzmann, M. et al. Reconstitution of Nup157 and Nup145N into the Nup84 complex. J. Biol. Chem. 280, 18442–18451 (2005).
Dube, P., Tavares, P., Lurz, R. & van Heel, M. The portal protein of bacteriophage SPP1: a DNA pump with 13-fold symmetry. EMBO J. 12, 1303–1309 (1993).
van Heel, M., Harauz, G., Orlova, E. V., Schmidt, R. & Schatz, M. A new generation of the IMAGIC image processing system. J. Struct. Biol. 116, 17–24 (1996).
Pettersen, E. F. et al. UCSF chimera —a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Acknowledgements
We are grateful to S. Merker, P. Ihrig and J. Lechner for performing mass spectrometry. E.H. is recipient of grants from the Deutsche Forschungsgemeinschaft (SFB 638/B2) and Fonds der Chemischen Industrie.
Author information
Authors and Affiliations
Contributions
Experiments were designed and data analysed and interpreted by P.S. and E.H. Strain construction, DNA recombination work, fluorescence microscopy and biochemical analyses (affinity purification, gel filtration, in vitro assays) were performed by P.S. and R.K. D.H. and P.P. performed double-flourescence microscopy. Negative-staining electron microscopy was conducted by D.F., M.D. and B.M. The manuscript was written by P.S. and E.H. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary figures S1, S2, S3, S4, S5, S6 and Supplementary table S1 (PDF 1204 kb)
Rights and permissions
About this article
Cite this article
Stelter, P., Kunze, R., Flemming, D. et al. Molecular basis for the functional interaction of dynein light chain with the nuclear-pore complex. Nat Cell Biol 9, 788–796 (2007). https://doi.org/10.1038/ncb1604
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1604
This article is cited by
-
In-cell architecture of the nuclear pore and snapshots of its turnover
Nature (2020)
-
Linker Nups connect the nuclear pore complex inner ring with the outer ring and transport channel
Nature Structural & Molecular Biology (2015)
-
Peroxisomes take shape
Nature Reviews Molecular Cell Biology (2013)
-
From networks of protein interactions to networks of functional dependencies
BMC Systems Biology (2012)
-
Coupling viruses to dynein and kinesin-1
The EMBO Journal (2011)