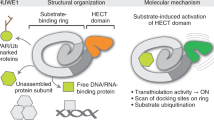
Abstract
Cells have quality-control mechanisms to recognize non-native protein structures and either help the proteins fold or promote their degradation1,2. Ubiquitin-conjugating enzymes (E2s) and ubiquitin ligases (E3s) work together to assemble polyubiquitin chains on misfolded or misassembled proteins, which are then degraded by the proteasome3,4. Here, we find that Ubc7, a yeast E2, can itself undergo degradation when its levels exceed that of its binding partner Cue1, a transmembrane protein that tethers Ubc7 to the endoplasmic reticulum5,6. Unassembled, and thus mislocalized, Ubc7 is targeted to the proteasome by Ufd4, a homologous to E6-AP C-terminus (HECT)-class E3. Ubc7 is autoubiquitinated by a novel mechanism wherein the catalytic cysteine, instead of a lysine residue, provides the polyubiquitin chain acceptor site, and this cysteine-linked chain functions as a degradation signal. The polyubiquitin chain can also be transferred to a lysine side chain, suggesting a mechanism for polyubiquitin chain assembly that precedes substrate modification.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wickner, S., Maurizi, M. R. & Gottesman, S. Posttranslational quality control: folding, refolding, and degrading proteins. Science 286, 1888–1893 (1999).
Meusser, B., Hirsch, C., Jarosch, E. & Sommer, T. ERAD: the long road to destruction. Nature Cell Biol 7, 766–772 (2005).
Hochstrasser, M. Ubiquitin-dependent protein degradation. Annu. Rev. Genetics 30, 405–439 (1996).
Pickart, C. M. Mechanisms underlying ubiquitination. Annu. Rev. Biochem. 70, 503–533 (2001).
Biederer, T., Volkwein, C. & Sommer, T. Role of Cue1p in ubiquitination and degradation at the ER surface. Science 278, 1806–1809 (1997).
Gardner, R. G., Shearer, A. G. & Hampton, R. Y. In vivo action of the HRD ubiquitin ligase complex: mechanisms of endoplasmic reticulum quality control and sterol regulation. Mol. Cell Biol. 21, 4276–4291 (2001).
Biederer, T., Volkwein, C. & Sommer, T. Degradation of subunits of the Sec61p complex, an integral component of the ER membrane, by the ubiquitin-proteasome pathway. EMBO J. 15, 2069–2076 (1996).
Laney, J. D., Mobley, E. F. & Hochstrasser, M. The short-lived Matα2 transcriptional repressor is protected from degradation in vivo by interactions with its corepressors Tup1 and Ssn6. Mol. Cell Biol. 26, 371–380 (2006).
Friedlander, R., Jarosch, E., Urban, J., Volkwein, C. & Sommer, T. A regulatory link between ER-associated protein degradation and the unfolded-protein response. Nature Cell Biol. 2, 379–384 (2000).
Johnson, E. S., Ma, P. C. M., Ota, I. M. & Varshavsky, A. A proteolytic pathway that recognizes ubiquitin as a degradation signal. J. Biol. Chem. 270, 17442–17456 (1995).
Xie, Y. & Varshavsky, A. UFD4 lacking the proteasome-binding region catalyses ubiquitination but is impaired in proteolysis. Nature Cell Biol. 4, 1003–1007 (2002).
Walter, J., Urban, J., Volkwein, C. & Sommer, T. Sec61p-independent degradation of the tail-anchored ER membrane protein Ubc6p. EMBO J. 20, 3124–3131 (2001).
Rape, M. & Kirschner, M. W. Autonomous regulation of the anaphase-promoting complex couples mitosis to S-phase entry. Nature 432, 588–595 (2004).
Yamanaka, A. et al. Cell cycle-dependent expression of mammalian E2-C regulated by the anaphase-promoting complex/cyclosome. Mol. Biol. Cell 11, 2821–2831 (2000).
Machida, Y. J. et al. UBE2T is the E2 in the Fanconi anemia pathway and undergoes negative autoregulation. Mol. Cell 23, 589–596 (2006).
Wu, P. Y. et al. A conserved catalytic residue in the ubiquitin-conjugating enzyme family. EMBO J. 22, 5241–5250 (2003).
Cook, W. J., Martin, P. D., Edwards, B. F., Yamazaki, R. K. & Chau, V. Crystal structure of a class I ubiquitin conjugating enzyme (Ubc7) from Saccharomyces cerevisiae at 2.9 angstroms resolution. Biochem 36, 1621–1627 (1997).
Swanson, R., Locher, M. & Hochstrasser, M. A conserved ubiquitin ligase of the nuclear envelope/endoplasmic reticulum that functions in both ER-associated and Matα2 repressor degradation. Genes Dev. 15, 2660–2674 (2001).
Ciechanover, A. & Ben-Saadon, R. N-terminal ubiquitination: more protein substrates join in. Trends Cell Biol. 14, 103–106 (2004).
Coulombe, P., Rodier, G., Bonneil, E., Thibault, P. & Meloche, S. N-Terminal ubiquitination of extracellular signal-regulated kinase 3 and p21 directs their degradation by the proteasome. Mol. Cell Biol. 24, 6140–6150 (2004).
Hodgins, R. R. W., Ellison, K. S. & Ellison, M. J. Expression of a ubiquitin derivative that conjugates to protein irreversibly produces phenotypes consistent with a ubiquitin deficiency. J. Biol. Chem. 267, 8807–8812 (1992).
Hodgins, R., Gwozd, C., Arnason, T., Cummings, M. & Ellison, M. J. The tail of a ubiquitin-conjugating enzyme redirects multi-ubiquitin chain synthesis from the lysine 48-linked configuration to a novel nonlysine-linked form. J. Biol. Chem. 271, 28766–28771 (1996).
Lin, Y., Hwang, W. C. & Basavappa, R. Structural and functional analysis of the human mitotic-specific ubiquitin-conjugating enzyme, UbcH10. J. Biol. Chem. 277, 21913–21921 (2002).
Cadwell, K. & Coscoy, L. Ubiquitination on nonlysine residues by a viral E3 ubiquitin ligase. Science 309, 127–130 (2005).
Haldeman, M. T., Xia, G., Kasperek, E. M. & Pickart, C. M. Structure and function of ubiquitin conjugating enzyme E2-25K: the tail is a core-dependent activity element. Biochem. 36, 10526–10537 (1997).
Hochstrasser, M. Lingering mysteries of ubiquitin-chain assembly. Cell 124, 27–34 (2006).
Chen, P., Johnson, P., Sommer, T., Jentsch, S. & Hochstrasser, M. Multiple ubiquitin-conjugating enzymes participate in the in vivo degradation of the yeast MATα2 repressor. Cell 74, 357–369 (1993).
Ravid, T., Kreft, S. G. & Hochstrasser, M. Membrane and soluble substrates of the Doa10 ubiquitin ligase are degraded by distinct pathways. EMBO J. 25, 533–543 (2006).
Varelas, X., Ptak, C. & Ellison, M. J. Cdc34 self-association is facilitated by ubiquitin thiolester formation and is required for its catalytic activity. Mol. Cell Biol. 23, 5388–5400 (2003).
Deng, M. & Hochstrasser, M. Spatially regulated ubiquitin ligation by an ER/nuclear membrane ligase. Nature 443, 827–831 (2006).
Acknowledgements
We thank: Y. Reiss, G. Jona and O. Kerscher for valuable discussions; Y. Xie for providing the yeast deletion strain plate; Y. Xie, R. Hampton and T. Sommer for plasmids; and T. Biederer, R. Felberbaum, G. Jona and S. Kreft for comments on the manuscript. This work was supported by a U.S. National Institutes of Health (NIH) grant (GM046904) to M.H.
Author information
Authors and Affiliations
Contributions
T.R. performed all the experiments, and T.R. and M.H. conceived and designed the experiments and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures S1, S2, S3, S4, Supplementary Table and Methods (PDF 483 kb)
Rights and permissions
About this article
Cite this article
Ravid, T., Hochstrasser, M. Autoregulation of an E2 enzyme by ubiquitin-chain assembly on its catalytic residue. Nat Cell Biol 9, 422–427 (2007). https://doi.org/10.1038/ncb1558
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1558
This article is cited by
-
Legionella pneumophila inhibits immune signalling via MavC-mediated transglutaminase-induced ubiquitination of UBE2N
Nature Microbiology (2018)
-
Ubc7/Ube2g2 ortholog in Entamoeba histolytica: connection with the plasma membrane and phagocytosis
Parasitology Research (2018)
-
Multiple E3s promote the degradation of histone H3 variant Cse4
Scientific Reports (2017)
-
Ubiquitylation of p62/sequestosome1 activates its autophagy receptor function and controls selective autophagy upon ubiquitin stress
Cell Research (2017)
-
E2 enzymes: more than just middle men
Cell Research (2016)