Abstract
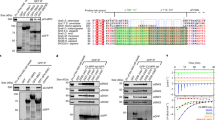
Clathrin-coated vesicles mediate diverse processes such as nutrient uptake, downregulation of hormone receptors, formation of synaptic vesicles, virus entry, and transport of biosynthetic proteins to lysosomes. Cycles of coat assembly and disassembly are integral features of clathrin-mediated vesicular transport (Fig. 1a). Coat assembly involves recruitment of clathrin triskelia, adaptor complexes and other factors that influence coat assembly, cargo sequestration, membrane invagination and scission1,2,3 (Fig. 1a). Coat disassembly is thought to be essential for fusion of vesicles with target membranes and for recycling components of clathrin coats to the cytoplasm for further rounds of vesicle formation. In vitro, cytosolic heat-shock protein 70 (Hsp70) and the J-domain co-chaperone auxilin catalyse coat disassembly4. However, a specific function of these factors in uncoating in vivo has not been demonstrated, leaving the physiological mechanism and significance of uncoating unclear. Here we report the identification and characterization of a Saccharomyces cerevisiae J-domain protein, Aux1. Inactivation of Aux1 results in accumulation of clathrin-coated vesicles, impaired cargo delivery, and an increased ratio of vesicle-associated to cytoplasmic clathrin. Our results demonstrate an in vivo uncoating function of a J domain co-chaperone and establish the physiological significance of uncoating in transport mediated by clathrin-coated vesicles.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Marsh, M. & McMahon, H. T. Science 285, 215–220 (1999).
Pishvaee, B. & Payne, G. S. Cell 95 , 443–446 (1998).
Schmid, S. L. Annu. Rev. Biochem. 66, 511–548 (1997).
Ungewickell, E. et al. Nature 378, 632–635 (1995).
Kelley, W. L. Trends Biochem. 23, 222–227 (1998).
Holstein, S. E. H., Ungewickell, H. & Ungewickell, E. J. Cell Biol. 135 , 925–937 (1996).
Blatch, G. L. & Lassle, M. Bioessays 21, 932–939 (1999).
Barlowe, C. et al. Cell 77, 895–907 (1994).
Yeung, B. G., Phan, H. L. & Payne, G. S. Mol. Biol. Cell 10, 3643– 3659 (1999).
Seeger, M. & Payne, G. S. EMBO J. 11, 2811–2818 (1992).
Bryant, N. J. & Stevens, T. H. Microbiol. Mol. Biol. Rev. 62, 230–247 ( 1998).
Tan, P. K., Davis, N. G., Sprague, G. F. & Payne, G. S. J. Cell Biol. 123, 1707–1716 (1993).
Raymond, C. K., Howald-Stevenson, I., Vater, C. A. & Stevens, T. H. Mol. Biol. Cell 3, 1389– 1402 (1992).
Jiang, R-F., Greener, T., Barouch, W., Greene, L. & Eisenberg, E. J. Biol. Chem. 272 , 6141–6145 (1997).
Honing, S., Kreimer, G., Robenek, H. & Jockusch, B. M. J. Cell Sci. 107, 1185–1196 (1994).
Umeda, A., Meyerholz, A. & Ungewickell, E. Eur. J. Cell Biol. 79, 336– 342 (2000).
Cremona, O. et al. Cell 99, 179–186 (1999).
DeLuca-Flaherty, C., McKay, D. B., Parham, P. & Hill, B. L. Cell 62, 875–887 ( 1990).
Harris, T. W., Hartwig, E., Horvitz, H. R. & Jorgensen, E. M. J. Cell Biol. 150, 589–599 (2000).
Owen, D. J. et al. Cell 97, 805–815 (1999).
Pishvaee, B., Munn, A. & Payne, G. S. EMBO J. 16, 2227– 2239 (1997).
Traub, L. M., Downs, M. A., Westrich, J. L. & Fremont, D. H. Proc. Natl Acad. Sci. USA 96, 8907– 8912 (1999).
Greener, T., Zhao, X., Nojima, H., Eisenberg, E. & Greene, L. E. J. Biol. Chem. 275, 1365–1370 (2000).
Baudin, A., Ozier-Kalogeropoulos, O., Denouel, A., Lacroute, F. & Cullin, C. Nucleic Acids Res. 21, 3329–3330 (1993).
Longtine, M. S. et al. Yeast 14, 953–961 (1998).
Vowels, J. J. & Payne, G. S. Mol. Biol. Cell 9, 1351–1365 (1998).
Ryazantsev, S., Tishchenko, V., Vasiliev, V., Zav'yalov, V. & Abramov, V. Eur. J. Biochem. 190, 393–399 ( 1990).
Bleazard, W. et al. Nature Cell Biol. 1, 298– 304 (1999).
Rieder, S. E., Banta, L. M., Kohrer, K., McCaffery, J. M. & Emr, S. D. Mol. Biol. Cell 7, 985–999 (1996).
Altschul, S. F. et al. Nucleic Acids Res. 25, 3389– 3402 (1997).
Thompson, J. D., Gibson, T. J., Plewniak, F., Jeanmougin, F. & Higgins, D. G. Nucleic Acids Res. 25, 4876–4882 ( 1997).
Bateman, A. et al. Nucleic Acids Res. 28, 263– 266 (2000).
Acknowledgements
We thank A. Nakamura for construction of the GST–Aux1 fusion, A. Rajasekaran for assistance with confocal microscopy, and A. van der Bliek and J. Hutton for comments on the manuscript. We also thank T. Graham for sharing unpublished results. This study was supported by NIH grant RO1 GM39040 to G.S.P.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Pishvaee, B., Costaguta, G., Yeung, B. et al. A yeast DNA J protein required for uncoating of clathrin-coated vesicles in vivo. Nat Cell Biol 2, 958–963 (2000). https://doi.org/10.1038/35046619
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/35046619
This article is cited by
-
J domain independent functions of J proteins
Cell Stress and Chaperones (2016)
-
Arabidopsis thaliana J-class heat shock proteins: cellular stress sensors
Functional & Integrative Genomics (2009)
-
The clathrin-binding motif and the J-domain of Drosophila Auxilin are essential for facilitating Notch ligand endocytosis
BMC Developmental Biology (2008)
-
Ptc1p regulates cortical ER inheritance via Slt2p
The EMBO Journal (2006)