Abstract
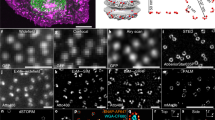
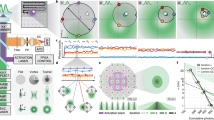
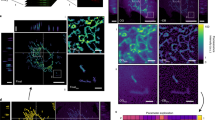
Live-cell imaging technology using fluorescent proteins (green fluorescent protein and its homologues) has revolutionized the study of cellular dynamics1,2. But tools that can quantitatively analyse complex spatiotemporal processes in live cells remain lacking. Here we describe a new technique — fast multi-colour four-dimensional imaging combined with automated and quantitative time-space reconstruction — to fill this gap. As a proof of principle, we apply this method to study the re-formation of the nuclear envelope in live cells3,4,5,6. Four-dimensional imaging of three spectrally distinct fluorescent proteins is used to simultaneously visualize three different cellular compartments at high speed and with high spatial resolution. The highly complex data, comprising several thousand images from a single cell, were quantitatively reconstructed in time–space by software developed in-house. This analysis reveals quantitative and qualitative insights into the highly ordered topology of nuclear envelope formation, in correlation with chromatin expansion — results that would have been impossible to achieve by manual inspection alone. Our new technique will greatly facilitate study of the highly ordered dynamic architecture of eukaryotic cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Lamond, A. I. & Earnshaw, W. C. Science 280, 547–553 (1998).
Misteli, T. Science 291, 843–847 (2001).
Ellenberg, J. et al. J. Cell Biol. 138, 1193–1206 (1997).
Haraguchi, T. et al. J. Cell Sci. 113, 779–794 (2000).
Collas, I. & Courvalin, J. C. Sorting nuclear membrane proteins at mitosis. Trends Cell Biol. 10, 5–8 (2000).
Marshall, I. C. B. & Wilson, K. L. Trends Cell Biol. 7, 69–74 (1997).
Eils, R., Gerlich, D., Tvarusko, W., Spector, D. L. & Misteli, T. Mol. Biol. Cell 11, 413–418 (2000).
Belmont, A. S., Dietzel, S., Nye, A. C., Strukov, Y. G. & Tumbar, T. Curr. Opin. Cell Biol. 11, 307–311 (1999).
Lippincott-Schwartz, J. et al. Methods Cell Biol. 58, 261–281 (1999).
Tvaruskó, W. et al. Proc. Natl Acad. Sci. USA 96, 7950–7955 (1999).
Qian, H., Sheetz, M. P. & Elson, E. L. Biophys. J. 60, 910–921 (1991).
Marshall, W. F. et al. Interphase chromosomes undergo constrained diffusional motion in living cells. Curr. Biol. 7, 930–939 (1997).
Rustom, A. et al. Biotechniques 28, 722–730 (2000).
Thomas, C. F. & White, J. G. Trends Biotechnol. 16, 175–182 (1998).
Black, M. J., Sapiro, G., Marimont, D. & Heeger, D. IEEE Trans. Image Process. 7, 421–432 (1998).
Watt, A. & Watt, M. Advanced Animation and Rendering Techniques (ed. Wegner, P.) (Addison-Wesley, New York, 1992).
Terasaki, M. Mol. Biol. Cell 11, 897–914 (2000).
Gerace, L. & Burke, B. Annu. Rev. Cell Biol. 4, 335–374 (1988).
Tsien, R. Y. Annu. Rev. Biochem. 67, 509–544 (1998).
Matz, M. V. et al. Nature Biotechnol. 17, 969–973 (1999); erratum ibid. 17, 1227 (1999).
Niswender, K. D., Blackman, S. M., Rohde, L., Magnuson, M. A. & Piston, D. W. J. Microsc. 180, 109–116 (1995).
Akey, C. W. J. Mol. Biol. 248, 273–293 (1995).
Besl, P. J. & McKay, N. D. IEEE Trans. Pattern Analysis Machine Intelligence 14, 239–256 (1992).
Bodoor, K. et al. J. Cell Sci. 112, 2253–2264 (1999).
Broers, J. L. et al. J. Cell Sci. 112, 3463–3475 (1999).
Chaudhary, N. & Courvalin, J.-C. J. Cell Biol. 122, 295–306 (1993).
Ellenberg, J. & Lippincott-Schwartz, J. Methods 19, 362–372 (1999).
Acknowledgements
We thank B. Oakley for the γ-tubulin cDNA; N. Daigle for technical assistance; J. Mattes for providing the matching algorithm; and C. Conrad and C. Athale for critical comments on the manuscript. The bioinformatics group acknowledges the support of the German Federal Ministry of Education and Research (BMBF) through the BioFuture grant (AZ 11880A) and of the German Research Council (Ei 358/2-1 and Ei 358/1-1). J.B. was supported by a fellowship through EMBL's international Ph.D. programme. Part of this work was performed in collaboration with T.I.L.L.-Photonics, Munich, Germany.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary figures
Figure S1 Detailed description of 3-D reconstruction technique (PDF 317 kb)
Figure S2 Integration of LBR into patches and the NE measured in single slices versus 4-D analysis
Figure S3 Comparison of volume expansion quantified with 4-D methods versus 2-D analysis in projected image stacks
Figure S4 Reliability of 4-D quantification with threshold segmentation
Supplementary movie
Movie 1 800 1000 Movie 1 Assembly of image stack from optical slices and reconstruction of a 3-D surface model of LBR-EYFP patches (green) on chromatin surface (red). First, the pre-processed optical image slices of a single image stack are displayed sequentially. Then, they are removed stepwise to reveal the 3-D reconstructed surface model. For further details see fig. 1. (MOV 285 kb)
Supplementary movie
Movie 2 Animated sequence of 3-D reconstructions of early LBR patch formation on chromatin surface. For details see fig. 2. (MOV 1659 kb)
Supplementary movie
Movie 3 Morphed reconstruction of NE and chromatin expansion. Four intermediate states of chromatin and NE surfaces per time frame were interpolated, thereby increasing temporal resolution. For details see fig. 3. (MOV 1022 kb)
Rights and permissions
About this article
Cite this article
Gerlich, D., Beaudouin, J., Gebhard, M. et al. Four-dimensional imaging and quantitative reconstruction to analyse complex spatiotemporal processes in live cells. Nat Cell Biol 3, 852–855 (2001). https://doi.org/10.1038/ncb0901-852
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb0901-852
This article is cited by
-
Mechanisms driving acentric chromosome transmission
Chromosome Research (2020)
-
A transient pool of nuclear F-actin at mitotic exit controls chromatin organization
Nature Cell Biology (2017)
-
Differential pH-dependent cellular uptake pathways among foamy viruses elucidated using dual-colored fluorescent particles
Retrovirology (2012)
-
Assays for mitotic chromosome condensation in live yeast and mammalian cells
Chromosome Research (2009)
-
Dynamic proteomics in individual human cells uncovers widespread cell-cycle dependence of nuclear proteins
Nature Methods (2006)