Abstract
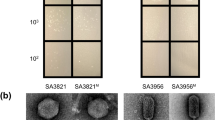
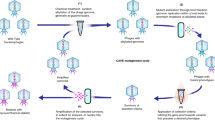
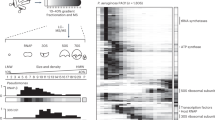
Over evolutionary time bacteriophages have developed unique proteins that arrest critical cellular processes to commit bacterial host metabolism to phage reproduction. Here, we apply this concept of phage-mediated bacterial growth inhibition to antibiotic discovery. We sequenced 26 Staphylococcus aureus phages and identified 31 novel polypeptide families that inhibited growth upon expression in S. aureus. The cellular targets for some of these polypeptides were identified and several were shown to be essential components of the host DNA replication and transcription machineries. The interaction between a prototypic pair, ORF104 of phage 77 and DnaI, the putative helicase loader of S. aureus, was then used to screen for small molecule inhibitors. Several compounds were subsequently found to inhibit both bacterial growth and DNA synthesis. Our results suggest that mimicking the growth-inhibitory effect of phage polypeptides by a chemical compound, coupled with the plethora of phages on earth, will yield new antibiotics to combat infectious diseases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Salgado, C.D., Farr, B.M. & Calfee, D.P. Community-acquired methicillin-resistant Staphylococcus aureus: a meta-analysis of prevalence and risk factors. Clin. Infect. Dis. 36, 131–139 (2003).
Tenover, F.C., Biddle, J.W. & Lancaster, M.V. Increasing resistance to vancomycin and other glycopeptides in Staphylococcus aureus. Emerg. Infect. Dis. 7, 327–332 (2001).
Khare, M. & Keady, D. Antimicrobial therapy of methicillin resistant Staphylococcus aureus infection. Expert Opin. Pharmacother. 4, 165–177 (2003).
Kuroda, M. et al. Whole genome sequencing of meticillin-resistant Staphylococcus aureus. Lancet 357, 1225–1240 (2001).
Moir, D.T., Shaw, K.J., Hare, R.S. & Vovis, G.F. Genomics and antimicrobial drug discovery. Antimicrob. Agents Chemother. 43, 439–446 (1999).
McDevitt, D. & Rosenberg, M. Exploiting genomics to discover new antibiotics. Trends Microbiol. 9, 611–617 (2001).
Ji, Y. et al. Identification of critical staphylococcal genes using conditional phenotypes generated by antisense RNA. Science 293, 2266–2269 (2001).
Forsyth, R.A. et al. A genome-wide strategy for the identification of essential genes in Staphylococcus aureus. Mol. Microbiol. 43, 1387–1400 (2002).
Kobayashi, K. et al. Essential Bacillus subtilis genes. Proc. Natl. Acad. Sci. USA 100, 4678–4683 (2003).
Nechaev, S. & Severinov, K. Inhibition of Escherichia coli RNA polymerase by bacteriophage T7 gene 2 protein. J. Mol. Biol. 289, 815–826 (1999).
Orsini, G., Ouhammouch, M., Le Caer, J.P. & Brody, E.N. The asiA gene of bacteriophage T4 codes for the anti-sigma 70 protein. J. Bacteriol. 175, 85–93 (1993).
Mallory, J.B., Alfano, C. & McMacken, R. Host virus interactions in the initiation of bacteriophage lambda DNA replication. Recruitment of Escherichia coli DnaB helicase by lambda P replication protein. J. Biol. Chem. 265, 13297–13307 (1990).
Odegrip, R., Schoen, S., Haggard-Ljungquist, E., Park, K. & Chattoraj, D.K. The interaction of bacteriophage P2 B protein with Escherichia coli DnaB helicase. J. Virol. 74, 4057–4063 (2000).
Biswas, B. et al. Bacteriophage therapy rescues mice bacteremic from a clinical isolate of vancomycin-resistant Enterococcus faecium. Infect. Immun. 70, 204–210 (2002).
Stone, R. Bacteriophage therapy. Stalin's forgotten cure. Science 298, 728–731 (2002).
Schuch, R., Nelson, D. & Fischetti, V.A. A bacteriolytic agent that detects and kills Bacillus anthracis. Nature 418, 884–889 (2002).
Loeffler, J.M., Nelson, D. & Fischetti, V.A. Rapid killing of Streptococcus pneumoniae with a bacteriophage cell wall hydrolase. Science 294, 2170–2172 (2001).
Ackermann, H.-W. & DuBow, M.S. Viruses of Prokaryotes, vol. 2 (CRC Press, Boca Raton, Florida, 1987).
Kaneko, J., Kimura, T., Narita, S., Tomita, T. & Kamio, Y. Complete nucleotide sequence and molecular characterization of the temperate staphylococcal bacteriophage phiPVL carrying Panton-Valentine leukocidin genes. Gene 215, 57–67 (1998).
Tauriainen, S., Karp, M., Chang, W. & Virta, M. Recombinant luminescent bacteria for measuring bioavailable arsenite and antimonite. Appl. Environ. Microbiol. 63, 4456–4461 (1997).
Kreiswirth, B.N. et al. The toxic shock syndrome exotoxin structural gene is not detectably transmitted by a prophage. Nature 305, 709–712 (1983).
Wang, I.N., Smith, D.L. & Young, R. Holins: the protein clocks of bacteriophage infections. Annu. Rev. Microbiol. 54, 799–825 (2000).
Young, I., Wang, I. & Roof, W.D. Phages will out: strategies of host cell lysis. Trends Microbiol. 8, 120–128 (2000).
Sonnhammer, E.L., von Heijne, G. & Krogh, A. A hidden Markov model for predicting transmembrane helices in protein sequences. Proc. Int. Conf. Intell. Syst. Mol. Biol. 6, 175–182 (1998).
Simpson, R.J. Proteins and proteomics: a laboratory manual (Cold Spring Harbor Laboratory Press, New York, 2002).
Bruand, C., Farache, M., McGovern, S., Ehrlich, S.D. & Polard, P. DnaB, DnaD and DnaI proteins are components of the Bacillus subtilis replication restart primosome. Mol. Microbiol. 42, 245–255 (2001).
Marsin, S., McGovern, S., Ehrlich, S.D., Bruand, C. & Polard, P. Early steps of Bacillus subtilis primosome assembly. J. Biol. Chem. 276, 45818–45825 (2001).
Velten, M. et al. A two-protein strategy for the functional loading of a cellular replicative DNA helicase. Mol. Cell 11, 1009–1020 (2003).
Mathis, G. Probing molecular interactions with homogeneous techniques based on rare earth cryptates and fluorescence energy transfer. Clin. Chem. 41, 1391–1397 (1995).
Jana, M., Luong, T.T., Komatsuzawa, H., Shigeta, M. & Lee, C.Y. A method for demonstrating gene essentiality in Staphylococcus aureus. Plasmid 44, 100–104 (2000).
Walsh, C. Antibiotics: Actions, Origins, Resistance (ASM Press, Washington, DC, 2003).
Sambrook, J., Fritsch, E.F. & Maniatis, T. Molecular Cloning: A laboratory manual, edn. 2 (Cold Spring Harbor Laboratory Press, New York, 1989).
Adams, M.H. Bacteriophages (Interscience Publishers, New York, NY, 1959).
Schenk, S. & Laddaga, R.A. Improved method for electroporation of Staphylococcus aureus. FEMS Microbiol. Lett. 73, 133–138 (1992).
Methot, N., Song, M.S. & Sonenberg, N. A region rich in aspartic acid, arginine, tyrosine, and glycine (DRYG) mediates eukaryotic initiation factor 4B (eIF4B) self-association and interaction with eIF3. Mol. Cell Biol. 16, 5328–5334 (1996).
Kaelin, W.G., Jr. et al. Expression cloning of a cDNA encoding a retinoblastoma-binding protein with E2F-like properties. Cell 70, 351–364 (1992).
De Crescenzo, G., Grothe, S., Lortie, R., Debanne, M.T. & O'Connor-McCourt, M. Real-time kinetic studies on the interaction of transforming growth factor alpha with the epidermal growth factor receptor extracellular domain reveal a conformational change model. Biochemistry 39, 9466–9476 (2000).
NCCLS specialty collection: susceptibility testing, edn. 5 (NCCLS, Wayne, PA, 2000).
Kaito, C., Kurokawa, K., Hossain, M.S., Akimitsu, N. & Sekimizu, K. Isolation and characterization of temperature-sensitive mutants of the Staphylococcus aureus dnaC gene. FEMS Microbiol. Lett. 210, 157–164 (2002).
Acknowledgements
We thank M. Virta for providing us with the pTOO21 vector and C.Y. Lee for vectors used in essentiality analysis. We thank all PhageTech technical staff for their assistance during the course of this research. We also would like to thank the National Research Council (Canada) Industrial Research Assistance Program for their support of part of our research. J.L., M.D., M.C., N.H., T.K., G.M. and R.S. are recipients of the Natural Sciences and Engineering Research Council of Canada Industrial Research Fellowship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
All the authors are/were employed or compensated by PhageTech, which owns the intellectual property related to the work reported in this paper.
Rights and permissions
About this article
Cite this article
Liu, J., Dehbi, M., Moeck, G. et al. Antimicrobial drug discovery through bacteriophage genomics. Nat Biotechnol 22, 185–191 (2004). https://doi.org/10.1038/nbt932
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt932
This article is cited by
-
Overview of the risks of Staphylococcus aureus infections and their control by bacteriophages and bacteriophage-encoded products
Brazilian Journal of Microbiology (2021)
-
Approaches to optimize therapeutic bacteriophage and bacteriophage-derived products to combat bacterial infections
Virus Genes (2020)
-
Peculiarities of Staphylococcus aureus phages and their possible application in phage therapy
Applied Microbiology and Biotechnology (2019)
-
Bacillus safensis FO-36b and Bacillus pumilus SAFR-032: a whole genome comparison of two spacecraft assembly facility isolates
BMC Microbiology (2018)