Abstract
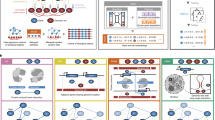
We describe an algorithm for discovering regulatory networks of gene modules, GRAM (Genetic Regulatory Modules), that combines information from genome-wide location and expression data sets. A gene module is defined as a set of coexpressed genes to which the same set of transcription factors binds. Unlike previous approaches1,2,3,4,5 that relied primarily on functional information from expression data, the GRAM algorithm explicitly links genes to the factors that regulate them by incorporating DNA binding data, which provide direct physical evidence of regulatory interactions. We use the GRAM algorithm to describe a genome-wide regulatory network in Saccharomyces cerevisiae using binding information for 106 transcription factors profiled in rich medium conditions data* from over 500 expression experiments. We also present a genome-wide location analysis data set for regulators in yeast cells treated with rapamycin, and use the GRAM algorithm to provide biological insights into this regulatory network.
*Note: In the version of this article initially published online, the word "and" was omitted from the fourth sentence of the abstract, altering the meaning. The sentence should read: "We use the GRAM algorithm to describe a genome-wide regulatory network in Saccharomyces cerevisiae using binding information for 106 transcription factors profiled in rich medium conditions and data from over 500 expression experiments." This mistake has been corrected for the HTML and print versions of the article.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Eisen, M.B., Spellman, P.T., Brown, P.O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl. Acad. Sci. USA 95, 14863–14868 (1998).
Segal, E. et al. Module networks: identifying regulatory modules and their condition-specific regulators from gene expression data. Nat. Genet. 34, 166–176 (2003).
Ihmels, J. et al. Revealing modular organization in the yeast transcriptional network. Nat. Genet. 31, 370–377 (2002).
Pilpel, Y., Sudarsanam, P. & Church, G.M. Identifying regulatory networks by combinatorial analysis of promoter elements. Nat. Genet. 29, 153–159 (2001).
Berman, B.P. et al. Exploiting transcription factor binding site clustering to identify cis-regulatory modules involved in pattern formation in the Drosophila genome. Proc. Natl. Acad. Sci. USA 99, 757–762 (2002).
Lee, T.I. et al. Transcriptional regulatory networks in Saccharomyces cerevisiae. Science 298, 799–804 (2002).
Mewes, H.W. et al. MIPS: a database for genomes and protein sequences. Nucleic Acids Res. 30, 31–34 (2002).
Holland, M.J., Yokoi, T., Holland, J.P., Myambo, K. & Innis, M.A. The GCR1 gene encodes a positive transcriptional regulator of the enolase and glyceraldehyde-3-phosphate dehydrogenase gene families in Saccharomyces cerevisiae. Mol. Cell Biol. 7, 813–820 (1987).
Baker, H.V. Glycolytic gene expression in Saccharomyces cerevisiae: nucleotide sequence of GCR1, null mutants, and evidence for expression. Mol. Cell Biol. 6, 3774–3784 (1986).
Forsburg, S.L. & Guarente, L. Identification and characterization of HAP4: a third component of the CCAAT-bound HAP2/HAP3 heteromer. Genes Dev. 3, 1166–1178 (1989).
Matys, V. et al. TRANSFAC: transcriptional regulation, from patterns to profiles. Nucleic Acids Res. 31, 374–378 (2003).
Jacinto, E. & Hall, M.N. Tor signalling in bugs, brain and brawn. Nat. Rev. Mol. Cell Biol. 4, 117–126 (2003).
Crespo, J.L. & Hall, M.N. Elucidating TOR signaling and rapamycin action: lessons from Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 66, 579–591 (2002).
Raught, B., Gingras, A.C. & Sonenberg, N. The target of rapamycin (TOR) proteins. Proc. Natl. Acad. Sci. USA 98, 7037–7044 (2001).
Hardwick, J.S., Kuruvilla, F.G., Tong, J.K., Shamji, A.F. & Schreiber, S.L. Rapamycin-modulated transcription defines the subset of nutrient-sensitive signaling pathways directly controlled by the Tor proteins. Proc. Natl. Acad. Sci. USA 96, 14866–14870 (1999).
Shamji, A.F., Kuruvilla, F.G. & Schreiber, S.L. Partitioning the transcriptional program induced by rapamycin among the effectors of the Tor proteins. Curr. Biol. 10, 1574–1581 (2000).
Cardenas, M.E., Cutler, N.S., Lorenz, M.C., Di Como, C.J. & Heitman, J. The TOR signaling cascade regulates gene expression in response to nutrients. Genes Dev. 13, 3271–3279 (1999).
Hasan, R. et al. The control of the yeast H2O2 response by the Msn2/4 transcription factors. Mol. Microbiol. 45, 233–241 (2002).
Rep, M., Krantz, M., Thevelein, J.M. & Hohmann, S. The transcriptional response of Saccharomyces cerevisiae to osmotic shock. Hot1p and Msn2p/Msn4p are required for the induction of subsets of high osmolarity glycerol pathway-dependent genes. J. Biol. Chem. 275, 8290–8300 (2000).
Boy-Marcotte, E., Perrot, M., Bussereau, F., Boucherie, H. & Jacquet, M. Msn2p and Msn4p control a large number of genes induced at the diauxic transition which are repressed by cyclic AMP in Saccharomyces cerevisiae. J. Bacteriol. 180, 1044–1052 (1998).
Martinez-Pastor, M.T. et al. The Saccharomyces cerevisiae zinc finger proteins Msn2p and Msn4p are required for transcriptional induction through the stress response element (STRE). EMBO J. 15, 2227–2235 (1996).
Schuller, H.J. Transcriptional control of nonfermentative metabolism in the yeast Saccharomyces cerevisiae. Curr. Genet. 43, 139–160 (2003).
Crespo, J.L., Powers, T., Fowler, B. & Hall, M.N. The TOR-controlled transcription activators GLN3, RTG1, and RTG3 are regulated in response to intracellular levels of glutamine. Proc. Natl. Acad. Sci. USA 99, 6784–6789 (2002).
Komeili, A., Wedaman, K.P., O'Shea, E.K. & Powers, T. Mechanism of metabolic control: target of rapamycin signaling links nitrogen quality to the activity of the Rtg1 and Rtg3 transcription factors. J. Cell Biol. 151, 863–878 (2000).
Liao, X. & Butow, R.A. RTG1 and RTG2: two yeast genes required for a novel path of communication from mitochondria to the nucleus. Cell 72, 61–71 (1993).
Pinkham, J.L. & Guarente, L. Cloning and molecular analysis of the HAP2 locus: a global regulator of respiratory genes in Saccharomyces cerevisiae. Mol. Cell Biol. 5, 3410–3416 (1985).
Dang, V.D., Bohn, C., Bolotin-Fukuhara, M. & Daignan-Fornier, B. The CCAAT box-binding factor stimulates ammonium assimilation in Saccharomyces cerevisiae, defining a new cross-pathway regulation between nitrogen and carbon metabolisms. J. Bacteriol. 178, 1842–1849 (1996).
Dang, V.D., Valens, M., Bolotin-Fukuhara, M. & Daignan-Fornier, B. Cloning of the ASN1 and ASN2 genes encoding asparagine synthetases in Saccharomyces cerevisiae: differential regulation by the CCAAT-box-binding factor. Mol. Microbiol. 22, 681–692 (1996).
Shen-Orr, S.S., Milo, R., Mangan, S. & Alon, U. Network motifs in the transcriptional regulation network of Escherichia coli. Nat. Genet. 31, 64–68 (2002).
Coffman, J.A., Rai, R., Cunningham, T., Svetlov, V. & Cooper, T.G. Gat1p, a GATA family protein whose production is sensitive to nitrogen catabolite repression, participates in transcriptional activation of nitrogen-catabolic genes in Saccharomyces cerevisiae. Mol. Cell Biol. 16, 847–858 (1996).
Acknowledgements
Z.B-J. is supported by the Program in Mathematics and Molecular Biology at Florida State University through the Burroughs Wellcome Fund Interfaces Program. G.G. is supported by a National Defense Engineering and Science graduate fellowship. This work was partially funded by a US National Institutes of Health grant.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Bar-Joseph, Z., Gerber, G., Lee, T. et al. Computational discovery of gene modules and regulatory networks. Nat Biotechnol 21, 1337–1342 (2003). https://doi.org/10.1038/nbt890
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt890
This article is cited by
-
Inflammatory and interferon gene expression signatures in patients with mitochondrial disease
Journal of Translational Medicine (2023)
-
Identification, characterization, and prognosis investigation of pivotal genes shared in different stages of breast cancer
Scientific Reports (2023)
-
The modular biochemical reaction network structure of cellular translation
npj Systems Biology and Applications (2023)
-
High-throughput screening identified miR-7-2-3p and miR-29c-3p as metastasis suppressors in gallbladder carcinoma
Journal of Gastroenterology (2020)
-
Markov chain Monte Carlo simulation of a Bayesian mixture model for gene network inference
Genes & Genomics (2019)