Abstract
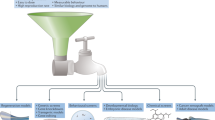
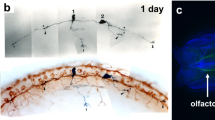
The zebrafish (Danio rerio) provides an excellent model for studying vertebrate development and human disease because of its ex utero, optically transparent embryogenesis and amenability to in vivo manipulation. The rapid embryonic developmental cycle, large clutch sizes and ease of maintenance at large numbers also add to the appeal of this species. Considerable genomic data has recently become publicly available that is aiding the construction of zebrafish microarrays, thus permitting global gene expression analysis. The zebrafish is also suitable for chemical genomics, in part as a result of the permeability of its embryos to small molecules and consequent avoidance of external confounding maternal effects. Finally, there is increasing characterization and analysis of zebrafish models of human disease. Thus, the zebrafish offers a high-quality, high-throughput bioassay tool for determining the biological effect of small molecules as well as for dissecting biological pathways.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Alaoui-Ismaili, M.H., Lomedico, P.T. & Jindal, S. Chemical genomics: discovery of disease genes and drugs. Drug Disc. Today 7, 292–294 (2002).
Törnell, J. & Snaith, M. Transgenic systems in drug discovery: from target identification to humanized mice. Drug Disc. Today 7, 461–470 (2002).
Gmuender, H. Perspectives and challenges for DNA microarrays in drug discovery and development. Biotechniques 32, 152–158 (2002).
Zambrowicz, B.P. & Sands, A.T. Knockouts model the 100 best-selling drugs—will they model the next 100? Nat. Rev. Drug Discov. 2, 38–51 (2003).
Grunwald, D.J. & Eisen, J.S. Headwaters of the zebrafish—emergence of a new model vertebrate. Nat. Rev. Genet. 3, 717–723 (2002).
Shin, J.T. & Fishman, M.C. From zebrafish to human: modular medical models. Annu. Rev. Genomics Hum. Genet. 3, 311–340 (2002).
Neumann, C.J. Vertebrate development: a view from zebrafish. Semin. Cell Dev. Biol. 13, 469 (2002).
Lawson, N.D. & Weinstein, B.M. Arteries and veins: making a difference with zebrafish. Nat. Rev. Genet. 3, 674–682 (2002).
Spitsbergen, J.M. & Kent, M.L. The state of the art of the zebrafish model for toxicology and toxicologic pathology research—advantages and current limitations. Toxicol. Pathol. 31, 62–87 (2003).
Neely, M.N., Pfeifer, J.D. & Caparon, M. Streptococcus-zebrafish model of bacterial pathogenesis. Infect. Immun. 70, 3904–3914 (2002).
Ju, B. et al. Faithful expression of green fluorescent protein (GFP) in transgenic zebrafish embryos under control of zebrafish gene promoters. Dev. Gene. 25, 158–167 (1999).
Golling, G. et al. Insertional mutagenesis in zebrafish rapidly identifies genes essential for early vertebrate development. Nat. Genet. 31, 135–140 (2002).
Nasevicius, A. & Ekker, S.C. The zebrafish as a novel system for functional genomics and therapeutic development applications. Curr. Opin. Mol. Ther. 3, 224–228 (2001).
Heasman, J. Morpholino oligos: making sense of antisense? Dev. Biol. 243, 209–214 (2002).
Halloran, M.C. et al. Laser-induced gene expression in specific cells of transgenic zebrafish. Development 127, 1953–1960 (2000).
Galloway, J.L. & Zon, L.I. Ontogeny of hematopoiesis: examining the emergence of hematopoietic cells in the vertebrate embryo. Curr. Top. Dev. Biol. 53, 139–158 (2003).
Sehnert, A.J. & Stainier, D.Y.R. A window to the heart: can zebrafish mutants help us understand heart disease in humans? Trends Genet. 18, 491–494 (2002).
Hove, J.R. et al. Intracardiac fluid forces are an essential epigenetic factor for embryonic cardiogenesis. Nature 421, 172–177 (2003).
Goldsmith, P. & Harris, W.A. The zebrafish as a tool for understanding the biology of visual disorders. Semin. Cell Dev. Biol. 14, 11–18 (2003).
Lawson, N.D. & Weinstein, B.M. Arteries and veins: making a difference with zebrafish. Nat. Rev. Genet. 3, 674–682 (2002).
Du, S.J., Frenkel, V., Kindschi, G. & Zohar, Y. Visualizing normal and defective bone development in zebrafish embryos using the fluorescent chromophore calcein. Dev. Biol. 238, 239–246 (2001).
Whitfield, T.T. Zebrafish as a model for hearing and deafness. J. Neurobiol. 53, 157–171 (2002).
Amatruda, J.F., Shepard, J.L., Stern, H.M & Zon, L.I. Zebrafish as a cancer model system. Cancer Cell 1, 229–231 (2002).
Langenau, D.M. et al. Myc-induced T-cell leukemia in transgenic zebrafish. Science 299, 887–890 (2003).
Tomasiewicz, H.G., Flaherty, D.B., Soria, J.P. & Wood, J.G. Transgenic zebrafish model of neurodegeneration. J. Neurosci. Res. 70, 734–745 (2002).
Guyon, J.R. et al. The dystrophin associated protein complex in zebrafish. Hum. Mol. Genet. 12, 601–615 (2003).
Chambers, S.P., Anderson, L.V.B., Maguire, G.M., Dodd, A. & Love, D.R. Sarcoglycans of the zebrafish: orthology and localization to the sarcolemma and myosepta of muscle. Biochem. Biophys. Res. Commun. 303, 488–495 (2003).
Tamai, K.T., Vardhanabhuti, V., Arthur, S., Foulkes, N.S. & Whitmore, D. Flies and fish: birds of a feather. J. Neuroendocrinol. 15, 344–349 (2003).
O'Kane, C.J. Modelling human diseases in Drosophila and Caenorhabditis. Semin. Cell Dev. Biol. 14, 3–10 (2003).
Brown, W.R.A., Hubbard, S.J., Tickle, C. & Wilson, S.A. The chicken as a model for large-scale analysis of vertebrate gene function. Nat. Rev. Genet. 4, 87–98 (2003).
Anderson, K.V. & Ingham, P.W. The transformation of the model organism: a decade of developmental genetics. Nat. Genet. 33 (Suppl.), 285–293 (2003).
Postlethwait, J.H. et al. Zebrafish comparative genomics and the origins of vertebrate chromosomes. Genome Res. 10, 1890–1902 (2000).
Smith, C. Drug target validation: hitting the target. Nature 422, 341–347 (2003).
Colangelo, V. et al. Gene expression profiling of 12633 genes in Alzheimer hippocampal CA1: transcription and neurotrophic factor down-regulation and up-regulation of apoptotic and pro-inflammatory signaling. J. Neurosci. Res. 70, 462–473 (2002).
Haslett, J.N. et al. Gene expression comparison of biopsies from Duchenne muscular dystrophy (DMD) and normal skeletal muscle. Proc. Natl. Acad. Sci. USA. 99, 15000–15005 (2002).
Orr, H.T. Microarrays and polyglutamine disorders: reports from the Hereditary Disease Array Group. Hum. Mol. Genet. 11, 1909–1910 (2002).
Clark, M.D. et al. An oligonucleotide fingerprint normalized and expressed sequence tag characterized zebrafish cDNA library. Genome Res. 11, 1594–1602 (2001).
Tabuchi, Y., Kondo, T., Ogawa, R. & Mori, H. DNA microarray analyses of genes elicited by ultrasound in human U937 cells. Biochem. Biophys. Res. Commun. 290, 498–503 (2002).
Zheng, X.F.S. & Chan, T.-F. Chemical genomics in the global study of protein functions. Drug Dev. Today 7, 197–205 (2002).
Hughes, T.R. et al. Functional discovery via a compendium of expression profiles. Cell 102, 109–126 (2000).
Carroll, A.S., Bishop, A.C., DeRisi, J.L., Shokat, K.M. & O'Shea, E.K. Chemical inhibition of the Pho85 cyclin-dependent kinase reveals a role in the environmental stress response. Proc. Natl. Acad. Sci. USA 98, 12578–12583 (2001).
Martin, D.A. et al. Defective CD95/APO-1/Fas signal complex formation in the human autoimmune lymphoproliferative syndrome, type Ia. Proc. Natl. Acad. Sci. USA 96, 4552–4557 (1999).
Ihle, J.N. Minireview: the challenges of translating knockout phenotypes into gene function. Cell 102, 131–134 (2000).
Peterson, R.T., Link, B.A., Dowling, J.E. & Schreiber, S.L. Small molecule developmental screens reveal the logic and timing of vertebrate development. Proc. Natl. Acad. Sci. USA 97, 12965–12969 (2000).
Chan, J., Bayliss, P.E., Wood, J.M. & Roberts, T.M. Dissection of angiogenic signaling in zebrafish using a chemical genetic approach. Cancer Cell 1, 257–267 (2002).
Ton, C., Stamatiou, D., Dzau, V.J. & Liew, C.-C. Construction of a zebrafish cDNA microarray: gene expression profiling of the zebrafish during development. Biochem. Biophys. Res. Commun. 296, 1134–1142 (2002).
Lo, J. et al. 15,000 unique zebrafish EST clusters and their future use in microarray for profiling gene expression patterns during embryogenesis. Genome Res. 13, 455–466 (2003).
Herwig, R., Aanstad, P., Clark, M. & Lehrach, H. Statistical evaluation of differential expression on cDNA nylon arrays with replicated experiments. Nucleic Acids Res. 29, E117 (2001).
Stickney, H.L. et al. Rapid mapping of zebrafish mutations with SNPs and oligonucleotide microarrays. Genome Res. 12, 1929–1934 (2002).
Saikumar, P. et al. Apoptosis: definition, mechanisms, and relevance to disease. Am. J. Med. 107, 489–506 (1999).
Yabu, T., Kishi, S., Okazaki, T. & Yamashita, M. Characterization of zebrafish caspase-3 and induction of apoptosis through ceramide generation in fish fathead minnow tailbud cells and zebrafish embryo. Biochem. J. 360, 39–47 (2001).
Mogi, A. et al. Caspase activities and tumor necrosis factor receptor R1 (p55) level are elevated in the substantia nigra from Parkinsonian brain. J. Neural Transmem. 107, 335–341 (2000).
Smale G., Nichols N.R., Brady D.R., Finch C.E. & Horton W.E. Jr. Evidence for apoptotic cell death in Alzheimer's disease. Exp. Neurol. 133, 225–230 (1995).
Ventimiglia, R. et al. Research overview role of caspases in Neuronal apoptosis. Drug Dev. Res. 52, 515–533 (2001).
Hanahan, D. & Weinberg, R.A. The hallmarks of cancer. Cell 100, 57–70 (2000).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Inohara, N. & Nuñez, G. Genes with homology to mammalian apoptosis regulators identified in zebrafish. Cell Death Differ. 7, 509–510 (2000).
Degterev, A. et al. Identification of small-molecule inhibitors of interaction between the BH3 domain and Bcl-XL . Nat. Cell Biol. 3, 173–182 (2001).
Los, G. et al. Using mRNA expression profiling to determine anticancer drug efficacy. Cytometry 47, 66–71 (2002).
Miyashita, T. et al. Tumor suppressor p53 is a regulator of BCL-2 and BAX gene expression in vitro and in vivo. Oncogene 9, 1799–1805 (1994).
Acknowledgements
We acknowledge funding support of the University of Auckland Vice Chancellor's Development Fund, University of Auckland Research Committee, Lottery Grants Board of New Zealand and the Maurice and Phyllis Paykel Trust. We thank Vivienne Ward for assistance in graphics production.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pichler, F., Laurenson, S., Williams, L. et al. Chemical discovery and global gene expression analysis in zebrafish. Nat Biotechnol 21, 879–883 (2003). https://doi.org/10.1038/nbt852
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt852
This article is cited by
-
Marine fungal metabolite butyrolactone I prevents cognitive deficits by relieving inflammation and intestinal microbiota imbalance on aluminum trichloride-injured zebrafish
Journal of Neuroinflammation (2022)
-
Detection of 8-oxoguanine and apurinic/apyrimidinic sites using a fluorophore-labeled probe with cell-penetrating ability
BMC Molecular and Cell Biology (2019)
-
First profiling of lysine crotonylation of myofilament proteins and ribosomal proteins in zebrafish embryos
Scientific Reports (2018)
-
Indomethacin treatment prior to pentylenetetrazole-induced seizures downregulates the expression of il1b and cox2 and decreases seizure-like behavior in zebrafish larvae
BMC Neuroscience (2016)
-
A zebrafish mosaic assay to study mammalian stem cells in real time in vivo
Journal of Molecular Histology (2016)