Abstract
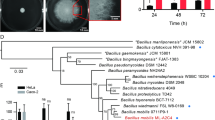
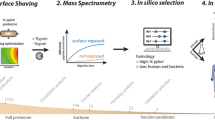
We have used DNA microarrays to follow Neisseria meningitidis serogroup B (MenB) gene regulation during interaction with human epithelial cells. Host-cell contact induced changes in the expression of 347 genes, more than 30% of which encode proteins with unknown function. The upregulated genes included transporters of iron, chloride, amino acids, and sulfate, many virulence factors, and the entire pathway of sulfur-containing amino acids. Approximately 40% of the 189 upregulated genes coded for peripherally located proteins, suggesting that cell contact promoted a substantial reorganization of the cell membrane. This was confirmed by fluorescence activated cell sorting (FACS) analysis on adhering bacteria using mouse sera against twelve adhesion-induced proteins. Of the 12 adhesion-induced surface antigens, 5 were able to induce bactericidal antibodies in mice, demonstrating that microarray technology is a valid approach for identifying new vaccine candidates and nicely complements other genome mining strategies used for vaccine discovery.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Gotschlich, E.C., Liu, T.Y. & Artenstein, M.S. Human immunity to the meningococcus. 3. Preparation and immunochemical properties of the group A, group B, and group C meningococcal polysaccharides. J. Exp Med. 129, 1349–1365 (1969).
Naess, A. et al. Sequelae one year after meningococcal disease. Acta Neurol. Scand. 89, 139–142 (1994).
Nassif, X., Pujol, C., Morand, P. & Eugène, E. Interaction of pathogenic Neisseria with host cells. Is it possible to assemble the puzzle? Mol. Microbiol. 32, 1121–1132 (1999).
Merz, A.J. & So, M. Interaction of pathogenic Neisseriae with epithelial cell membranes. Annu. Rev. Cell. Dev. Biol. 16, 423–457 (2000).
Mahan, M.J., Slauch, J.M. & Mekalanos, J.J. Selection of bacterial virulence genes that are specifically induced in host tissues. Science 259, 686–688 (1993).
Hensel, M. et al. Simultaneous identification of bacterial virulence genes by negative selection. Science 269, 400–403 (1995).
Schena, M., Shalon, D., Davs, R.W. & Brown, P.O. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science 270, 467–470 (1995).
Pandey, A. & Mann, M. Proteomics to study genes and genomes. Nature 405, 837–846 (2000).
Tettelin, H. et al. Complete genome sequence of Neisseria meningitidis serogroup B strain MC58. Science 287, 1809–1815 (2000).
Sun, Y.-H., Bakshi, S., Chalmers, R. & Tang, C.M. Functional genomics of Neisseria meningitidis pathogenesis. Nat. Med. 6, 1269–1273 (2000).
Zollinger, W.D. New and improved vaccines against meningococcal disease. in New Generation Vaccines (eds Levine, M.M., Wodrow, G.C., Kaper, J.B. & Cobon, G.S.) 469–488 (Dekker, New York, 1997).
Martin, D., Cadieux, N., Hamel, J. & Brodeur, B.R. Highly conserved Neisseria meningitidis surface protein confers protection against experimental infection. J. Exp. Med. 185, 1173–1183 (1997).
Pizza, M.G. et al. Whole genome sequencing to identify vaccines candidates against serogroup B meningococcus. Science 287, 1816–1820 (2000).
Pujol, C., Eugène, E., de Saint Martin, L. & Nassif, X. Interaction of Neisseria meningitidis with a polarized monolayer of epithelial cells. Infect. Immun. 65, 4836–4842 (1997).
Deghmane, Al.-E. et al. Intimate adhesion of Neisseria meningitidis to human epithelial cells is under the control of the crgA gene, a novel Lys-R type transcriptional regulator. EMBO J. 19, 1068–1078 (2000).
Pujol, C., Eugène, E., Marceau, M. & Nassif, X. The meningococcal pilT protein is required for induction of intimate attachment to epithelial cells following pilus-mediated adhesion. Proc. Nat. Acad. Sci. USA 96, 4017–4022 (1999).
Shea, J.E., Santangelo, J.D. & Feldman, R. Signature-tagged mutagenesis in the identification of virulence genes in pathogens. Curr. Opin. Microbiol. 3, 451–458 (2000).
Saunders, N.J. et al. Repeat–associated phase variable genes in the complete genome sequence of Neisseria meningitidis strain MC58. Mol. Microbiol. 37, 207–215 (2000).
Braaten, B., Nou, Y., Kaltenbach, L. & Low, D. Methylation patterns in pap regulatory DNA control pyelonephritis-associated pili phase variation in E. coli. Cell 76, 577–588 (1994).
Heithoff, D.M., Sinsheimer, R.L., Low, D.A. & Mahan, M.J. An essential role for DNA adenine methylation in bacterial virulence. Science 284, 967–970 (1999).
Garcia-Del Portillo, F., Pucciarelli, M.G. & Casadesus, J. DNA adenine methylase mutants of Salmonella typhimurium show defects in protein secretion, cell invasion, and M cell cytotoxicity. Proc. Natl. Acad. Sci. USA 96, 11578–11583 (1999).
Klebanoff, S.J., Locksley, R.M., Jong, E.C. & Rosen, H. Oxidative response of phagocytes to parasitic invasion. Ciba Found. Symp. 99, 92–112 (1983).
Ramarao, N., Gray-Owen, S.D. & Meyer, T.F. Helicobacter pylori induces but survives the extracellular release of oxygen radicals from professional phagocytes using its catalase activity. Mol. Microbiol. 38, 103–113 (2000).
Lottenberg, R. et al. Cloning, sequence analysis, and expression in Escherichia coli of a streptococcal plasmin receptor. J. Bacteriol. 174, 5204–5210 (1992).
De Matteis, M.A. et al. Stimulation of endogenous ADP-ribosylation by brefeldin A. Proc. Natl. Acad. Sci. USA 91, 1114–1118 (1994).
Pancholi, V. & Fischetti, V.A. Regulation of the phosphorylation of human pharyngeal cell proteins by group A streptococcal surface dehydrogenase: signal transduction between streptococci and pharyngeal cells. J. Exp. Med. 186, 1633–1643 (1997).
Winram, S.B. & Lottemberg, R. The plasmin-binding protein Plr of group A streptococci is identified as glyceraldheyde-3-phosphate dehydrogenase. Microbiology 142, 2311–2320 (1996).
Zhang, Y.L., Ong, C.T. & Leung, K.Y. Molecular analysis of genetic differences between virulent and avirulent strains of Aeromonas hydrophila isolated from diseased fish. Microbiology 146, 999–1009 (2000).
Zhang, H.Z. & Donnenberg, M.S. DsbA is required for stability of the type IV pilin of enteropathogenic Escherichia coli. Mol. Microbiol. 21, 787–797 (1996).
Yu, I., Edwards-Jones, B., Neyrolles, O. & Kroll, J.S. Key role for DsbA in cell-to-cell spread of Shigella flexneri, permitting secretion of Ipa proteins to interepithelial protrusions. Infect. Immun. 68, 6449–6456 (2000).
Susa, M., Hacker, J. & Marre, R. De novo synthesis of Legionella pneumophila antigens during intracellular growth in phagocytic cells. Infect. Immun. 64, 1679–1684 (1996).
Wintemeyer, E. et al. Influence of site specifically altered Mip proteins on intracellular survival of Legionella pneumophila in eukaryotic cells. Infect. Immun. 63, 4576–4583 (1995).
Horne, S.M., Kottom, T.J., Nolan, L.K. & Young, K.D. Decreased intracellular survival of an fkpA mutant of Salmonella typhimurium Copenhagen. Infec. Immun. 65, 806–810 (1997).
Adegbola, R.A., Mulholland, E.K., Secka, O., Jaffar, S. & Greenwood, B.M. Vaccination with a Haemophilus influenzae Type b conjugate vaccine reduces oropharyngeal carriage of H. influenzae Type b among Gambian children. J. Infec. Immun. 177, 1758–1761 (1998).
Dagan, R. et al. Reduction of nasopharyngeal carriage of Streptococcus pneumoniae after administration of a 9-valent pneumococcal conjugate vaccine to toddlers attending day care centers. J. Infec. Dis. 185, 927–936 (2002).
Grandi, G. Antibacterial vaccine design using genomics and proteomics. Trends Biotechnol. 19, 181–188 (2001).
Cummings, C.A. & Relman, D.A. Using DNA microarrays to study host microbe interaction. Emerg. Infect. Dis. 6, 513–522 (2000).
Belcher, C.E. et al. The transcriptional response of respiratory epithelial cells to Berdetella pertussis reveal host defensive and pathogen counter defensive strategies. Proc. Natl. Acad. Sci. USA 97, 13847–13857 (2000).
Rappuoli, R. Pushing the limits of cellular microbiology: microarrays to study bacteria-host cell intimate contacts. Proc. Natl. Acad. Sci. USA 97, 13468–13469 (2000).
Rappuoli, R. Reverse vaccinology. Curr. Opin. Microbiol. 3, 445–450 (2000).
Mathiassen, S., Lauemoller, S.L., Ruhwald, M., Claesson, M.H. & Buus, S. Tumor-associated antigens identified by mRNA expression profiling induce protective anti-tumor immunity. Eur. J. Immunol. 31, 1239–1246 (2001).
Etz, H. et al. Identification of in vivo expressed vaccine candidate antigens from Staphylococcus aureus. Proc. Natl. Acad. Sci. USA 99, 6573–6578 (2002).
Long, A.D. et al. Improved statistical inference from DNA microarray data using analysis of variance and a Bayesian statistical framework. Analysis of global gene expression in Escherichia coli K12. J. Biol. Chem. 276, 19937–19944 (2001).
Trieu-Cuot, P., Poyart-Salmeron, C., Carlier, C. & Courvalin, P. Nucleotide sequence of the erythromycin resistance gene of the conjugative transposon Tn1545. Nucleic Acids Res. 18, 3660 (1990).
Trieu-Cuot, P., Gerbaud, G., Lambert, T. & Courvalin, P. In vivo transfer of genetic information between gram-positive and gram-negative bacteria. EMBO J. 4, 3583–3587 (1985).
Peeters, C.C. et al. Immunogenicity of various presentation forms of PorA outer membrane protein of Neisseria meningitidis in mice. Vaccine 17, 2702–2712 (1999).
Seiler, A., Reichardt, R., Sarkari, J., Caugant, D.A. & Achtman, M. Allelic polymorphism and site specific recombination in the opc locus of Neisseria meningitids. Mol. Microbiol. 19, 841–856 (1996).
Acknowledgements
We thank J. Telford for the critical reading of the manuscript and R. Moxon and D.A. Caugant for providing bacterial strains. We also thank M. Comanducci for providing genomic DNAs, V. Masignani, M. Scarselli, and R. Beltrami for assistance in computer analysis, S. Censini and S. Guidotti for nucleotide sequencing, G. Corsi for artwork, and A. Maiorino for expert secretarial assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Grifantini, R., Bartolini, E., Muzzi, A. et al. Previously unrecognized vaccine candidates against group B meningococcus identified by DNA microarrays. Nat Biotechnol 20, 914–921 (2002). https://doi.org/10.1038/nbt728
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt728
This article is cited by
-
Association between antibodies against group B Streptococcus surface proteins and recto-vaginal colonisation during pregnancy
Scientific Reports (2017)
-
TolC plays a crucial role in immune protection conferred by Edwardsiella tarda whole-cell vaccines
Scientific Reports (2016)
-
Burkholderia pseudomallei transcriptional adaptation in macrophages
BMC Genomics (2012)
-
The role of glyceraldehyde 3-phosphate dehydrogenase (GapA-1) in Neisseria meningitidis adherence to human cells
BMC Microbiology (2010)