Abstract
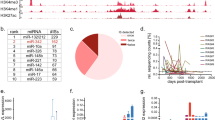
We have shown previously that transgene expression can be suppressed in hematopoietic cells using vectors that are responsive to microRNA (miRNA) regulation. Here we investigate the potential of this approach for more sophisticated control of transgene expression. Analysis of the relationship between miRNA expression levels and target mRNA suppression suggested that suppression depends on a threshold miRNA concentration. Using this information, we generated vectors that rapidly adjust transgene expression in response to changes in miRNA expression. These vectors sharply segregated transgene expression between closely related states of therapeutically relevant cells, including dendritic cells, hematopoietic and embryonic stem cells, and their progeny, allowing positive/negative selection according to the cells' differentiation state. Moreover, two miRNA target sites were combined to restrict transgene expression to a specific cell type in the liver. Notably, the vectors did not detectably perturb endogenous miRNA expression or regulation of natural targets. The properties of miRNA-regulated vectors should allow for safer and more effective therapeutic applications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Singec, I., Jandial, R., Crain, A., Nikkhah, G. & Snyder, E.Y. The leading edge of stem cell therapeutics. Annu. Rev. Med. 58, 313–328 (2007).
Aiuti, A. et al. Correction of ADA-SCID by stem cell gene therapy combined with nonmyeloablative conditioning. Science 296, 2410–2413 (2002).
Gaspar, H.B. et al. Gene therapy of X-linked severe combined immunodeficiency by use of a pseudotyped gammaretroviral vector. Lancet 364, 2181–2187 (2004).
Hacein-Bey-Abina, S. et al. Sustained correction of X-linked severe combined immunodeficiency by ex vivo gene therapy. N. Engl. J. Med. 346, 1185–1193 (2002).
Ott, M.G. et al. Correction of X-linked chronic granulomatous disease by gene therapy, augmented by insertional activation of MDS1–EVI1, PRDM16 or SETBP1. Nat. Med. 12, 401–409 (2006).
Goverdhana, S. et al. Regulatable gene expression systems for gene therapy applications: progress and future challenges. Mol. Ther. 12, 189–211 (2005).
Haviernik, P. & Bunting, K.D. Safety concerns related to hematopoietic stem cell gene transfer using retroviral vectors. Curr. Gene Ther. 4, 263–276 (2004).
Hawley, T.S., Fong, A.Z., Griesser, H., Lyman, S.D. & Hawley, R.G. Leukemic predisposition of mice transplanted with gene-modified hematopoietic precursors expressing flt3 ligand. Blood 92, 2003–2011 (1998).
Brown, B.D. & Lillicrap, D. Dangerous liaisons: the role of “danger” signals in the immune response to gene therapy. Blood 100, 1133–1140 (2002).
Follenzi, A. et al. Targeting lentiviral vector expression to hepatocytes limits transgene-specific immune response and establishes long-term expression of human antihemophilic factor IX in mice. Blood 103, 3700–3709 (2004).
Yuasa, K. et al. Adeno-associated virus vector-mediated gene transfer into dystrophin-deficient skeletal muscles evokes enhanced immune response against the transgene product. Gene Ther. 9, 1576–1588 (2002).
Sadeghi, H. & Hitt, M.M. Transcriptionally targeted adenovirus vectors. Curr. Gene Ther. 5, 411–427 (2005).
Waehler, R., Russell, S.J. & Curiel, D.T. Engineering targeted viral vectors for gene therapy. Nat. Rev. Genet. 8, 573–587 (2007).
De Palma, M. et al. Promoter trapping reveals significant differences in integration site selection between MLV and HIV vectors in primary hematopoietic cells. Blood 105, 2307–2315 (2005).
ENCODE Project Consortium. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447, 799–816 (2007).
Ambros, V. microRNAs: tiny regulators with great potential. Cell 107, 823–826 (2001).
Bartel, D.P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Bartel, D.P. & Chen, C.Z. Micromanagers of gene expression: the potentially widespread influence of metazoan microRNAs. Nat. Rev. Genet. 5, 396–400 (2004).
Lagos-Quintana, M. et al. Identification of tissue-specific microRNAs from mouse. Curr. Biol. 12, 735–739 (2002).
Landgraf, P. et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 129, 1401–1414 (2007).
Plasterk, R.H. Micro RNAs in animal development. Cell 124, 877–881 (2006).
Hobert, O. Common logic of transcription factor and microRNA action. Trends Biochem. Sci. 29, 462–468 (2004).
Xiao, C. et al. MiR-150 controls B cell differentiation by targeting the transcription factor c-Myb. Cell 131, 146–159 (2007).
Brown, B.D., Venneri, M.A., Zingale, A., Sergi Sergi, L. & Naldini, L. Endogenous microRNA regulation suppresses transgene expression in hematopoietic lineages and enables stable gene transfer. Nat. Med. 12, 585–591 (2006).
Mansfield, J.H. et al. MicroRNA-responsive 'sensor' transgenes uncover Hox-like and other developmentally regulated patterns of vertebrate microRNA expression. Nat. Genet. 36, 1079–1083 (2004).
Cao, X., Pfaff, S.L. & Gage, F.H. A functional study of miR-124 in the developing neural tube. Genes Dev. 21, 531–536 (2007).
Didiano, D. & Hobert, O. Perfect seed pairing is not a generally reliable predictor for miRNA-target interactions. Nat. Struct. Mol. Biol. 13, 849–851 (2006).
Amendola, M., Venneri, M.A., Biffi, A., Vigna, E. & Naldini, L. Coordinate dual-gene transgenesis by lentiviral vectors carrying synthetic bidirectional promoters. Nat. Biotechnol. 23, 108–116 (2005).
Steiner, F.A. et al. Structural features of small RNA precursors determine Argonaute loading in Caenorhabditis elegans. Nat. Struct. Mol. Biol. 14, 927–933 (2007).
Forstemann, K., Horwich, M.D., Wee, L., Tomari, Y. & Zamore, P.D. Drosophila microRNAs are sorted into functionally distinct argonaute complexes after production by dicer-1. Cell 130, 287–297 (2007).
Bhattacharyya, S.N., Habermacher, R., Martine, U., Closs, E.I. & Filipowicz, W. Relief of microRNA-mediated translational repression in human cells subjected to stress. Cell 125, 1111–1124 (2006).
Leung, A.K. & Sharp, P.A. microRNAs: a safeguard against turmoil? Cell 130, 581–585 (2007).
Fazi, F. et al. A minicircuitry comprised of microRNA-223 and transcription factors NFI-A and C/EBPalpha regulates human granulopoiesis. Cell 123, 819–831 (2005).
Cimmino, A. et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc. Natl. Acad. Sci. USA 102, 13944–13949 (2005).
Doench, J.G., Petersen, C.P. & Sharp, P.A. siRNAs can function as miRNAs. Genes Dev. 17, 438–442 (2003).
Banchereau, J. & Palucka, A.K. Dendritic cells as therapeutic vaccines against cancer. Nat. Rev. Immunol. 5, 296–306 (2005).
Steinman, R.M., Hawiger, D. & Nussenzweig, M.C. Tolerogenic dendritic cells. Annu. Rev. Immunol. 21, 685–711 (2003).
Taganov, K.D., Boldin, M.P., Chang, K.J. & Baltimore, D. NF-kappaB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc. Natl. Acad. Sci. USA 103, 12481–12486 (2006).
O'Connell, R.M., Taganov, K.D., Boldin, M.P., Cheng, G. & Baltimore, D. MicroRNA-155 is induced during the macrophage inflammatory response. Proc. Natl. Acad. Sci. USA 104, 1604–1609 (2007).
Chen, C.Z., Li, L., Lodish, H.F. & Bartel, D.P. MicroRNAs modulate hematopoietic lineage differentiation. Science 303, 83–86 (2004).
Fukao, T. et al. An evolutionarily conserved mechanism for microRNA-223 expression revealed by microRNA gene profiling. Cell 129, 617–631 (2007).
Suh, M.R. et al. Human embryonic stem cells express a unique set of microRNAs. Dev. Biol. 270, 488–498 (2004).
Grimm, D. et al. Fatality in mice due to oversaturation of cellular microRNA/short hairpin RNA pathways. Nature 441, 537–541 (2006).
Koller, E. et al. Competition for RISC binding predicts in vitro potency of siRNA. Nucleic Acids Res. 34, 4467–4476 (2006).
Chen, K. & Rajewsky, N. The evolution of gene regulation by transcription factors and microRNAs. Nat. Rev. Genet. 8, 93–103 (2007).
Hutvagner, G. & Zamore, P.D. A microRNA in a multiple-turnover RNAi enzyme complex. Science 297, 2056–2060 (2002).
Liu, J. et al. Argonaute2 is the catalytic engine of mammalian RNAi. Science 305, 1437–1441 (2004).
Haley, B. & Zamore, P.D. Kinetic analysis of the RNAi enzyme complex. Nat. Struct. Mol. Biol. 11, 599–606 (2004).
Li, Q.J. et al. miR-181a is an intrinsic modulator of T cell sensitivity and selection. Cell 129, 147–161 (2007).
Ambros, V. & Chen, X. The regulation of genes and genomes by small RNAs. Development 134, 1635–1641 (2007).
Tomari, Y., Du, T. & Zamore, P.D. Sorting of Drosophila small silencing RNAs. Cell 130, 299–308 (2007).
Griffiths-Jones, S., Grocock, R.J., van Dongen, S., Bateman, A. & Enright, A.J. miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res. 34, D140–D144 (2006).
Baskerville, S. & Bartel, D.P. Microarray profiling of microRNAs reveals frequent coexpression with neighboring miRNAs and host genes. RNA 11, 241–247 (2005).
Yang, L. & Baltimore, D. Long-term in vivo provision of antigen-specific T cell immunity by programming hematopoietic stem cells. Proc. Natl. Acad. Sci. USA 102, 4518–4523 (2005).
Brennecke, J., Stark, A., Russell, R.B. & Cohen, S.M. Principles of microRNA-target recognition. PLoS Biol. 3, e85 (2005).
Grimson, A. et al. MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol. Cell 27, 91–105 (2007).
Neilson, J.R., Zheng, G.X., Burge, C.B. & Sharp, P.A. Dynamic regulation of miRNA expression in ordered stages of cellular development. Genes Dev. 21, 578–589 (2007).
Chen, C. et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res. 33, e179 (2005).
De Palma, M., Venneri, M.A., Roca, C. & Naldini, L. Targeting exogenous genes to tumor angiogenesis by transplantation of genetically modified hematopoietic stem cells. Nat. Med. 9, 789–795 (2003).
Acknowledgements
HUES were kindly provided by Doug Melton from Harvard Stem Cell Institute (Cambridge, MA, USA), under specific Material Transfer Agreement to C.G. We are grateful to Irene Bozzoni, Desiree Bonci and Roger Tsien for providing reagents, Silvia Gregori and Daniela Tomasoni for human monocytes, Angelo Lombardo, Michele De Palma and Roberta Mazzieri for helpful discussions, and Lucia Sergi Sergi and Giulia Schira for technical help. We would also like to thank the San Raffaele Centre of Statistics for Biomedical Sciences for suggestions on data analysis. This work was supported by grants from Telethon (TIGET grant), EU (Projects LSHB-CT-2004-005276, RIGHT and LSHB-CT-2004-005242, CONSERT) and the Italian Ministry of Scientific Research to L.N., and EU Grant LSB-CT-2004-503257 to G.L. B.D.B. is the recipient of a Natural Science and Engineering Research Council of Canada (NSERC) fellowship. B.G. is the recipient of a research fellowship from the German Research Foundation (DFG Forschungsstipendium).
Author information
Authors and Affiliations
Contributions
B.D.B. designed and performed research and analyzed data concerning vector design, miRNA regulation and dendritic cells, and wrote the paper. B.G. designed and performed research and analyzed data on HS cells and differentiation state-specific and suicide vectors, and wrote the paper. A.C. performed research and analyzed data concerning vector design and miRNA regulation. S.C. and G.L. performed research and analyzed data regarding the ES cell studies. M.A., A.Z. and A.B. performed research. C.G. coordinated ES cell work. L.N. coordinated the project, designed research, analyzed data and wrote the paper.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figs. 1–7; Supplementary Table 1; Supplementary Materials and Methods (PDF 1068 kb)
Rights and permissions
About this article
Cite this article
Brown, B., Gentner, B., Cantore, A. et al. Endogenous microRNA can be broadly exploited to regulate transgene expression according to tissue, lineage and differentiation state. Nat Biotechnol 25, 1457–1467 (2007). https://doi.org/10.1038/nbt1372
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1372
This article is cited by
-
Insight into extracellular vesicles in vascular diseases: intercellular communication role and clinical application potential
Cell Communication and Signaling (2023)
-
Development of microglia-targeting adeno-associated viral vectors as tools to study microglial behavior in vivo
Communications Biology (2022)
-
Mapping and targeted viral activation of pancreatic nerves in mice reveal their roles in the regulation of glucose metabolism
Nature Biomedical Engineering (2022)
-
In vivo self-assembled small RNAs as a new generation of RNAi therapeutics
Cell Research (2021)
-
Regulated control of gene therapies by drug-induced splicing
Nature (2021)