Abstract
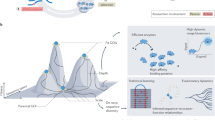
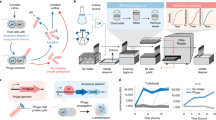
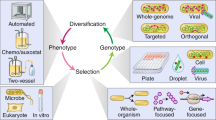
We describe an efficient method for generating combinatorial libraries with a high percentage of unique and functional mutants. Combinatorial libraries have been successfully used in the past to express ensembles of mutant proteins in which all possible amino acids are encoded at a few positions in the sequence. However, as more positions are mutagenized the proportion of functional mutants is expected to decrease exponentially. Small groups of residues were randomized in parallel to identify, at each altered position, amino acids which lead to functional proteins. By using optimized nucleotide mixtures deduced from the sequences selected from the random libraries, we have simultaneously altered 16 sites in a model pigment binding protein: approximately one percent of the observed mutants were functional. Mathematical formalization and extrapolation of our experimental data suggests that a 107-fold increase in the throughput of functional mutants has been obtained relative to the expected frequency from a random combinatorial library. Exponential ensemble mutagenesis should be advantageous in cases where many residues must be changed simultaneously to achieve a specific engineering goal, as in the combinatorial mutagenesis of phage displayed antibodies. With the enhanced functional mutant frequencies obtained by this method, entire proteins could be mutagenized combinatorially.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Oliphant, A.R., Nussbaum, A.L. and Struhl, K. 1986. Cloning of random sequence Oligonucleotides. Gene 44: 177–183.
Reidhaar-Olson, J.F., Bowie, J.U., Breyer, R.M., Hu, J.C., Knight, K.L., Lim, W.A., Mossing, M.C., Parsell, D.A., Shoemaker, K.R. and Sauer, R.T. 1991. Random mutagenesis of protein sequences using oligonucleotide cassettes. Methods in Enzymol. 208: 564–587.
Gram, H., Marconi, L.-A., Barbas, C.F., Collet, T.A., Lerner, R.A. and Kang, A.S. 1992. In vitro selection and affinity maturation of antibodies from a naive combinatorial immunoglobulin library. Proc. Natl. Acad. Sci. USA 89: 3576–3580.
Barbas, C.F., Bain, J.D., Hoekstra, D.M. and Lerner, R.A. 1992. Scmisynthetic combinatorial antibody libraries: A chemical solution to the diversity problem. Proc. Natl. Acad. Sci. USA 89: 4457–4461.
Lam, K.S., Salmon, S.E., Hersh, E.M., Hruby, V.J., Kazmiersky, W.M. and Knapp, R.J. 1991. A new type of synthetic peptide library for identifying ligand-binding activity. Nature 354: 82–84.
Houghten, R.A., Pinilla, C., Blondelle, S.E., Appel, J.R., Dooley, C.T. and Cuervo, J.H. 1991. Generation and use of synthetic peptide combinatorial libraries for basic research and drug discovery. Nature 354: 84–86.
Roberts, B.L., Markland, W., Ley, A.C., Kent, R.B., White, D.W., Guterman, S.K. and Ladner, R.C. 1992. Directed evolution of a protein: Selection of potent neutrophil elastase inhibitors displayed on M13 fusion phage. Proc. Natl. Acad. Sci. USA 89: 2429–2433.
Lowman, H.B., Bass, S.H., Simpson, N. and Wells, J.A. 1991. Selecting high-affinity binding proteins by monovalcnt phage display. Biochemistry 30: 10832–10838.
Smith, G.P. 1985. Filamentous fusion phage: Novel expression vectors that display cloned antigens on the virion surface. Science 228: 1315–1317.
Hoogenboom, H.R., Griffiths, A.D., Johnson, K.S., Chiswell, D.J., Hudson, P. and Winter, G. 1991. Multi-subunit proteins on the surface of filamentous phage: Methodologies for displaying antibody (Fab) heavy and light chains. Nucleic Acids Res. 19: 4133–4137.
Kang, A.S., Barbas, C.F., Janda, K.D., Benkovic, S.J. and Lerner, R.A. 1991. Linkage of recognition and replication functions by assembling combinatorial antibody Fab libraries on phage surfaces. Proc. Natl. Acad. Sci. USA 88: 4363–4366.
Youvan, D.C. and Ismail, S. 1985. Light-harvesting II B800–B850 complex structural genes from Rhodopseudomonas capsulata. Proc. Natl. Acad. Sci. USA 82: 63–67.
Zuber, H. 1990. Consideration on the structural principles of the antenna complexes of phototrophic bacteria, p. 161–180. In: Molecular Biology of Membrane-Bound Complexes in Phototrophic Bacteria. G. Drews and E. A. Dawes (Eds.). Plenum Press, New York.
Arkin, A.P., Goldman, E.R., Robles, S.J., Coleman, W., Goddard, C.A., Yang, M.M. and Youvan, D.C. 1990. Applications of imaging spectroscopy in molecular biology: Colony screening based on absorption spectra. Bio/Technology 8: 746–749.
Yang, M.M. and Youvan, D.C. 1988. Applications of imaging spectroscopy in molecular biology. I. Screening photosynthetic bacteria. Bio/Technology 6: 939–942.
Youvan, D.C., Goldman, E.R., Delagrave, S. and Yang, M.M. 1993. Digital imaging spectroscopy for massively parallel screening of mutants. Methods in Enzymol. In press.
Goldman, E.R. and Youvan, D.C. An algorithmically optimized combinatorial library screened by digital imaging spectroscopy. Bio/Technology 10: 1557–1561.
Delagrave, S., Goldman, E.R. and Youvan, D.C. 1993 Recursive ensemble mutagenesis. Prot. Engng. 6: 327–331.
Arkin, A.P. and Youvan, D.C. 1992. A combinatorial optimization procedure for protein engineering: Simulation of recursive ensemble mutagenesis. Proc. Natl. Acad. Sci. USA 89: 7811–7815.
Arkin, A.P. and Youvan, D.C. 1992. Optimizing nucleotide mixtures to encode specific subsets of amino acids for semi-random mutagenesis. Bio/Technology 10: 297–300.
Youvan, D.C., Arkin, A.P. and Yang, M.M. 1992. Recursive ensemble mutagenesis: A combinatorial optimization technique for protein engineering, p. 401–410. In: Parallel Problem Solving from Nature, 2. R. Maenner and B. Manderick (Eds.). Elsevier Publishing Co., Amsterdam.
Lim, W.A. and Sauer, R.T. 1991. The role of internal packing interactions in determining the structure and stability of a protein. J. Mol. Biol. 219: 359–376.
Robles, S.J., Ranck, T. and Youvan, D.C. 1992. Symmetrical intragenic suppressors of the bacterial reaction center cd-helix exchange mutants, p. 21–23. In: Structure of the Bacterial Photosynthetic Reaction Center (II). J. Breton and A. Vermeglio (Eds.). Plenum, New York.
Wells, J.A. 1990. Additivity of mutational effects in proteins. Biochemistry 29: 8509–8516.
Robles, S.J. and Youvan, D.C. 1993. Hydropathy and molar volume constraints on combinatorial mutants of the photosynthetic reaction center. J. Mol. Biol. 232: 242–252.
Chothia, C., Lesk, A.M., Gherardi, E., Tomlinson, I.M., Walter, G., Marks, J.D., Llewelyn, M.B. and Winter, G. 1992. Structural repertoire of the human Vh segments. J. Mol. Biol. 227: 799–817.
Burton, D.R., Barbas, C.F, Persson, M.A.A., Koenig, S., Chanock, R.M. and Lerner, R.A. 1992. A large array of human monoclonal antibodies to type 1 human immunodeficiency virus from combinatorial libraries of asymptomatic seropositive individuals. Proc. Natl. Acad. Sci. USA 88: 10134–10137.
Sambrook, J., Pritsch, E.F. and Maniatis, T. 1990. Cloning: A Laboratory Manual, 2nd Edition. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Delagrave, S., Youvan, D. Searching Sequence Space to Engineer Proteins: Exponential Ensemble Mutagenesis. Nat Biotechnol 11, 1548–1552 (1993). https://doi.org/10.1038/nbt1293-1548
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nbt1293-1548