Abstract
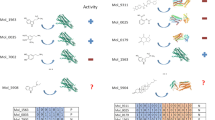
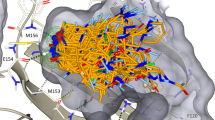
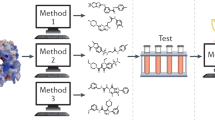
Lead generation is a major hurdle in small-molecule drug discovery, with an estimated 60% of projects failing from lack of lead matter or difficulty in optimizing leads for drug-like properties. It would be valuable to identify these less-druggable targets before incurring substantial expenditure and effort. Here we show that a model-based approach using basic biophysical principles yields good prediction of druggability based solely on the crystal structure of the target binding site. We quantitatively estimate the maximal affinity achievable by a drug-like molecule, and we show that these calculated values correlate with drug discovery outcomes. We experimentally test two predictions using high-throughput screening of a diverse compound collection. The collective results highlight the utility of our approach as well as strategies for tackling difficult targets.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Brown, D. & Superti-Furga, G. Rediscovering the sweet spot in drug discovery. Drug Discov. Today 8, 1067–1077 (2003).
Hopkins, A.L. & Groom, C.R. The druggable genome. Nat. Rev. Drug Disc. 1, 727–730 (2002).
Fauman, E.B., Hopkins, A. & Groom, C.R. Structural Bioinformatics in Drug Discovery. in Structural Bioinformatics (eds. Weissig, H. & Bourne, P.) 477–498 (Wiley-Liss, Hoboken, NJ 2003).
Vassilev, L.T. et al. In vivo activation of the p53 pathway by small-molecule antagonists of MDM2. Science 303, 844–848 (2004).
Lipinski, C.A., Lombardo, F., Dominy, B.W. & Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 46, 3–26 (2001).
Palm, K., Stenberg, P., Luthman, K. & Artursson, P. Polar molecular surface properties predict the intestinal absorption of drugs in humans. Pharm. Res. 14, 568–571 (1997).
Veber, D.F. et al. Molecular properties that influence the oral bioavailability of drug candidates. J. Med. Chem. 45, 2615–2623 (2002).
Wermuth, C.G. The Practice of Medicinal Chemistry, edn. 2 (Academic Press, London, UK and San Diego, 2003)
Kuntz, I.D., Chen, K., Sharp, K.A. & Kollman, P.A. The maximal affinity of ligands. Proc. Natl. Acad. Sci. USA 96, 9997–10002 (1999).
Chothia, C. Hydrophobic bonding and accessible surface area in proteins. Nature 248, 338–339 (1974).
Karplus, P.A. Hydrophobicity regained. Protein Sci. 6, 1302–1307 (1997).
Sharp, K.A., Nicholls, A., Fine, R.F. & Honig, B. Reconciling the magnitude of the microscopic and macroscopic hydrophobic effects. Science 252, 106–109 (1991).
Cheng, Y-K. & Rossky, P.J. Surface topography dependence of biomolecular hydrophobic hydration. Nature 392, 696–699 (1998).
Southall, N.T. & Dill, K.A. The mechanism of hydrophobic solvation depends on solute radius. J. Phys. Chem. B 104, 1326–1331 (2000).
De Young, L.R. & Dill, K.A. Partitioning of nonpolar solutes into bilayers and amorphous n-Alkanes. J. Phys. Chem. 94, 801–809 (1990).
Hendsch, Z.S. & Tidor, B. Do salt bridges stabilize proteins? A continuum electrostatic analysis. Protein Sci. 3, 211–226 (1994).
Coleman, R.G., Burr, M.A., Souvaine, D.L. & Cheng, A.C. An intuitive approach to measuring protein surface curvature. Proteins 61, 1068–1074 (2005).
Liang, J., Edelsbrunner, H. & Woodward, C. Anatomy of protein pockets and cavities: measurement of binding site geometry and implications for ligand design. Protein Sci. 7, 1884–1897 (1998).
Ettmayer, P., Amidon, G.L., Clement, B. & Testa, B. Lessons learned from marketed and investigational prodrugs. J. Med. Chem. 47, 2393–2404 (2004).
Kim, C.U. et al. Influenza neuraminidase inhibitors possessing a novel hydrophobic interaction in the enzyme active site: design, synthesis, and structural analysis of carbocyclic sialic acid analogues with potent anti-influenza activity. J. Am. Chem. Soc. 119, 681–690 (1997).
Teague, S.J. Implications of protein flexibility for drug discovery. Nat. Rev. Drug Disc. 2, 527–541 (2003).
Arkin, M.R. & Wells, J.A. Small-molecule inhibitors of protein–protein interactions: progressing towards the dream. Nat. Rev. Drug Disc. 3, 301–317 (2004).
Davis, A.M. & Teague, S.J. Hydrogen bonding, hydrophobic interactions, and failure of the rigid receptor hypothesis. Agnew. Chem. Int. Ed. 38, 736– 749 (1999).
Nayal, M. & Honig, B. On the nature of cavities on protein surfaces: application to the identification of drug binding sites. Proteins 63, 892–906 (2006).
Hajduk, P.J., Huth, J.R. & Fesik, S.W. Druggability indices for protein targets derived from NMR-based screening data. J. Med. Chem. 48, 2518–2525 (2005).
Deininger, M., Buchdunger, E. & Druker, B.J. The development of imatinib as a therapeutic agent for chronic myeloid leukemia. Blood 105, 2640–2653 (2005).
Istvan, E.S. & Deisenhofer, J. Structural mechanism for statin inhibition of HMG-CoA reductase. Science 292, 1160–1164 (2001).
Schachter, M. Chemical, pharmacokinetic and pharmacodynamic properties of statins: an update. Fundam. Clin. Pharmacol. 19, 117–125 (2004).
Armstrong, K.A., Tidor, B. & Cheng, A.C. Optimal charges in lead progression: a structure-based neuraminidase case study. J. Med. Chem. 49, 2470–2477 (2006).
Jacques, S.L. et al. Characterization of yeast homoserine dehydrogenase, an antifungal target: the invariant histidine 309 is important for enzyme integrity. Biochim. Biophys. Acta 1544, 28–41 (2001).
Nissink, J.W. et al. A new test for validating predictions of protein ligand interaction. Proteins 49, 457–471 (2002).
Acknowledgements
We thank colleagues across Pfizer Global R&D for discussions, and Jill Milne and Ralph Lambalot for supporting the HTS experiment. A.C.C. additionally thanks Ken Dill for advice. This work was entirely funded by Pfizer.
Author information
Authors and Affiliations
Contributions
A.C.C. developed the method, designed experiments and analyzed data and wrote the manuscript; R.G.C. developed computational geometry algorithms and analyzed data; K.T.S. helped design the HTS experiment, performed screening for HSD, and analyzed results of the screening comparison; P.S. helped design the HTS experiment, developed the H-PGDS assay and performed screening; Q.C. and D.R.C. helped analyze data for Figures 1 and 3a; A.C.S. helped implement computational methods; E.S.H. discussed results and helped design the HTS experiment. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
Details of targets used in study.
Supplementary Table 2
References and notes for best affinity values.
Rights and permissions
About this article
Cite this article
Cheng, A., Coleman, R., Smyth, K. et al. Structure-based maximal affinity model predicts small-molecule druggability. Nat Biotechnol 25, 71–75 (2007). https://doi.org/10.1038/nbt1273
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1273
This article is cited by
-
Predicting drug–protein interactions by preserving the graph information of multi source data
BMC Bioinformatics (2024)
-
Drug–target affinity prediction with extended graph learning-convolutional networks
BMC Bioinformatics (2024)
-
SSELM-neg: spherical search-based extreme learning machine for drug–target interaction prediction
BMC Bioinformatics (2023)
-
Advancing drug–target interaction prediction: a comprehensive graph-based approach integrating knowledge graph embedding and ProtBert pretraining
BMC Bioinformatics (2023)
-
Graph regularized non-negative matrix factorization with \(L_{2,1}\) norm regularization terms for drug–target interactions prediction
BMC Bioinformatics (2023)