Abstract
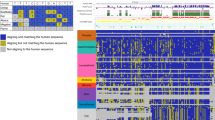
We have analyzed issues of reliability in studies in which comparative genomic approaches have been applied to the discovery of regulatory elements at a genome-wide level in vertebrates. We point out some potential problems with such studies, including difficulties in accurately identifying orthologous promoter regions. Many of these subtle analytical problems have become apparent only when studying the more complex vertebrate genomes. By determining motif reliability, we compared existing tools when applied to the discovery of vertebrate regulatory elements. We then used a statistical clustering method to produce a computational catalog of high quality putative regulatory elements from vertebrates, some of which are widely conserved among vertebrates and many of which are novel regulatory elements. The results provide a glimpse into the wealth of information that comparative genomics can yield and suggest the need for further improvement of genome-wide comparative computational techniques.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Tagle, D. et al. Embryonic ε and γ globin genes of a prosimian primate (Galago crassicaudatus); nucleotide and amino acid sequences, developmental regulation and phylogenetic footprints. J. Mol. Biol. 203, 439–455 (1988).
Cliften, P. et al. Finding functional features in Saccharomyces genomes by phylogenetic footprinting. Science 301, 71–76 (2003).
Kellis, M., Patterson, N., Endrizzi, M., Birren, B. & Lander, E.S. Sequencing and comparison of yeast species to identify genes and regulatory elements. Nature 423, 241–254 (2003).
Karlin, S. & Altschul, S.F. Methods for assessing the statistical significance of molecular sequence features by using general scoring schemes. Proc. Natl. Acad. Sci. USA 87, 2264–2268 (1990).
Dieterich, C., Wang, H., Rateitschak, K., Luz, H. & Vingron, M. CORG: a database for COmparative Regulatory Genomics. Nucleic Acids Res. 31, 55–57 (2003).
Elemento, O. & Tavazoie, S. Fast and systematic genome-wide discovery of conserved regulatory elements using a non-alignment based approach. Genome Biol. 6, R18 (2005).
Xie, X. et al. Systematic discovery of regulatory motifs in human promoters and 3′ UTRs by comparison of several mammals. Nature 434, 338–345 (2005).
Birney, E. et al. An overview of Ensembl. Genome Res. 14, 925–928 (2004).
Chenna, R. et al. Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Res. 31, 3497–3500 (2003).
Bray, N. & Pachter, L. MAVID: Constrained ancestral alignment of multiple sequences. Genome Res. 14, 693–699 (2004).
Brudno, M. et al. LAGAN and Multi-LAGAN: Efficient tools for large-scale multiple alignment of genomic DNA. Genome Res. 13, 721–731 (2003).
Morgenstern, B. DIALIGN 2: Improvement of the segment-to-segment approach to multiple sequence alignment. Bioinformatics 15, 211–218 (1999).
Blanchette, M. et al. Aligning multiple genomic sequences with the threaded blockset aligner. Genome Res. 14, 708–715 (2004).
Blanchette, M. & Tompa, M. Footprinter: A program designed for phylogenetic footprinting. Nucleic Acids Res. 31, 3840–3842 (2003).
Fitch, W.M. Toward defining the course of evolution: Minimum change for a specified tree topology. Syst. Zool. 20, 406–416 (1971).
Pollard, D.A., Bergman, C.M., Stoye, J., Celniker, S.E. & Eisen, M.B. Benchmarking tools for the alignment of functional noncoding DNA. BMC Bioinformatics 5, 6 (2004).
Margulies, E., Blanchette, M., Haussler, D. & Green, E. NISC Comparative Sequencing Program, Haussler, D. & Green, E. Identification and characterization of multi-species conserved sequences. Genome Res. 13, 2507–2518 (2003).
Matys, V. et al. TRANSFAC: transcriptional regulation, from patterns to profiles. Nucleic Acids Res. 31, 374–378 (2003).
International Chicken Genome Sequencing Consortium. Sequence and comparative analysis of the chicken genome provide unique perspectives on vertebrate evolution. Genome Biol. 432, 695–716 (2004).
Zhang, B., Schmoyer, D., Kirov, S. & Snoddy, J. GOTree Machine (GOTM): a web-based platform for interpreting sets of interesting genes using Gene Ontology hierarchies. BMC Bioinformatics 5, 16 (2004).
Thiel, G., Sarraj, J.A. & Stefano, L. cAMP response element binding protein (CREB) activates transcription via two distinct genetic elements of the human glucose-6-phosphatase gene. BMC Mol. Biol. 6, 2 (2005).
Yamazaki, Y., Kubota, H., Nozaki, M. & Nagata, K. Transcriptional regulation of the cytosolic chaperonin θ subunit gene, Cctq, by Ets domain transcription factors Elk-1, Sap-1a, and Net in the absence of serum response factor. J. Biol. Chem. 278, 30642–30651 (2003).
Scholz, H. & Kirschner, K.M. A role for the Wilms' tumor protein WT1 in organ development. Physiology (Bethesda) 20, 54–59 (2005).
Schäffer, A.A. et al. Improving the accuracy of PSI-BLAST protein database searches with composition-based statistics and other refinements. Nucleic Acids Res. 29, 2994–3005 (2001).
Olson, M.V. & Varki, A. Sequencing the chimpanzee genome: insights into human evolution and disease. Nat. Rev. Genet. 4, 20–28 (2003).
Acknowledgements
We thank Mathieu Blanchette, Nan Li, Michal Linial, Larry Ruzzo, Saurabh Sinha, Zasha Weinberg, Zizhen Yao, the Ensembl Help Desk (in particular, Michael Schuster and Ewan Birney) and the anonymous reviewers for their contributions to this work. This material is based upon work supported in part by the National Science Foundation under grant DBI-0218798 and by the National Institutes of Health under grant R01 HG02602.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Prakash, A., Tompa, M. Discovery of regulatory elements in vertebrates through comparative genomics. Nat Biotechnol 23, 1249–1256 (2005). https://doi.org/10.1038/nbt1140
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1140
This article is cited by
-
MGcV: the microbial genomic context viewer for comparative genome analysis
BMC Genomics (2013)
-
GPMiner: an integrated system for mining combinatorial cis-regulatory elements in mammalian gene group
BMC Genomics (2012)
-
Computational Challenges in Deciphering Genomic Structures of Bacteria
Journal of Computer Science and Technology (2010)
-
MATLIGN: a motif clustering, comparison and matching tool
BMC Bioinformatics (2007)
-
How to infer gene networks from expression profiles
Molecular Systems Biology (2007)