Abstract
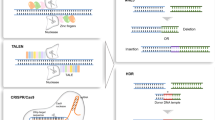
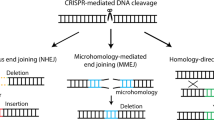
The ability to achieve site-specific manipulation of the mammalian genome has widespread implications for basic and applied research. Gene targeting is a process in which a DNA molecule introduced into a cell replaces the corresponding chromosomal segment by homologous recombination, and thus presents a precise way to manipulate the genome. In the past, the application of gene targeting to mammalian cells has been limited by its low efficiency. Zinc finger nucleases (ZFNs) show promise in improving the efficiency of gene targeting by introducing DNA double-strand breaks in target genes, which then stimulate the cell's endogenous homologous recombination machinery. Recent results have shown that ZFNs can be used to create targeting frequencies of up to 20% in a human disease-causing gene. Future work will be needed to translate these in vitro findings to in vivo applications and to determine whether zinc finger nucleases create undesired genomic instability.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Bob Crimi


Similar content being viewed by others
References
Hinnen, A., Hicks, J.B. & Fink, G.R. Transformation of yeast. Proc. Natl. Acad. Sci. USA 75, 1929–1933 (1978).
Rothstein, R.J. One-step gene disruption in yeast. Methods Enzymol. 101, 202–211 (1983).
Orr-Weaver, T.L., Szostak, J.W. & Rothstein, R.J. Yeast transformation: a model system for the study of recombination. Proc. Natl. Acad. Sci. USA 78, 6354–6358 (1981).
Capecchi, M. Altering the genome by homologous recombination. Science 244, 1288–1292 (1989).
Bradley, A., Ramirez-Solis, R., Zheng, H., Hasty, P. & Davis, A. Genetic manipulation of the mouse via gene targeting in embryonic stem cells. Ciba Found. Symp. 165, 256–269 (1992).
Mansour, S.L., Thomas, K.R. & Capecchi, M.R. Disruption of the proto-oncogene int-2 in mouse embryo-derived stem cells: a general strategy for targeting mutations to non-selectable genes. Nature 336, 348–352 (1988).
Sedivy, J.M. & Sharp, P.A. Positive genetic selection for gene disruption in mammalian cells by homologous recombination. Proc. Natl. Acad. Sci. USA 86, 227–231 (1989).
Hacein-Bey-Abina, S. et al. LMO2-associated clonal T cell proliferation in two patients after gene therapy for SCID-X1. Science 302, 415–419 (2003).
Cavazzana-Calvo, M. et al. Gene therapy of human severe combined immunodeficiency (SCID)-X1 disease. Science 288, 669–672 (2000).
Gaspar, H.B. et al. Gene therapy of X-linked severe combined immunodeficiency by use of a pseudotyped gammaretroviral vector. Lancet 364, 2181–2187 (2004).
Jasin, M. Genetic manipulation of genomes with rare-cutting endonucleases. Trends Genet. 12, 224–228 (1996).
Rouet, P., Smih, F. & Jasin, M. Introduction of double-strand breaks into the genome of mouse cells by expression of rare-cutting endonuclease. Mol. Cell. Biol. 14, 8096–8106 (1994).
Rouet, P., Smih, F. & Jasin, M. Expression of a site-specific endonuclease stimulates homologous recombination in mammalian cells. Proc. Natl. Acad. Sci. USA 91, 6064–6068 (1994).
Smih, F., Rouet, P., Romanienko, P.J. & Jasin, M. Double-strand breaks at the target locus stimulate gene targeting in embryonic stem cells. Nucleic Acids Res. 23, 5012–5019 (1995).
Choulika, A., Perrin, A., Dujon, B. & Nicolas, J.-F. Induction of homologous recombination in mammalian chromosomes by using the I-SceI system of Saccaromyces cerevisiae. Mol. Cell. Biol. 15, 1968–1973 (1995).
Sargent, R.G., Brenneman, M.A. & Wilson, J.H. Repair of site-specific double-strand breaks in a mammalian chromosome by homologous and illegitimate recombination. Mol. Cell. Biol. 17, 267–277 (1997).
Donoho, G., Jasin, M. & Berg, P. Analysis of gene targeting and intrachromosomal homologous recombination stimulated by genomic double-strand breaks in mouse embryonic stem cells. Mol. Cell. Biol. 18, 4070–4078 (1998).
Cohen-Tannoudji, M. et al. I-SceI-induced gene replacement at a natural locus in embryonic stem cells. Mol. Cell. Biol. 18, 1444–1448 (1998).
Porteus, M.H. & Baltimore, D. Chimeric nucleases stimulate gene targeting in human cells. Science 300, 763 (2003).
Kuan, J.Y. & Glazer, P.M. Targeted gene modification using triplex-forming oligonucleotides. Methods Mol. Biol. 262, 173–194 (2004).
Wurtz, N.R. & Dervan, P.B. Sequence specific alkylation of DNA by hairpin pyrrole-imidazole polyamide conjugates. Chem. Biol. 7, 153–161 (2000).
Dervan, P.B. & Edelson, B.S. Recognition of the DNA minor groove by pyrrole-imidazole polyamides. Curr. Opin. Struct. Biol. 13, 284–299 (2003).
Kaihatsu, K., Janowski, B.A. & Corey, D.R. Recognition of chromosomal DNA by PNAs. Chem. Biol. 11, 749–758 (2004).
Kim, Y.G., Cha, J. & Chandrasegaran, S. Hybrid restriction enzymes: zinc finger fusions to Fok I cleavage domain. Proc. Natl. Acad. Sci. USA 93, 1156–1160 (1996).
Chandrasegaran, S. & Smith, J. Chimeric restriction enzymes: what is next? Biol. Chem. 380, 841–848 (1999).
Kandavelou, K., Mani, M., Durai, S. & Chandrasegaran, S. in Nucleic Acids and Molecular Biology, vol. 14 (ed. Pingoud, A.M.) 413–434 (Springer-Verlag, Heidelberg, Germany, 2004).
Kim, Y.G. & Chandrasegaran, S. Chimeric restriction endonuclease. Proc. Natl. Acad. Sci. USA 91, 883–887 (1994).
Kim, Y.G., Smith, J., Durgesha, M. & Chandrasegaran, S. Chimeric restriction enzyme: Gal4 fusion to FokI cleavage domain. Biol. Chem. 379, 489–495 (1998).
Smith, J., Berg, J.M. & Chandrasegaran, S. A detailed study of the substrate specificity of a chimeric restriction enzyme. Nucleic Acids Res. 27, 674–681 (1999).
Smith, J. et al. Requirements for double-strand cleavage by chimeric restriction enzymes with zinc finger DNA-recognition domains. Nucleic Acids Res. 28, 3361–3369 (2000).
Bibikova, M. et al. Stimulation of homologous recombination through targeted cleavage by chimeric nucleases. Mol. Cell. Biol. 21, 289–297 (2001).
Pabo, C.O., Peisach, E. & Grant, R.A. Design and selection of novel Cys2His2 zinc finger proteins. Annu. Rev. Biochem. 70, 313–340 (2001).
Beerli, R.R. & Barbas, C.F. III. Engineering polydactyl zinc-finger transcription factors. Nat. Biotechnol. 20, 135–141 (2002).
Isalan, M. & Choo, Y. Engineering nucleic acid-binding proteins by phage display. Methods Mol. Biol. 148, 417–429 (2001).
Bibikova, M., Golic, M., Golic, K.G. & Carroll, D. Targeted chromosomal cleavage and mutagenesis in Drosophila using zinc-finger nucleases. Genetics 161, 1169–1175 (2002).
Bibikova, M., Beumer, K., Trautman, J.K. & Carroll, D. Enhancing gene targeting with designed zinc finger nucleases. Science 300, 764 (2003).
Lloyd, A., Plaisier, C.L., Carroll, D. & Drews, G.N. Targeted mutagenesis using zinc-finger nucleases in Arabidopsis. Proc. Natl. Acad. Sci. USA 102, 2232–2237 (2005).
Wilson, J.H. Pointing fingers at the limiting step in gene targeting. Nat. Biotechnol. 21, 759–760 (2003).
Porteus, M.H. in Abstract 709 presented at the Annual Meeting of the American Society of Gene Therapy, Minneapolis, Minnesota, June, 2004.
Urnov, F.D. et al. Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 435, 646–651 (2005).
Notarangelo, L.D. et al. Of genes and phenotypes: the immunological and molecular spectrum of combined immune deficiency. Defects of the gamma(c)-JAK3 signaling pathway as a model. Immunol. Rev. 178, 39–48 (2000).
Jamieson, A.C., Miller, J.C. & Pabo, C.O. Drug discovery with engineered zinc-finger proteins. Nat. Rev. Drug Discov. 2, 361–368 (2003).
Bitinaite, J., Wah, D.A., Aggarwal, A.K. & Schildkraut, I. FokI dimerization is required for DNA cleavage. Proc. Natl. Acad. Sci. USA 95, 10570–10575 (1998).
Richardson, C. & Jasin, M. Frequent chromosomal translocations induced by DNA double-strand breaks. Nature 405, 697–700 (2000).
Elliott, B. & Jasin, M. Double-strand breaks and translocations in cancer. Cell. Mol. Life Sci. 59, 373–385 (2002).
Tan, S. et al. Zinc-finger protein-targeted gene regulation: genomewide single-gene specificity. Proc. Natl. Acad. Sci. USA 100, 11997–12002 (2003).
Jantz, D., Amann, B.T., Gatto, G.J. Jr. & Berg, J.M. The design of functional DNA-binding proteins based on zinc finger domains. Chem. Rev. 104, 789–799 (2004).
van Gent, D.C., Hoeijmakers, J.H. & Kanaar, R. Chromosomal stability and the DNA double-stranded break connection. Nat. Rev. Genet. 2, 196–206 (2001).
Jasin, M. Chromosome breaks and genomic instability. Cancer Invest. 18, 78–86 (2000).
Paques, F. & Haber, J.E. Multiple pathways of recombination induced by double-strand breaks in Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 63, 349–404 (1999).
West, S.C. et al. Double-strand break repair in human cells. Cold Spring Harb. Symp. Quant. Biol. 65, 315–321 (2000).
McCormack, W.T., Tjoelker, L.W. & Thompson, C.B. Avian B-cell development: generation of an immunoglobulin repertoire by gene conversion. Annu. Rev. Immunol. 9, 219–241 (1991).
Diakun, G.P., Fairall, L. & Klug, A. EXAFS study of the zinc-binding sites in the protein transcription factor IIIA. Nature 324, 698–699 (1986).
Pavletich, N.P. & Pabo, C.O. Zinc finger-DNA recognition: crystal structure of a Zif268-DNA complex at 2.1 A. Science 252, 809–817 (1991).
Wolfe, S.A., Nekludova, L. & Pabo, C.O. DNA recognition by Cys2His2 zinc finger proteins. Annu. Rev. Biophys. Biomol. Struct. 29, 183–212 (2000).
Segal, D.J. & Barbas, C.F. 3rd Custom DNA-binding proteins come of age: polydactyl zinc-finger proteins. Curr. Opin. Biotechnol. 12, 632–637 (2001).
Segal, D.J. et al. Evaluation of a modular strategy for the construction of novel polydactyl zinc finger DNA-binding proteins. Biochemistry 42, 2137–2148 (2003).
Liu, Q., Xia, Z., Zhong, X. & Case, C.C. Validated zinc finger protein designs for all 16 GNN DNA triplet targets. J. Biol. Chem. 277, 3850–3856 (2002).
Segal, D.J., Dreier, B., Beerli, R.R. & Barbas, C.F. III. Toward controlling gene expression at will: selection and design of zinc finger domains recognizing each of the 5′-GNN-3′ DNA target sequences. Proc. Natl. Acad. Sci. USA 96, 2758–2763 (1999).
Sera, T. & Uranga, C. Rational design of artificial zinc-finger proteins using a nondegenerate recognition code table. Biochemistry 41, 7074–7081 (2002).
Kim, J.S. & Pabo, C.O. Getting a handhold on DNA: design of poly-zinc finger proteins with femtomolar dissociation constants. Proc. Natl. Acad. Sci. USA 95, 2812–2817 (1998).
Liu, Q., Segal, D.J., Ghiara, J.B. & Barbas, C.F. III. Design of polydactyl zinc-finger proteins for unique addressing within complex genomes. Proc. Natl. Acad. Sci. USA 94, 5525–5530 (1997).
Moore, M., Klug, A. & Choo, Y. Improved DNA binding specificity from polyzinc finger peptides by using strings of two-finger units. Proc. Natl. Acad. Sci. USA 98, 1437–1441 (2001).
Chevalier, B.S. & Stoddard, B.L. Homing endonucleases: structural and functional insight into the catalysts of intron/intein mobility. Nucleic Acids Res. 29, 3757–3774 (2001).
Gimble, F.S. Invasion of a multitude of genetic niches by mobile endonuclease genes. FEMS Microbiol. Lett. 185, 99–107 (2000).
Belfort, M. & Roberts, R.J. Homing endonucleases: keeping the house in order. Nucleic Acids Res. 25, 3379–3388 (1997).
Chevalier, B.S. et al. Design, activity, and structure of a highly specific artificial endonuclease. Mol. Cell 10, 895–905 (2002).
Epinat, J.C. et al. A novel engineered meganuclease induces homologous recombination in yeast and mammalian cells. Nucleic Acids Res. 31, 2952–2962 (2003).
Sussman, D. et al. Isolation and characterization of new homing endonuclease specificities at individual target site positions. J. Mol. Biol. 342, 31–41 (2004).
Gimble, F.S., Moure, C.M. & Posey, K.L. Assessing the plasticity of DNA target site recognition of the PI-SceI homing endonuclease using a bacterial two-hybrid selection system. J. Mol. Biol. 334, 993–1008 (2003).
Acknowledgements
We thank Scott Cameron, Jim Amatruda, Brian Cauff, Shondra Pruett, Patrick Connelly, Michael Holmes and Philip Gregory for reading the manuscript and for their helpful comments. The work in the Porteus lab is supported by the Burroughs-Wellcome Fund, a K08 award from the National Heart and Blood Institute, and UT Southwestern Medical Center. Work in the Carroll lab is supported by research grants from the US Public Health Service.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
D.C. is an inventor on US and international patent applications that describe the ZFN-based gene-targeting technology.
Rights and permissions
About this article
Cite this article
Porteus, M., Carroll, D. Gene targeting using zinc finger nucleases. Nat Biotechnol 23, 967–973 (2005). https://doi.org/10.1038/nbt1125
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1125