Abstract
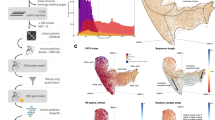
Accurate determination of functional interactions among proteins at the genome level remains a challenge for genomic research. Here we introduce a genome-scale approach to functional protein annotation—phylogenomic mapping—that requires only sequence data, can be applied equally well to both finished and unfinished genomes, and can be extended beyond single genomes to annotate multiple genomes simultaneously. We have developed and applied it to more than 200 sequenced bacterial genomes. Proteins with similar evolutionary histories were grouped together, placed on a three dimensional map and visualized as a topographical landscape. The resulting phylogenomic maps display thousands of proteins clustered in mountains on the basis of coinheritance, a strong indicator of shared function. In addition to systematic computational validation, we have experimentally confirmed the ability of phylogenomic maps to predict both mutant phenotype and gene function in the delta proteobacterium Myxococcus xanthus.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hartwell, L.H., Hopfield, J.J., Leibler, S. & Murray, A.W. From molecular to modular cell biology. Nature 402, C47–C52 (1999).
Marcotte, E.M. et al. Detecting protein function and protein-protein interactions from genome sequences. Science 285, 751–753 (1999).
Gertz, J. et al. Inferring protein interactions from phylogenetic distance matrices. Bioinformatics 19, 2039–2045 (2003).
Pazos, F. & Valencia, A. In silico two-hybrid system for the selection of physically interacting protein pairs. Proteins 47, 219–227 (2002).
Overbeek, R., Fonstein, M., D'Souza, M., Pusch, G.D. & Maltsev, N. The use of gene clusters to infer functional coupling. Proc. Natl. Acad. Sci. USA 96, 2896–2901 (1999).
Huynen, M.A., Snel, B., von Mering, C. & Bork, P. Function prediction and protein networks. Curr. Opin. Cell Biol. 15, 191–198 (2003).
Pellegrini, M., Marcotte, E.M., Thompson, M.J., Eisenberg, D. & Yeates, T.O. Assigning protein functions by comparative genome analysis: protein phylogenetic profiles. Proc. Natl. Acad. Sci. USA 96, 4285–4288 (1999).
Eisen, M.B., Spellman, P.T., Brown, P.O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl. Acad. Sci. USA 95, 14863–14868 (1998).
Barabasi, A.L. & Oltvai, Z.N. Network biology: understanding the cell's functional organization. Nat. Rev. Genet. 5, 101–113 (2004).
Alter, O., Brown, P.O. & Botstein, D. Singular value decomposition for genome-wide expression data processing and modeling. Proc. Natl. Acad. Sci. USA 97, 10101–10106 (2000).
Davidson, G.S., Wylie, B.N. & Boyack, K. Cluster stability and the use of noise in interpretation of clustering. in Proceedings of the IEEE Symposium on Information Visualization 2001 (INFOVIS'01), 23–30 (IEEE Computer Society, 2001).
Werner-Washburne, M. et al. Comparative analysis of multiple genome-scale data sets. Genome Res. 12, 1564–1573 (2002).
Kim, S.K. et al. A gene expression map for Caenorhabditis elegans. Science 293, 2087–2092 (2001).
Marcotte, E.M., Xenarios, I., van Der Bliek, A.M. & Eisenberg, D. Localizing proteins in the cell from their phylogenetic profiles. Proc. Natl. Acad. Sci. USA 97, 12115–12120 (2000).
Enault, F., Suhre, K., Poirot, O., Abergel, C. & Claverie, J.M. Phydbac2: improved inference of gene function using interactive phylogenomic profiling and chromosomal location analysis. Nucleic Acids Res. 32, W336–W339 (2004).
Altschul, S.F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
de Hoon, M.J., Imoto, S., Nolan, J. & Miyano, S. Open source clustering software. Bioinformatics 20, 1453–1454 (2004).
Julien, B. & Shah, S. Heterologous expression of epothilone biosynthetic genes in Myxococcus xanthus. Antimicrob. Agents Chemother. 46, 2772–2778 (2002).
Gerth, K. et al. The myxalamids, new antibiotics from Myxococcus xanthus (Myxobacterales). I. Production, physico-chemical and biological properties, and mechanism of action. J. Antibiot. (Tokyo) 36, 1150–1156 (1983).
Pospiech, A., Cluzel, B., Bietenhader, J. & Schupp, T. A new Myxococcus xanthus gene cluster for the biosynthesis of the antibiotic saframycin Mx1 encoding a peptide synthetase. Microbiology 141, 1793–1803 (1995).
Shi, W. & Zusman, D.R. The two motility systems of Myxococcus xanthus show different selective advantages on various surfaces. Proc. Natl. Acad. Sci. USA 90, 3378–3382 (1993).
Kaiser, D. & Welch, R. Dynamics of fruiting body morphogenesis. J. Bacteriol. 186, 919–927 (2004).
Kaiser, D. Coupling cell movement to multicellular development in myxobacteria. Nat. Rev. Microbiol. 1, 45–54 (2003).
Wu, S.S. & Kaiser, D. Genetic and functional evidence that Type IV pili are required for social gliding motility in Myxococcus xanthus. Mol. Microbiol. 18, 547–558 (1995).
Lowe, J., van den Ent, F. & Amos, L.A. Molecules of the bacterial cytoskeleton. Annu. Rev. Biophys. Biomol. Struct. 33, 177–198 (2004).
Wolgemuth, C., Hoiczyk, E., Kaiser, D. & Oster, G. How myxobacteria glide. Curr. Biol. 12, 369–377 (2002).
Raetz, C.R. & Whitfield, C. Lipopolysaccharide endotoxins. Annu. Rev. Biochem. 71, 635–700 (2002).
Gaspar, J.A., Thomas, J.A., Marolda, C.L. & Valvano, M.A. Surface expression of O-specific lipopolysaccharide in Escherichia coli requires the function of the TolA protein. Mol. Microbiol. 38, 262–275 (2000).
Fink, J.M. & Zissler, J.F. Defects in motility and development of Myxococcus xanthus lipopolysaccharide mutants. J. Bacteriol. 171, 2042–2048 (1989).
Youderian, P., Burke, N., White, D.J. & Hartzell, P.L. Identification of genes required for adventurous gliding motility in Myxococcus xanthus with the transposable element mariner. Mol. Microbiol. 49, 555–570 (2003).
Caberoy, N.B., Welch, R.D., Jakobsen, J.S., Slater, S.C. & Garza, A.G. Global mutational analysis of NtrC-like activators in Myxococcus xanthus: identifying activator mutants defective for motility and fruiting body development. J. Bacteriol. 185, 6083–6094 (2003).
Kroos, L., Kuspa, A. & Kaiser, D. Defects in fruiting body development caused by Tn5 lac insertions in Myxococcus xanthus. J. Bacteriol. 172, 484–487 (1990).
Harris, M.A. et al. The Gene Ontology (GO) database and informatics resource. Nucleic Acids Res. 32 Database issue, D258–261 (2004).
Michalickova, K. et al. SeqHound: biological sequence and structure database as a platform for bioinformatics research. BMC Bioinformatics 3, 32 (2002).
Camon, E. et al. The Gene Ontology Annotation (GOA) Database: sharing knowledge in Uniprot with Gene Ontology. Nucleic Acids Res. 32, D262–D266 (2004).
Boyle, E.I. et al. GO:TermFinder – open source software for accessing Gene Ontology information and finding significantly enriched Gene Ontology terms associated with a list of genes. Bioinformatics 20, 3710–3715 (2004).
Venter, J.C. et al. Environmental genome shotgun sequencing of the Sargasso Sea. Science 304, 66–74 (2004).
McAdams, H.H., Srinivasan, B. & Arkin, A.P. The evolution of genetic regulatory systems in bacteria. Nat. Rev. Genet. 5, 169–178 (2004).
Holder, M. & Lewis, P.O. Phylogeny estimation: traditional and Bayesian approaches. Nat. Rev. Genet. 4, 275–284 (2003).
Daubin, V., Moran, N.A. & Ochman, H. Phylogenetics and the cohesion of bacterial genomes. Science 301, 829–832 (2003).
Florea, L., McClelland, M., Riemer, C., Schwartz, S. & Miller, W. EnteriX 2003: Visualization tools for genome alignments of Enterobacteriaceae. Nucleic Acids Res. 31, 3527–3532 (2003).
Galperin, M.Y. & Koonin, E.V. Who's your neighbor? New computational approaches for functional genomics. Nat. Biotechnol. 18, 609–613 (2000).
Stuart, J.M., Segal, E., Koller, D. & Kim, S.K. A gene-coexpression network for global discovery of conserved genetic modules. Science 302, 249–255 (2003).
Acknowledgements
We thank Harley McAdams, Lucy Shapiro, William Nierman and Dale Kaiser for helpful discussions. We thank the Monsanto Corporation and the Institute for Genomics Research for providing access to the genome sequence of M. xanthus DK1622. This work was supported in part by National Science Foundation (NSF) Grant MCB-0444154 to A.G.G. B.S.S. was supported by a Department of Defense National Defense Science and Engineering Graduate Fellowship through the Army Research Office. Sequencing of M. xanthus DK1622 was accomplished with support from the NSF.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
Global computational validation of phylogenomic mapping (PDF 153 kb)
Supplementary Note 1
Similarity matrix generation (PDF 91 kb)
Supplementary Note 2
Motility assays and plasmid insertion (PDF 54 kb)
Supplementary Note 3
Gene ontology analysis (PDF 61 kb)
Rights and permissions
About this article
Cite this article
Srinivasan, B., Caberoy, N., Suen, G. et al. Functional genome annotation through phylogenomic mapping. Nat Biotechnol 23, 691–698 (2005). https://doi.org/10.1038/nbt1098
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1098
This article is cited by
-
Disparate data fusion for protein phosphorylation prediction
Annals of Operations Research (2010)
-
Complete genome sequence of the myxobacterium Sorangium cellulosum
Nature Biotechnology (2007)