Abstract
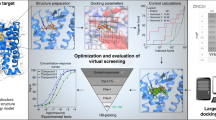
The need to decrease the time scale for clinical compound discovery has led to innovations at several stages in the process, including genomics/proteomics for target identification, ultrahigh-throughput screening1 for lead identification, and structure-based drug design2 and combinatorial chemistry3 for lead optimization. A critical juncture in the process is the identification of a proper lead compound, because a poor choice may generate costly difficulties at later stages. Lead compounds are commonly identified from high-throughput screens of large compound libraries, derived from known substrates/inhibitors, or identified in computational prescreeusing X-ray crystal structures2,4,5,6. Structural information is often consulted to efficiently optimize leads, but under the current paradigm, such data require preidentification and confirmation of compound binding. Here, we describe a new X-ray crystallography–driven screening technique that combines the steps of lead identification, structural assessment, and optimization. The method is rapid, efficient, and high-throughput, and it results in detailed crystallographic structure information. The utility of the method is demonstrated in the discovery and optimization of a new orally available class of urokinase inhibitors for the treatment of cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Sundberg, S. High-throughput and ultra-high-throughput screening: solution- and cell-based approaches. Curr. Opin. Biotechnol. 11, 47–53 (2000).
Verlinde, C.L. & Hol, W.G. Structure-based drug design: progress, results and challenges. Structure 2, 577–87 (1994).
Salemme, F., Spurlino, J. & Bone, R. Serendipity meets precision: the integration of structure-based drug design and combinatorial chemistry for efficient drug discovery. Structure 15, 319–324 (1997).
Kuntz, I. Structure-based strategies for drug design and discovery. Science 257, 1078–1082 (1992).
Shuker, S.B., Hajduk, P.J., Meadows, R.P. & Fesik, S.W. Discovering high affinity ligands for proteins: SAR by NMR. Science 274, 1531–1534 (1996).
Fejzo, J. et al. The SHAPES strategy: an NMR-based approach for lead generation in drug discovery. Chem. Biol. 6, 755–769 (1999).
Allen, K.N. et al. An experimental approach to mapping the binding surfaces of crystalline proteins. J. Phys. Chem. 100, 2605–2611 (1996).
Goodford, P.J. A computational procedure for determining energetically favorable binding sites on biologically important molecules. J. Med. Chem. 28, 849–857 (1985).
Caflish, A., Miranker, A. & Karplus, M. Multiple copy simultaneous search and construction of ligands in binding sites: application to inhibitors of HIV-1 aspartic proteinase. J. Med. Chem. 36, 2142–2167 (1993).
Kuntz, I.D., Meng, E.C. & Shoichet, B.K. Structure-based drug design. Accounts Chem. Res. 27, 117–123 (1994).
Verlinde, C., Rudenko, G. & Hol, W. In search of new lead compounds for trypanosomiasis drug desgin: a protein structure-based linked-fragment approach. J. Comput. Aided Mol. Des. 6, 131–147 (1992).
Cheng, X. et al. Using electrospray ionization FTICR mass spectrometry to study competitive binding of inhibitors to carbonic anhydrase. J. Am. Chem. Soc. 117, 8859–8860 (1995).
Lin, M., Shapiro, M.J. & Wareing, J.R. Screening mixtures by affinity NMR. J. Org. Chem. 62, 8930–8931 (1997).
Muchmore, S. et al. Automated crystal mounting and data collection for protein crystallography. Structure, in press.
Verlinde, C., Kim, H., Berstein, B., Mande, S. & Hol, W. Antitrypanosomiasis drug development based on structures of glycolytic enzymes. In Structure-based drug design (ed. Veerapandian, P.) 365–394 (Marcel Dekker, Inc, New York, NY; 1997).
Reuning, U. et al. Multifunctional potential of the plasminogen activation system in tumor invasion and metastasis. Int. J. Oncol. 13, 896–906 (1998).
Carmeliet, P. et al. Physiological consequences of loss of plasminogen activator gene function in mice. Nature 368, 419–24 (1994).
Nienaber, V., Wang, J., Davidson, D. & Henkin, J. Re-engineering of human urokinase provides a system for structure-based drug design at high resolution and reveals a novel bindng sub-site. J. Biol. Chem. 275, 7239–7248 (2000).
Nienaber, V. et al. Structure-directed discovery of potent non-peptidic inhibitors of human urokinase which access a novel binding sub-site. Structure 8, 553–563 (2000).
Menear, K. Progress towards the discovery of orally active thrombin inhibitors. Curr. Med. Chem. 5, 457–468 (1998).
Daylight chemical information systems, I. (Mission Viejo, CA,).
Price, S. & Nagai, K. Protein engineering as a tool for crystallography. Cur. Opin. Biotechnol. 6, 425–430 (1995).
Berman, H.M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Otwinowski, Z. & Minor, W. Processing of X-ray dIffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Brunger, A.T. X-PLOR version 3.1 manual. (Yale University Press, New Haven, CT; 1993).
Acknowledgements
We thank R. Meadows, S. Fesik, and S. Betz for helpful discussions and comments on the manuscript; J. Wang for preparation of the urokinase protein; and S. Muchmore and T. Rockway for discussions on the technique.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nienaber, V., Richardson, P., Klighofer, V. et al. Discovering novel ligands for macromolecules using X-ray crystallographic screening. Nat Biotechnol 18, 1105–1108 (2000). https://doi.org/10.1038/80319
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/80319
This article is cited by
-
Rapid optimisation of fragments and hits to lead compounds from screening of crude reaction mixtures
Communications Chemistry (2020)
-
NMR-based fragment screening and lead discovery accelerated by principal component analysis
Journal of Biomolecular NMR (2019)
-
The crystal structure of the AgamOBP1•Icaridin complex reveals alternative binding modes and stereo-selective repellent recognition
Cellular and Molecular Life Sciences (2017)
-
Biophysics in drug discovery: impact, challenges and opportunities
Nature Reviews Drug Discovery (2016)
-
Catalytic activation of pre-substrates via dynamic fragment assembly on protein templates
Nature Communications (2014)