Abstract
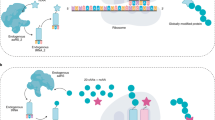
Altering protein structure via the techniques of protein engineering has already allowed the development of proteins displaying both modified and novel activities. The only limitation of conventional site-directed mutagenesis, the cornerstone of protein engineering, is that substitutions are restricted to the 20 naturally occurring, proteinogenic amino acids. However, the discovery of a 21st amino acid, selenocysteine, and the development of novel in vitro translation systems have demonstrated that considerably more substitutions are possible. To this end, a number of experimental approaches have been developed that allow the incorporation of synthetic amino acids into proteins. Some of these have already been successfully applied in vitro and efforts to transfer this technology to in vivo systems are now underway.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Smith, M.S. 1985. In vitro mutagenesis. Annu. Rev. Genet. 19: 423–462.
Medynski, D. 1992. Genetic approaches to protein structure and function: point mutations as modifiers of protein function. Bio/Technology 10: 1002–1006.
Creighton, T.E. 1993. Proteins: structures and molecular properties, 2nd Ed. W.H. Freeman and Company, New York.
Matthews, B.W. 1993. Structural and genetic analysis of protein stability. Annu. Rev. Biochem. 62: 139–160.
Böck, A., Forchhammer, K., Heider, J., Leinfelder, W., Sawers, G., Veprek, B. and Zinoni, F. 1991. Selenocysteine: the 21st amino acid. Mol. Microbiol. 5: 515–520.
Noren, C.J., Anthony-Cahill, S.J., Griffith, M.C. and Schulte, P.G. 1989. A general method for site-specific incoiporation of unnatural amino acids into proteins. Science 244: 182–188.
Mazel, D., Pochet, S. and Marlière, P. 1994. Genetic characterization of polypeptide defomiylase, a distinctive enzyme of eubacterial translation. EMBO J. 13: 914–923.
Rajbhandary, U.L. 1994. Initiator transfer RNAs. J. Bacteriol. 176: 547–552.
Zinoni, P., Birkmann, A., Leinfelder, W. and Böck, A. 1987. Cotranslational insertion of selenocysteine into formate dehydrogenase from Escherichia coli directed by a UGA codon. Proc. Natl. Acad. Sci. USA 84: 3156–3160.
Böck, A., Forchhammer, K., Heider, J. and Baron, C. 1991. Selenoprotein synthesis: an expansion of the genetic code. Trends Biochem. Sci. 16: 463–467.
Ringquist, S., Schneider, D., Gibson, T., Baron, C., Böck, A. and Gold, L. 1994. Recognition of the mRNA selenocysteine insertion sequence by the specialized translation elongation factor SELB. Genes Dev. 8: 376–385.
Hendrickson, W.A., Horton, J.R. and LeMaster, D.M. 1990. Selenomethionyl proteins produced for analysis by multiwavelength anomalous diffraction (MAD): a vehicle for direct determination of three-dimensional structure. EMBO J. 9: 1665–1672.
Hendrickson, W.A. 1991. Determination of macromolecular structures from anomalous diffraction of synchrotron radiation. Science 254: 51–58.
Müller, S., Senn, H., Gsell, B., Vetter, W., Baron, C. and Böck, A. 1994. The formation of diselenide bridges in proteins by incorporation of selenocysteine residues–Biosynthesis and characterization of (Se) (2)-thioredoxin. Biochemistry 33: 3404–3412.
Hilvert, D. 1991. Extending the chemistry of enzymes and abzymes. Trends Biotechnol. 9: 11–17.
Bell, I.M. and Hilvert, D. 1993. Peroxide dependence of the semisynthetic enzyme selenosubtilisin. Biochemistry 32: 13969–13973.
Mottagui-Tabar, S., Bjornsson, A. and Isaksson, L.A. 1994. The second to last amino acid in the nascent peptide as a codon context determinant. EMBO J. 13: 249–257.
Hatfield, D. and Diamond, A. 1993. UGA: a split personality in the genetic code. Trends Genet. 9: 69–70.
Osawa, S., Jukes, T.H., Watanabe, K. and Muto, A. 1992. Recent evidence for evolution of the genetic code. Microbiol. Rev. 56: 229–264.
Richmond, M.H. 1962. The effect of amino acid analogues on growth and protein synthesis in microorganisms. Bacteriol. Rev. 26: 398–420.
Tam, R. and Saier, M.H. Jr., 1993. Structural, functional, and evolutionary relationships among extracellular solute-binding receptors of bacteria. Microbiol. Rev. 57: 320–346.
Reizer, J., Finley, K., Kakuda, D., Macleod, C.L., Reizer, A. and Saier, M.H. Jr., 1993. Mammalian integral membrane receptors are homologous to facilitators and antiporters of yeast, funghi, and eubacteria. Protein Sci. 2: 20–30.
Hennecke, H. and Böck, A. 1975. Altered α subunits in phenylalanyl-tRNA synthetases from p-fluorophenylalanine-resistant strains of Escherichia coli. Eur. J. Biochem. 55: 431–437.
Jakubowski, H. and Goldman, E. 1992. Editing of errors in selection of amino acids for protein synthesis. Microbiol. Rev. 56: 412–429.
Kim, H.Y., Ghosh, G., Schulman, L.H., Brunie, S. and Jakubowski, H. 1993. The relationship between synthetic and editing functions of the active site of an aminoacyl-tRNA synthetase. Proc. Natl. Acad. Sci. USA 90: 11553–11557.
Thompson, R.C. 1988. EFTu provides an internal kinetic standard for translational accuracy. Trends Biochem. Sci. 13: 91–93.
Stanzel, M., Schön, A. and Sprinzl, M. 1994. Discrimination against misacylated tRNA by chloroplast elongation factor Tu. Eur. J. Biochem. 219: 435–439.
Ibba, M., Kast, P. and Hennecke, H. 1994. Substrate specificity is determined by amino acid binding pocket size in Escherichia coli phenylalanyl-tRNA synthetase. Biochemistry. In press.
Hortin, G. and Boime, I. 1983. Applications of amino acid analogs for studying co-and posttranslational modifications of proteins. Methods Enzymol. 96: 777–784.
Wilson, M.J. and Hatfield, D.L. 1984. Incorporation of modified amino acids into proteins in vivo. Biochim. Biophys. Acta 781: 205–215.
Heinemeyer, W., Kleinschmidt, J.A., Saidowsky, J., Escher, C. and Wolf, D.H. 1991. Proteinase yscE, the yeast proteosome/multicatalytic-multifunctional proteinase: mutants unravel its function in stress-induced proteolysis and uncover its necessity for survival. EMBO J. 10: 555–562.
Kohno, T., Kohda, D., Haruki, M., Yokoyama, S. and Miyazawa, T. 1990. Nonprotein amino acid furanomycin, unlike isoleucine in chemical structure, is charged to isoleucine tRNA by isoleucyl-tRNA synthetase and incorporated into protein. J. Biol. Chem. 265: 6931–6935.
Lemeignan, B., Sonigo, P. and Marlière, P. 1993. Phenotypic suppression by incorporation of an alien amino acid. J. Mol. Biol. 231: 161–166.
Koide, H., Yokoyama, S., Kawai, G., Ha, J-M., Oka, T., Kawai, S., Miyake, T., Fuwa, T. and Miyazawa, T. 1988. Biosynthesis of a protein containing a nonprotein amino acid by Escherichia coli: L-2 amino hexanoic acid at position 21 in human epidermal growth factor. Proc. Natl. Acad. Sci. USA 85: 6237–6241.
Hinds, M.G., King, R.W. and Feeney, J. 1992. 19F n.m.r. studies of conformational changes accompanying cyclic AMP binding to 3-fluorophenylalanine-containing cyclic AMP receptor protein from Escherichia coli. Biochem. J. 287: 627–632.
Kanemori, M., Mori, H. and Yura, T. 1994. Induction of Escherichia coli heat shock proteins by abnormal proteins results solely from stabilization of σ32, p. 142. In: Biology of Heat Shock Proteins and Molecular Chaperones. Proceedings of the 1994 meeting on Cold Spring Harbor Laboratory, Cold Spring Harbor, New York.
Ross, J.B.A., Senear, D.F., Waxman, E., Kombo, B.B., Rusinova, E., Huang, Y.E., Laws, W.R., Ross, J.B.A., and Hasselbacher, C.A. 1992. Spectral enhancement of proteins: biological incorporation and fluorescene characterization of 5-hydroxytryptophan in bacteriophage γ cI represser. Proc. Natl. Acad. Sci. USA 89: 12023–12027.
Holman, C. and Benisek, W.F. 1994. Extent of proton transfer in the transition states of the reaction catalyzed by the Δ5-3-ketosteroid isomerase of Comamonas (Pseudomonas) testosteroni: site-specific replacement of the active site base, aspartate 38, by the weaker base alanine-3-sulfinate. Biochemistry 33: 2672–2681.
Kim, D-W., Yoshimura, T., Esaki, N., Satoh, E. and Soda, K. 1994. Studies of the active-site lysyl residue of thermostable aspartate aminotransferase: combination of site-directed mutagenesis and chemical modification. J. Biochem. (Tokyo) 115: 93–97.
Normanly, J., Kleina, L.G., Masson, J-M., Abelson, J. and Miller, J.H. 1990. Construction of Escherichia coli amber suppressor tRNA genes. J. Mol. Biol. 213: 719–726.
Michaels, M.L., Kim, C.W., Matthews, D.A. and Miller, J.H. 1990. Escherichia coli thymidyiate synthase: amino acid substitutions by suppression of amber nonsense mutations. Proc. Natl. Acad. Sci. USA 87: 3957–3961.
Ellman, J., Mendel, D., Anthony-Cahill, S., Noren, C.J. and Schultz, P.G. 1991. Biosynthetic methods for introducing unnatural amino acids site-specifically into proteins. Methods Enzymol. 202: 301–336.
Brunner, J. 1993. Biosynthetic incorporation of non-natural amino acids into proteins. Chem. Soc. Rev. 22: 183–189.
Mendel, D., Ellman, J. and Schultz, P.G. 1993. Protein biosynthesis with conformationally restricted amino acids. J. Am. Chem. Soc. 115: 4359–4360.
Spirin, A.S., Baranov, V.I., Ryabova, L.A., Ovodov, S.Y. and Alakhov, Y.B. 1988. A continuous cell-free translation system capable of producing poly peptides in high yield. Science 242: 1162–1164.
Ying, W., Zhang, D.Y. and Kramer, F.R. 1992. Amplifiable messenger RNA. Proc. Natl. Acad. Sci. USA 89: 11769–11773.
Sonar, S., Patel, N., Fischer, W. and Rothschild, K.J. 1993. Cell-free synthesis, functional refolding, and spectroscopic characterization of bacteriorhodopsin, an integral membrane protein. Biochemistry 32: 13777–0781.
Brunner, J. 1993. New photolabelling and crosslinking methods. Annu. Rev. Biochem. 62: 483–514.
High, S., Martoglio, B., Görlich, D., Andersen, S.S.L., Ashford, A.J., Giner, A., Hartmann, E., Prehn, S., Rapoport, T.A., Dobberstein, B. and Brunner, J. 1993. Site-specific photocross-linking reveals that sec61p and TRAM contact different regions of a membrane-inserted signal sequence. J. Biol. Chem. 268: 26745–26751.
Ellman, J.A., Mendel, D. and Schultz, P.G. 1992. Site-specific incorporation of novel backbone structures into proteins. Science 255: 197–200.
Mendel, D., Ellman, J.A., Chang, Z., Veenstra, D.L., Kollman, P.A. and Schultz, P.G. 1992. Probing protein stability with unnatural amino acids. Science 256: 1798–1802.
Chung, H.-H., Benson, D.R. and Schultz, P.G. 1993. Probing the structure and mechanism of ras protein with an expanded genetic code. Science 259: 806–809.
Judice, J.K., Gamble, T.R., Murphy, E.C., de Vos, A. M. and Schultz, P.G. 1993. Probing the mechanism of staphylococcal nuclease with unnatural amino acids: kinetic and structural studies. Science 261: 1578–1581.
Bain, J.D., Diala, E.S., Glabe, C.G.D.A., Lyttle, M.H., Dix, T.A. and Chamberlin, A.R. 1991. Site-specific incorporation of nonnatural residues during in vitro protein biosynthesis with semisynthetic aminoacyl-tRNAs. Biochemistry 30: 5411–5421.
Piccirilli, J.A., Krauch, T., Moroney, S.E. and Benner, S.A. 1990. Enzymatic incorporation of a new base pair into DNA and RNA extends the genetic alphabet. Nature 343: 33–37.
Bain, J.D., Switzer, C., Chamberlain, A.R. and Benner, S.A. 1992. Ribo-some-mediated incorporation of a non-standard amino acid into a peptide through expansion of the genetic code. Nature 356: 537–539.
Schon, A., Kannangara, C.G., Gough, S. and Soil, D. 1988. Protein biosynthesis in organelles requires misaminoacylation of tRNA. Nature 331: 187–190.
Carter, C.W., Jr. 1993. Cognition, mechanism, and evolutionary relationships in aminoacyl-tRNA synthetases. Annu. Rev. Biochem. 62: 715–748.
Kast, P. and Hennecke, H. 1991. Amino acid substrate specificity of Escherichia coli phenylalanyl-tRNA synthetase altered by distinct mutations. J. Mol. Biol. 222: 99–124.
Kast, P. 1991. Investigations on Escherichia coli phenylalanyl-tRNA synthetase at the molecular level. Ph.D. Thesis ETH Nr. 9468.
McClain, W.H. 1993. Rules that govern tRNA identity in protein synthesis. J. Mol. Biol. 234: 257–280.
Saks, M.E., Sampson, J.R. and Abelson, J.N. 1994. The transfer RNA identity problem: a search for rules. Science 263: 191–197.
Rould, M.A., Perona, J.J. and Steitz, T.A. 1991. Structural basis of anticodon loop recognition by glutaminyl-tRNA synthetase. Nature 352: 213–218.
Weygand-Durasevic, L., Schwob, E. and Söil, D. 1993. Acceptor end binding domain interactions ensure correct aminoacylation of transfer RNA. Proc. Natl. Acad. Sci. USA 90: 2010–2014.
Jacobsen, J.R., Prudent, J.R., Kochersperger, L., Yonkovich, S. and Schultz, P.G. 1992. An efficient antibody-catalyzed aminoacylation reaction. Science 256: 365–367.
Kast, P. 1994. pKSS-a second-generation general purpose cloning vector for efficient positive selection of recombinant clones. Gene 138: 109–114.
Kurzchalia, T.V., Wiedmann, M., Breter, H., Zimmermann, W., Bauschke, E. and Rapoport, T.A. 1988. tRNA-mediated labelling of proteins with biotin. Eur. J. Biochem. 172: 663–668.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ibba, M., Hennecke, H. Towards Engineering Proteins by Site-Directed Incorporation In Vivo of Non-Natural Amino Acids. Nat Biotechnol 12, 678–682 (1994). https://doi.org/10.1038/nbt0794-678
Issue Date:
DOI: https://doi.org/10.1038/nbt0794-678