Abstract
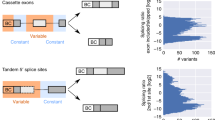
The human transcriptome is marked by extensive alternative mRNA splicing and the expression of many closely related genes, which may be difficult to distinguish using standard microarray techniques. Here we describe a sensitive and specific assay for parallel analysis of mRNA isoforms on a fiber-optic microarray platform. The method permits analysis of mRNA transcripts without prior RNA purification or cDNA synthesis. Using an endogenously expressed viral transcript as a model, we demonstrated that the assay readily detects mRNA isoforms from as little as 10–100 pg of total cellular RNA or directly from a few cells. Multiplexed analysis of human cancer cell lines revealed differences in mRNA splicing and suggested a potential autocrine mechanism in the development of choriocarcinomas. Our approach may be useful in the large-scale analysis of the role of alternative splicing in development and disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Lander, E.S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Venter, J.C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Graveley, B.R. Alternative splicing: increasing diversity in the proteomic world. Trends Genet. 17, 100–107 (2001).
Schmucker, D. et al. Drosophila Dscam is an axon guidance receptor exhibiting extraordinary molecular diversity. Cell 101, 671–684 (2000).
Lopez, A.J. Alternative splicing of pre-mRNA: developmental consequences and mechanisms of regulation. Annu. Rev. Genet. 32, 279–305 (1998).
Smith, C.W. & Valcárcel, J. Alternative pre-mRNA splicing: the logic of combinatorial control. Trends Biochem. Sci. 25, 381–388 (2000).
Black, D.L. Protein diversity from alternative splicing: a challenge for bioinformatics and post-genome biology. Cell 103, 367–370 (2000).
Shoemaker, D.D. et al. Experimental annotation of the human genome using microarray technology. Nature 409, 922–927 (2001).
Hu, G.K. et al. Predicting splice variant from DNA chip expression data. Genome Res. 11, 1237–1245 (2001).
Michael, K.L., Taylor, L.C., Schultz, S.L. & Walt, D.R. Randomly ordered addressable high-density optical sensor arrays. Anal. Chem. 70, 1242–1248 (1998).
Ferguson, J.A., Steemers, F.J. & Walt, D.R. High-density fiber-optic DNA random microsphere array. Anal. Chem. 72, 5618–5624 (2000).
Walt, D.R. Bead-based fiber-optic arrays. Science 287, 451–452 (2000).
Gerry, N.P. et al. Universal DNA microarray method for multiplex detection of low abundance point mutations. J. Mol. Biol. 292, 251–262 (1999).
Fan, J.B. et al. Parallel genotyping of human SNPs using generic high-density oligonucleotide tag arrays. Genome Res. 10, 853–860 (2000).
Landegren, U., Kaiser, R., Sanders, J. & Hood, L. A ligase-mediated gene detection technique. Science 241, 1077–1080 (1988).
Nilsson, M., Antson, D.O., Barbany, G. & Landegren, U. RNA-templated DNA ligation for transcript analysis. Nucleic Acids Res. 29, 578–581 (2001).
Kleppe, K., Van de Sande, J.H. & Khorana, H.G. Polynucleotide ligase-catalyzed joining of deoxyribo-oligonucleotides on ribopolynucleotide templates and of ribo-oligonucleotides on deoxyribopolynucleotide templates. Proc. Natl. Acad. Sci. USA 67, 68–73 (1970).
Harper, J.E. & Manley, J.L. Multiple activities of the human splicing factor ASF. Gene Expression 2, 19–29 (1992).
Cáceres, J.F., Stamm, S., Helfman, D.M. & Krainer, A.R. Regulation of alternative splicing in vivo by overexpression of antagonistic splicing factors. Science 265, 1706–1709 (1994).
James, M.C. & Peters, G. Alternative product of the p16/CKDN2A locus connects the Rb and p53 tumor suppressors. Prog. Cell Cycle Res. 4, 71–81 (2000).
Liggett, W.H. Jr., & Sidransky, D. Role of the p16 tumor suppressor gene in cancer. J. Clin. Oncol. 16, 1197–1206 (1998).
Jiang, Z.H. & Wu, J.Y. Alternative splicing and programmed cell death. Proc. Soc. Exp. Biol. Med. 220, 64–72 (1999).
Trowbridge, I.S. & Thomas, M.L. CD45: an emerging role as a protein tyrosine phosphatase required for lymphocyte activation and development. Annu. Rev. Immunol. 12, 85–116 (1994).
Miki, T. et al. Expression cDNA cloning of the KGF receptor by creation of a transforming autocrine loop. Science 251, 72–75 (1991).
Xu, X., Weinstein, M., Li, C. & Deng, C. Fibroblast growth factor receptors (FGFRs) and their roles in limb development. Cell Tiss. Res. 296, 33–43 (1999).
De Moerlooze, L. et al. An important role for the IIIb isoform of fibroblast growth factor receptor 2 (FGFR2) in mesenchymal–epithelial signaling during mouse organogenesis. Development 127, 483–492 (2000).
Hajihosseini, M.K., Wilson, S., De Moerlooze, L. & Dickson, C. A splicing switch and gain-of-function mutation in FgfR2-IIIc hemizygotes causes Apert/Pfeiffer-syndrome-like phenotypes. Proc. Natl. Acad. Sci. USA 98, 3855–3860 (2001).
Miki, T. et al. Determination of ligand-binding specificity by alternative splicing: two distinct growth factor receptors encoded by a single gene. Proc. Natl. Acad. Sci. USA 89, 246–250 (1992).
Yan, G., Fukabori, Y., McBride, G., Nikolaropolous, S. & McKeehan, W.L. Exon switching and activation of stromal and embryonic fibroblast growth factor (FGF)–FGF receptor genes in prostate epithelial cells accompany stromal independence and malignancy. Mol. Cell. Biol. 13, 4513–4522 (1993).
Carstens, R.P., Eaton, J.V., Krigman, H.R., Walther, P.J. & Garcia-Blanco, M.A. Alternative splicing of fibroblast growth factor receptor 2 (FGF-R2) in human prostate cancer. Oncogene 15, 3059–3065 (1997).
Watanabe, M., Ishiwata, T., Nishigai, K., Moriyama, Y. & Asano, G. Overexpression of keratinocyte growth factor in cancer cells and enterochromaffin cells in human colorectal cancer. Pathol. Intl. 50, 363–372 (2000).
Taniguchi, F. et al. Paracrine effects of bFGF and KGF on the process of mouse blastocyst implantation. Mol. Reprod. Dev. 50, 54–62 (1998).
Taniguchi, F. et al. Keratinocyte growth factor in the promotion of human chorionic gonadotropin production in human choriocarcinoma cells. Am. J. Obstet. Gynecol. 182, 692–698 (2000).
Acknowledgements
The authors thank Y.-S. Kwon and other members of the Fu laboratory for their assistance, E. Chudin from Illumina for statistical analyses, and S. Goodison, C. Nelson, H. Li, C. Buckmaster, A. Boyer, M. Ricote, J. Brown, C. Snyder, and E. Brinkman-van der Linden for their gifts of cell lines. D. Che, F. Garcia, and J. Haas provided expert assistance with imaging systems and image analysis. X-D. F. is a Scholar of the Leukemia and Lymphoma Society. This work was supported by grants from the American Cancer Society (J.M.Y) and National Institutes of Health (X-D. F. and M.C.).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Yeakley, J., Fan, JB., Doucet, D. et al. Profiling alternative splicing on fiber-optic arrays. Nat Biotechnol 20, 353–358 (2002). https://doi.org/10.1038/nbt0402-353
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nbt0402-353
This article is cited by
-
Xpress sequencing for express screening
Science-Business eXchange (2012)
-
Timing of plant immune responses by a central circadian regulator
Nature (2011)
-
REMAS: a new regression model to identify alternative splicing events from exon array data
BMC Bioinformatics (2009)