Abstract
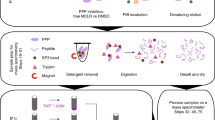
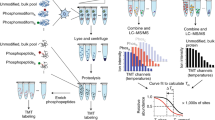
The current progression from genomics to proteomics is fueled by the realization that many properties of proteins (e.g., interactions, post-translational modifications) cannot be predicted from DNA sequence1. Although it has become feasible to rapidly identify proteins from crude cell extracts using mass spectrometry after two-dimensional electrophoretic separation, it can be difficult to elucidate low-abundance proteins of interest in the presence of a large excess of relatively abundant proteins2,3. Therefore, for effective proteome analysis it becomes critical to enrich the sample to be analyzed in subfractions of interest. For example, the analysis of protein kinase substrates can be greatly enhanced by enriching the sample of phosphorylated proteins. Although enrichment of phosphotyrosine-containing proteins has been achieved through the use of high-affinity anti-phosphotyrosine antibodies4, the enrichment of phosphoserine/threonine-containing proteins has not been routinely possible. Here, we describe a method for enriching phosphoserine/threonine-containing proteins from crude cell extracts, and for subsequently identifying the phosphoproteins and sites of phosphorylation. The method, which involves chemical replacement of the phosphate moieties by affinity tags, should be of widespread utility for defining signaling pathways and control mechanisms that involve phosphorylation or dephosphorylation of serine/threonine residues.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lander, E.S. The new genomics: global views of biology. Science 274, 536–539 (1996).
Gygi, S.P., Corthals, G.L., Zhang, Y., Rochon, Y. & Aebersold, R. Evaluation of two-dimensional gel electrophoresis-based proteome analysis technology. Proc. Natl. Acad. Sci. USA 97, 9390–9395 (2000).
Corthals, G.L., Wasinger, V.C., Hochstrasser, D.F. & Sanchez, J.C. The dynamic range of protein expression: a challenge for proteomic research. Electrophoresis 21, 1104–1115 (2000).
Pandey, A. et al. Analysis of receptor signaling pathways by mass spectrometry: identification of vav-2 as a substrate of the epidermal and platelet-derived growth factor receptors. Proc. Natl. Acad. Sci. USA 97, 179–184 (2000).
Taborsky, G. Phosphoproteins. Adv. Protein. Chem. 28, 1–210 (1974).
Annan, W.D., Manson, W. & Nimmo, J.A. The identification of phosphoseryl residues during the determination amino acid sequence in phosphoproteins. Anal. Biochem. 121, 62–68 (1982).
Meyer, H.E., Hoffmann-Posorske, E., Korte, H. & Heilmeyer, L.M., Jr. Sequence analysis of phosphoserine-containing peptides. Modification for picomolar sensitivity. FEBS Lett. 204, 61–66 (1986).
Holmes, C.F. A new method for the selective isolation of phosphoserine-containing peptides. FEBS Lett. 215, 21–24 (1987).
Mega, T., Nakamura, N. & Ikenaka, T. Modifications of substituted seryl and threonyl residues in phosphopeptides and a polysialoglycoprotein by beta-elimination and nucleophile additions. J. Biochem. (Tokyo) 107, 68–72 (1990).
Meyer, H.E., Hoffmann-Posorske, E. & Heilmeyer, L.M., Jr. Determination and location of phosphoserine in proteins and peptides by conversion to S-ethylcysteine. Methods Enzymol. 201, 169–185 (1991).
Meyer, H.E. et al. Strategies for nonradioactive methods in the localization of phosphorylated amino acids in proteins. FASEB J. 7, 776–782 (1993).
Moorhead, G., MacKintosh, R.W., Morrice, N., Gallagher, T. & MacKintosh, C. Purification of type 1 protein (serine/threonine) phosphatases by microcystin–Sepharose affinity chromatography. FEBS Lett. 356, 46–50 (1994).
Fadden, P. & Haystead, T.A. Quantitative and selective fluorophore labeling of phosphoserine on peptides and proteins: characterization at the attomole level by capillary electrophoresis and laser-induced fluorescence. Anal. Biochem. 225, 81–88 (1995).
Jaffe, H., Veeranna & Pant, H.C. Characterization of serine and threonine phosphorylation sites in beta-elimination/ethanethiol addition-modified proteins by electrospray tandem mass spectrometry and database searching. Biochemistry 37, 16211–16224 (1998).
Patchornik, A. & Sokolovsky, M. Nonenzymatic cleavages of peptide chains at the cysteine and serine residues through their conversion into dehydroalanine. I. Hydrolytic and oxidative cleavage of dehydroalanine residues. J. Am. Chem. Soc. 86, 1206–1212 (1964).
Zhang, W. & Chait, B.T. ProFound: an expert system for protein identification using mass spectrometric peptide mapping information. Anal. Chem. 72, 2482–2489 (2000).
Fenyo, D., Qin, J. & Chait, B.T. Protein identification using mass spectrometric information. Electrophoresis 19, 998–1005 (1998).
Sechi, S. & Chait, B.T. Modification of cysteine residues by alkylation. A tool in peptide mapping and protein identification. Anal. Chem. 70, 5150–5158 (1998).
Biemann, K. Contributions of mass spectrometry to peptide and protein structure. Biomed. Environ. Mass Spectrom. 16, 99–111 (1988).
Acknowledgements
This work was supported by the National Center for Research Resources, National Institutes of Health (Grant RR00862 to B.T.C.), and the Eisai Co., Ltd. We thank Fred Cross, Angus Nairn, Derek McLachlin, Yasutaka Takase, and Takashi Owa for useful discussions, Wenzhu Zhang and David Fenyö for development of the protein identification software, and Kappei Tsukahara for assistance in yeast culture and cell handling.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Oda, Y., Nagasu, T. & Chait, B. Enrichment analysis of phosphorylated proteins as a tool for probing the phosphoproteome. Nat Biotechnol 19, 379–382 (2001). https://doi.org/10.1038/86783
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/86783
This article is cited by
-
In situ digestion-assisted multi-template imprinted nanoparticles for efficient analysis of protein phosphorylation
Microchimica Acta (2023)
-
Dendrimer-Modified Silica Nanoparticles for Efficient Enrichment of Low-Concentration Peptides
Applied Biochemistry and Biotechnology (2022)
-
A novel mass spectrometric strategy “BEMAP” reveals Extensive O-linked protein glycosylation in Enterotoxigenic Escherichia coli
Scientific Reports (2016)
-
Low-bias phosphopeptide enrichment from scarce samples using plastic antibodies
Scientific Reports (2015)
-
Mass Spectrometric Analysis of the Cell Surface N-Glycoproteome by Combining Metabolic Labeling and Click Chemistry
Journal of the American Society for Mass Spectrometry (2015)