Abstract
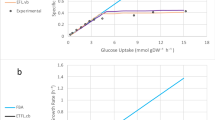
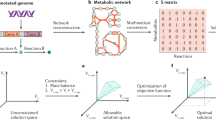
A significant goal in the post-genome era is to relate the annotated genome sequence to the physiological functions of a cell. Working from the annotated genome sequence, as well as biochemical and physiological information, it is possible to reconstruct complete metabolic networks. Furthermore, computational methods have been developed to interpret and predict the optimal performance of a metabolic network under a range of growth conditions. We have tested the hypothesis that Escherichia coli uses its metabolism to grow at a maximal rate using the E. coli MG1655 metabolic reconstruction. Based on this hypothesis, we formulated experiments that describe the quantitative relationship between a primary carbon source (acetate or succinate) uptake rate, oxygen uptake rate, and maximal cellular growth rate. We found that the experimental data were consistent with the stated hypothesis, namely that the E. coli metabolic network is optimized to maximize growth under the experimental conditions considered. This study thus demonstrates how the combination of in silico and experimental biology can be used to obtain a quantitative genotype–phenotype relationship for metabolism in bacterial cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
21 March 2003
Supplementary Table 1 HTML format was replaced with an Excel version
Notes
* Note:The original Supplementary Table 1 (HTML format) has been replaced with an Excel file.
References
TIGR Microbial database: a listing of published microbial genomes and chromosomes and those in progress. (The Institute for Genomic Research, Rockville, MD, 2000). http://www.tigr.org/tdb/mdb/mdbcomplete.html
Karp, P.D. et al. The EcoCyc and MetaCyc databases. Nucleic Acids Res. 28, 56–59 (2000).
Selkov, E. Jr.,, Grechkin, Y., Mikhailova, N. & Selkov, E. MPW: the Metabolic Pathways Database. Nucleic Acids Res. 26, 43–45 (1998).
Ogata, H. et al. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 27, 29–34 (1999).
Karp. P.D., Ouzounis, C. & Paley, S. HinCyc: a knowledge base of the complete genome and metabolic pathways of H. influenzae. Proceedings of the ISMB-96 Conference 4, 116–124 (1996).
Overbeek, R. et al. WIT: integrated system for high-throughput genome sequence analysis and metabolic reconstruction. Nucleic Acids Res. 28, 123–125 (2000).
Edwards, J.S. & Palsson, B.O. Systems Properties of the Haemophilus influenzae Rd metabolic genotype. J. Biol. Chem. 274, 17410–17416 (1999).
Edwards, J.S. & Palsson, B.O. The Escherichia coli MG1655 in silico metabolic genotype: its definition, characteristics, and capabilities. Proc. Natl. Acad. Sci. USA 97, 5528–5533 (2000).
Karp, P.D., Krummenacker, M., Paley, S. & Wagg, J. Integrated pathway-genome databases and their role in drug discovery. Trends Biotechnol. 17, 275–281 (1999).
Goryanin, I., Hodgman, T.C. & Selkov, E. Mathematical simulation and analysis of cellular metabolism and regulation. Bioinformatics 15, 749–758 (1999).
Tomita, M. et al. E-CELL: software environment for whole-cell simulation. Bioinformatics 15, 72–84 (1999).
Bailey, J.E. Mathematical modeling and analysis in biochemical engineering: past accomplishments and future opportunities. Biotechnol. Prog. 14, 8–20 (1998).
Fell, D. Understanding the control of metabolism. (Portland Press, London; 1996).
Domach, M.M., Leung, S.K., Cohn, R.E. & Shuler, M.M. Computer model for glucose-limited growth of a single cell of Escherichia coli B/r-A. Biotechnol. Bioeng. 26, 203–216 (1984).
Palsson, B.O. & Lee, I.D. Model complexity has a significant effect on the numerical value and interpretation of metabolic sensitivity coefficients. J. Theor. Biol. 161, 299–315 (1993).
Barkai, N. & Leibler, S. Robustness in simple biochemical networks. Nature 387, 913–917 (1997).
Liao, J.C. Modelling and analysis of metabolic pathways. Curr. Opin. Biotechnol. 4, 211–216 (1993).
Novak, B. et al. Finishing the cell cycle. J. Theor. Biol. 199, 223–233 (1999).
Kompala, D.S., Ramkrishna, D., Jansen, N.B. & Tsao, G.T. Investigation of bacterial growth on mixed substrates. Experimental evaluation of cybernetic models. Biotechnol. Bioeng. 28, 1044–1056 (1986).
McAdams, H.H. & Arkin, A. It's a noisy business! Genetic regulation at the nanomolar scale. Trends Genet. 15, 65–69 (1999).
Palsson, B.O., Joshi, A. & Ozturk, S.S. Reducing complexity in metabolic networks: making metabolic meshes manageable. Fed. Proc. 46, 2485–2489 (1987).
Jamashidi, N., Edwards, J., Fahland, T., Church, G. & Palsson, B. A computer model of human red blood cell metabolism. Bioinformatics, in press (2001).
Lee, I.D. & Palsson, B.O. A Macintosh software package for simulation of human red blood cell metabolism. Comput. Methods Programs Biomed. 38, 195–226 (1992).
Mulquiney, P.J., Bubb, W.A. & Kuchel, P.W. Model of 2,3-bisphosphoglycerate metabolism in the human erythrocyte based on detailed enzyme kinetic equations: in vivo kinetic characterization of 2,3-bisphosphoglycerate synthase/phosphatase using 13C and 31P NMR. Biochem. J. 342, 567–580 (1999).
Fell, D.A. & Small, J.A. Fat synthesis in adipose tissue. An examination of stoichiometric constraints. J. Biochem. 238, 781–786 (1986).
Edwards, J.S., Ramakrishna, R., Schilling, C.H. & Palsson, B.O. In Metabolic engineering. (eds Lee, S.Y. & Papoutsakis, E.T.) 13–57 (Marcel Dekker, New York, NY; 1999).
Pramanik, J. & Keasling, J.D. Stoichiometric model of Escherichia coli metabolism: incorporation of growth-rate dependent biomass composition and mechanistic energy requirements. Biotechnol. Bioeng. 56, 398–421 (1997).
Sauer, U., Cameron, D.C. & Bailey, J.E. Metabolic capacity of Bacillus subtilis for the production of purine nucleosides, riboflavin, and folic acid. Biotechnol. Bioeng. 59, 227–238 (1998).
Bonarius, H.P.J., Schmid, G. & Tramper, J. Flux analysis of underdetermined metabolic networks: the quest for the missing constraints. Trends Biotechnol. 15, 308–314 (1997).
Varma, A. & Palsson, B.O. Metabolic flux balancing: basic concepts, scientific and practical use. Bio/Technology 12, 994–998 (1994).
Edwards, J.S. Functional genomics and the computational analysis of bacterial metabolism (PhD Thesis). (Department of Bioengineering, University of California San Diego, La Jolla, CA; 1999).
Chvatal, V. Linear programming. (W.H. Freeman, New York, NY; 1983).
Varma, A. & Palsson, B.O. Metabolic capabilities of Escherichia coli: II. Optimal growth patterns. J. Theor. Biol. 165, 503–522 (1993).
Schilling, C.H. & Palsson, B.O. Assessment of the metabolic capabilities of Haemophilus influenzae Rd through a genome-scale pathway analysis. J. Theor. Biol. 203, 249–83 (2000).
Schilling, C.H., Schuster, S., Palsson, B.O. & Heinrich, R. Metabolic pathway analysis: basic concepts and scientific applications in the post-genomic era. Biotechnol. Prog. 15, 296–303 (1999).
Schilling, C.H., Edwards, J.S., Letscher, D. & Palsson, B.O. Pathway analysis and flux balance analysis: a comprehensive study of metabolic systems. Biotechnol. Bioeng. in press (2001).
Sauer, U. et al. Metabolic flux ratio analysis of genetic and environmental modulations of Escherichia coli central carbon metabolism. J. Bacteriol. 181, 6679–6688 (1999).
Klapa, M.I., Park, S.M., Sinskey, A.J. & Stephanopoulos, G. Metabolite and isotopomer balancing in the analysis of metabolic cycles: I. Theory. Biotechnol. Bioeng. 62, 375–391 (1999).
DeRisi, J.L., Iyer, V.R. & Brown, P.O. Exploring the metabolic and genetic control of gene expression on a genomic scale. Science 278, 680–686 (1997).
Richmond, C.S., Glasner, J.D., Mau, R., Jin, H. & Blattner, F.R. Genome-wide expression profiling in Escherichia coli K-12. Nucleic Acids Res. 27, 3821–3835 (1999).
Hacia, J.G., Brody, L.C., Chee, M.S., Fodor, S.P.A. & Collins, F.S. Detection of heterozygous mutations in BRCA1 using high-density oligonucleotide arrays and two-colour fluorescence analysis. Nat. Genet. 14, 441–447 (1996).
Link, A.J., Robison, K. & Church, G.M. Comparing the predicted and observed properties of proteins encoded in the genome of Escherichia coli K-12. Electrophoresis 18, 1259–1313 (1997).
Vanbogelen, R.A., Abshire, K.Z., Moldover, B., Olson, E.R. & Neidhardt, F.C. Escherichia coli proteome analysis using the gene-protein database. Electrophoresis 18, 1243–1251 (1997).
Gygi, S.P. et al. Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat. Biotechnol. 17, 994–999 (1999).
Schilling, C.H., Edwards, J.S. & Palsson, B.O. Towards metabolic phenomics: analysis of genomic data using flux balances. Biotechnol. Prog. 15, 288–295 (1999).
Neidhardt, F.C. (ed.) Escherichia coli and Salmonella: cellular and molecular biology. (ASM Press, Washington, DC; 1996).
Maniatis, T., Fritsch, E.F. & Sambrook, J. Molecular cloning: a laboratory manual. (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY; 1982).
Neidhardt, F.C. & Umbarger, H.E. In Escherichia coli and Salmonella: cellular and molecular biology. (ed. Neidhardt, F.C.) 13–16 (ASM Press, Washington, DC; 1996).
Strang, G. Linear algebra and its applications. (Saunders, Fort Worth, TX; 1988).
Varma, A., Boesch, B.W. & Palsson, B.O. Stoichiometric interpretation of Escherichia coli glucose catabolism under various oxygenation rates. Appl. Environ. Microbiol. 59, 2465–2473 (1993).
Covert, M.W. et al. Metabolic modeling of microbial strains in silico. Trends Biochem. Sci. in press (2001).
Acknowledgements
This work was funded by the NIH (GM57089) and the NSF (MCB 98-73384 and BES 98-14092). We would like to thank Christophe Schilling and George M. Church for insightful discussions during the preparation of this manuscript, and Markus Covert and Iman Famili for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Table 1
Note: The original Supplementary Table 1 (HTML format) has been replaced with an Excel file. (XLS 145 kb)
Rights and permissions
About this article
Cite this article
Edwards, J., Ibarra, R. & Palsson, B. In silico predictions of Escherichia coli metabolic capabilities are consistent with experimental data. Nat Biotechnol 19, 125–130 (2001). https://doi.org/10.1038/84379
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/84379
This article is cited by
-
Dynamic proteome trade-offs regulate bacterial cell size and growth in fluctuating nutrient environments
Communications Biology (2023)
-
Periodic synchronization of isolated network elements facilitates simulating and inferring gene regulatory networks including stochastic molecular kinetics
BMC Bioinformatics (2022)
-
Addressing uncertainty in genome-scale metabolic model reconstruction and analysis
Genome Biology (2021)
-
Microeconomics of Metabolism: The Warburg Effect as Giffen Behaviour
Bulletin of Mathematical Biology (2021)
-
Synergistic effects of repair, resilience and retention of damage determine the conditions for replicative ageing
Scientific Reports (2020)