Abstract
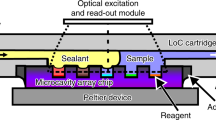
We have developed a method for anchored amplification on a microchip array that allows amplification and detection of multiple targets in an open format. Electronic anchoring of sets of amplification primers in distinct areas on the microchip permitted primer-primer interactions to be reduced and distinct zones of amplification created, thereby increasing the efficiency of the multiplex amplification reactions. We found strand displacement amplification (SDA) to be ideal for use in our microelectronic chip system because of the isothermal nature of the assay, which provides a rapid amplification system readily compatible with simple instrumentation. Anchored SDA supported multiplex DNA or RNA amplification without decreases in amplification efficiency. This microelectronic chip-based amplification system allows multiplexed amplification and detection to be performed on the same platform, streamlining development of any nucleic acid-based assay.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Debouck, C. & Goodfellow, P.N. DNA microarrays in drug discovery and development. Nat. Genet. 21, 48–50 (1999).
Schena, M., Shalon, D., Davis, R.W. & Brown, P.O. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science 270, 467–470 (1995).
Eggers, M. & Ehrlich, D. A review of microfabricated devices for gene-based diagnostics. Hematol. Pathol. 9, 1–15 (1995).
Chee, M. et al. Accessing genetic information with high-density DNA arrays. Science 274, 610–614 (1996).
Lockhart, D.J. et al. Expression monitoring by hybridization to high-density oligonucleotide arrays. Nat. Biotechnol. 14, 1675–1680 (1996).
Ferguson, J.A, Boles, T.C., Adams, C.P. & Walt, D.R. A fiber-optic DNA biosensor microarray for the analysis of gene expression. Nat. Biotechnol. 14, 1681–1684 (1996).
Weiler, J. & Hoheisel, J.D. Combining the preparation of oligonucleotide arrays and synthesis of high quality primers. Anal. Biochem., 15, 218–227 (1996).
Edwards, M.C. & Gibbs, R.A. Multiplex PCR: advantages, development and applications. PCR Methods Appl. 3, S65–S75 (1994).
Rychlik, W. Selection of primers for polymerase chain reaction. Mol. Biotechnol. 3, 129–134 (1995).
Shuber, A.P., Grondin, V.J. & Klinger, K.W. A simplified procedure for developing multiplex PCRs. Genome Res. 5, 488–493 (1995).
Brownie, J. et al. The elimination of primer–dimer accumulation in PCR. Nucleic Acids Res. 25, 3235–3241 (1997).
Sosnowski, R.G., Tu, E., Butler, W.F., O'Connell, J.P. & Heller, M.J. Rapid determination of single-base mismatch mutations in DNA hybrids by direct electric field control. Proc. Natl. Acad. Sci. USA 94, 1119–1123 (1997).
Edman, C.F. et al. Electric field directed nucleic acid hybridization on microchips. Nucleic Acids Res. 25, 4907–4914 (1997).
Walker, G.T, Little, M.C., Nadeau, J.G. & Shank, D.D. Isothermal, in vitro amplification of DNA by a restriction enzyme/DNA polymerase system. Proc. Natl. Acad. Sci. USA 89, 392–396 (1992).
Romano, J.W., Williams, K.G., Shurtliff, R.N., Ginocchio, C. & Kaplan, M. NASBA technology: isothermal RNA amplification in qualitative and quantitative diagnostics. Immunol. Invest. 26, 15–28 (1997).
Pasternack, R., Vuorinen, P, & Miettinen, A. Evaluation of the gen-probe Chlamydia trachomatis transcription-mediated amplification assay with urine specimens from women. J. Clin. Microbiol. 35, 676–678 (1997).
Lizardi, P.M. et al. Mutation detection and single molecule counting using isothermal rolling-circle amplification. Nat. Genet. 19, 225–232 (1998).
Spargo, C.A. et al. Detection of M. tuberculosis DNA using thermophilic strand displacement amplification. Mol. Cell. Probes 10, 247–256 (1996).
Svensson, P.J. & Dahlback, B. Resistance to activated protein C as a basis for venous thrombosis. N. Engl. J. Med., 300, 517–522 (1994).
Bertina, R.M. et al. Mutation in blood coagulation factor V associated with resistance to activated protein C. Nature 369, 64–67 (1994).
Nycz, C.M., Dean, C.H., Haaland, P.D., Spargo, C.A. & Walker, G.T. Quantitative reverse transcription strand displacement amplification: quantitation of nucleic acids using an isothermal amplification technique. Anal. Biochem. 259, 226–234 (1998).
Hatch, A., Sano, T., Misasi, J. & Smith, C.L. Rolling circle amplification of DNA immobilized on solid surfaces and its application to multiplex mutation detection. Genet. Anal. 15, 35–40 (1999).
Adams, C.P. & Kron, S.J. Method for performing amplification of nucleic acid with two primers bound to a single solid support. US PTO, US 5,641,658 (1997).
Van Gelder R.N. et al. Amplified RNA synthesized from limited quantities of heterogeneous cDNA. Proc. Natl. Acad. Sci. USA 87, 1663–1667 (1990).
Halpern, N.A. & Brentjens, T. Point of care testing informatics: The critical care-hospital interface. Crit. Care Clin. 15, 577–591 (1999).
Acknowledgements
The authors would like to gratefully acknowledge the intellectual contributions and support of Drs. Terry Walker, Dan MacLaurin, and Doug Malinowski, and Cathy Spargo of Becton-Dickinson, and Drs. Tina S. Nova, Jim P. O'Connell, Michael Heller, John Carrino, and Prashant Mehta of Nanogen, Inc. throughout the course of this work. The authors are especially grateful to Drs. Ray Radtke, Lana Feng, and Geoff Landis, and Dana Vollmer of Nanogen, Inc. for their contributions in support of permeation layer, primer design, and SDA in earlier stages of this work. Supported under Award number 97-LB-VX-0004 from the Office of Justice Programs, National Institute of Justice, Department of Justice. Points of view in this document are those of the authors and do not necessarily represent the official position of the US Department of Justice.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Westin, L., Xu, X., Miller, C. et al. Anchored multiplex amplification on a microelectronic chip array. Nat Biotechnol 18, 199–204 (2000). https://doi.org/10.1038/72658
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/72658
This article is cited by
-
On chip quadruplex priming amplification for quantitative isothermal diagnostics
Biomedical Microdevices (2018)
-
Parallel solid-phase isothermal amplification and detection of multiple DNA targets in microliter-sized wells of a digital versatile disc
Microchimica Acta (2016)
-
Extraction, amplification and detection of DNA in microfluidic chip-based assays
Microchimica Acta (2014)
-
Integration of sample pretreatment, μPCR, and detection for a total genetic analysis microsystem
Microchimica Acta (2014)
-
Improved loop-mediated isothermal amplification for HLA-DRB1 genotyping using RecA and a restriction enzyme for enhanced amplification specificity
Immunogenetics (2013)