Abstract
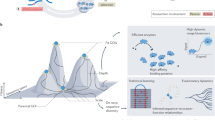
We describe a new method of random mutagenesis that employs the addition of peptide tails with random sequences to the C–terminal of enzyme molecules. A mutant population of catalase I from Bacillus stearothermophilus prepared by this method has a diversity in thermostability and enzyme activity equal to that obtained after random point mutagenesis. When a triple mutant of catalase I (I108T/D130N/I222T)—the thermostability of which is much lower than that of the wild type—was subjected to random elongation mutagenesis, we generated a mutant population containing only mutants with higher thermostability than the triple mutant. Some had an even higher stability than the wild–type enzyme, whose thermostability is considered to be optimized. These results indicate that peptide addition expands the protein sequence space resulting in a new fitness landscape. The enzyme can then move along the routes of the new landscape until it reaches a new optimum. The combination of random elongation mutagenesis with random point mutagenesis should be a useful approach to the in vitro evolution of proteins with new properties.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
You, L. and Arnold, F.H. 1996. Directed evolution of subtilisin E in Bacillus subtilis to enhance total activity in aqueous dimethylformamide. Protein Eng. 9: 77–83.
Moore, J.C. and Arnold, F.H. 1996. Directed evolution of a para–nitrobenzyl esterase for aqueous–organic solvents. Nat. Biotechnol. 14: 458– 467.
Zhang, J.–H., Dawes, G., and Stemmer, W.P.C. 1997. Directed evolution of a fucosidase from a galactosidase by DNA shuffling and screening. Proc. Natl. Acad. Sci. USA 94: 4504–4509.
Matsuura, T., Yomo, T., Trakulnaleamsai, S., Ohashi, Y., Yamamoto, K., and Urabe, I. 1998. Nonadditivity of mutational effects on the properties of catalase I and its application to efficient directed evolution. Protein Eng. 11: 789– 795.
Aita, T. and Husimi, Y. 1996. Fitness spectrum among random mutants on Mt. Fuji–type fitness landscape. J. Theor. Biol. 182: 469–485.
Trakulnaleamsai, S., Yomo, T., Yoshikawa, M., Aihara, S., and Urabe, I. 1995. Experimental sketch of landscapes in protein sequence space. J. Ferment. Bioeng. 79: 107–118.
Trakulnaleamsai, S., Aihara, S., Miyai, K., Suga, Y., Sota, M., Yomo, T. et al. 1992. Revised sequence and activity of Bacillus stearothermophilus catalase I (formerly peroxidase). J. Ferment. Bioeng. 74: 234–237.
Yomo, T., Yamano, T., Yamamoto, K., and Urabe, I. 1997. General equation of steady–state enzyme kinetics using net rate constants and its application to the kinetic analysis of catalase reaction. J. Theor. Biol. 188: 301–312.
Kobayashi, C., Suga, Y., Yamamoto, K., Yomo, T., Ogasahara, K., Yutani, K. et al. 1997. Thermal conversion from low– to high–activity forms of catalase I from Bacillus stearothermophilus. J. Biol. Chem. 272: 23011–23016.
Wells, J.A. 1990. Additivity of mutational effects in proteins. Biochemistry 29: 8509–8517 .
Triggs–Rain, B.L. and Loewen, P.C. 1987 . Physical characterization of katG encoding catalase HPI of Escherichia coli. Gene 52: 121–128.
Loprasert, S., Urabe, I., and Okada, H. 1990. Overproduction and single–step purification of Bacillus stearothermophilus peroxidase in Escherichia coli. Appl. Microbiol. Biotechnol. 32: 690– 692.
Maniatis, T., Fritsch, E.F., and Sambrook, J. 1982. Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
Laemmli, U.K. 1970. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227: 680–685.
Acknowledgements
This work was supported in part by Grants–in–Aid (10450310, 10145234, and 07280101) from the Ministry of Education, Science, Sports, and Culture, Japan, and was performed as a part of the Research and Development Project of Industrial Science and Technology Frontier Program supported by NEDO.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Matsuura, T., Miyai, K., Trakulnaleamsai, S. et al. Evolutionary molecular engineering by random elongation mutagenesis. Nat Biotechnol 17, 58–61 (1999). https://doi.org/10.1038/5232
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/5232
This article is cited by
-
A novel framework for engineering protein loops exploring length and compositional variation
Scientific Reports (2021)
-
Enhancing Catalytic Efficiency of an Actinoplanes utahensis Echinocandin B Deacylase through Random Mutagenesis and Site-Directed Mutagenesis
Applied Biochemistry and Biotechnology (2020)
-
C-Terminal Flanking Peptide Stabilized the Catalytic Domain of a Recombinant Bacillus subtilis Endo-β-1, 4-Glucanase
The Protein Journal (2013)
-
Enhanced thermal stability and specific activity of Pseudomonas aeruginosa lipoxygenase by fusing with self-assembling amphipathic peptides
Applied Microbiology and Biotechnology (2013)
-
Effect of the C-terminal domains and terminal residues of catalytic domain on enzymatic activity and thermostability of lichenase from Clostridium thermocellum
Biotechnology Letters (2010)