Abstract
We present an extension of the Minimum Information about any (x) Sequence (MIxS) standard for reporting sequences of uncultivated virus genomes. Minimum Information about an Uncultivated Virus Genome (MIUViG) standards were developed within the Genomic Standards Consortium framework and include virus origin, genome quality, genome annotation, taxonomic classification, biogeographic distribution and in silico host prediction. Community-wide adoption of MIUViG standards, which complement the Minimum Information about a Single Amplified Genome (MISAG) and Metagenome-Assembled Genome (MIMAG) standards for uncultivated bacteria and archaea, will improve the reporting of uncultivated virus genomes in public databases. In turn, this should enable more robust comparative studies and a systematic exploration of the global virosphere.
Similar content being viewed by others
Main
Current estimates are that virus particles massively outnumber live cells in most habitats1,2, but only a tiny fraction of viruses have been cultivated in the laboratory. An unprecedented diversity of viruses are being discovered through culture-independent sequencing3. Progress has been made in reconstructing genomes of uncultivated viruses de novo, from biotic and abiotic environments, without laboratory isolation of the virus–host system. For example, in the past 2 years, more than 750,000 uncultivated virus genomes (UViGs) have been identified in metagenome and metatranscriptome datasets4,5,6,7,8,9, five times the total number of genomes sequenced from virus isolates (Fig. 1), and UViGs already represent ≥95% of the taxonomic diversity in publicly available virus sequences10,11. Although double-stranded DNA (dsDNA) genomes are over-represented in UViGs because most metagenomic protocols exclusively target dsDNA, UViGs nonetheless enable an assessment of global virus diversity and an evaluation of structure and drivers of viral communities. UViGs also contribute to improving our understanding of the evolutionary history of viruses and virus–host interactions.
Genome sequences from isolates (blue and green) or from UViGs (yellow) are shown. For genomes from isolates, the total number of genomes (blue) and the number of 'reference' genomes (green) are shown. Data were downloaded using the queries “Viruses[Organism] AND srcdb_refseq[PROP] NOT wgs[PROP] NOT cellular organisms[ORGN] NOT AC_000001:AC_999999[PACC]” for reference genomes and “Viruses[Organism] NOT cellular organisms[ORGN] NOT wgs[PROP] NOT AC_000001:AC_999999[pacc] NOT gbdiv syn[prop] AND nuccore genome samespecies[Filter]” for total number of virus genomes, on the NCBI nucleotide database portal (https://www.ncbi.nlm.nih.gov/nuccore) in January 2018. Genomes from the influenza virus database (https://www.ncbi.nlm.nih.gov/genomes/FLU/Database/nph-select.cgi?go=genomeset) were also added to the total number of virus genomes. UViGs can be assembled from metagenomes, from proviruses identified in microbial genomes, or from single-virus genomes, and estimated total UViG numbers were obtained by compiling data from the literature and from the total number of sequences in the IMG/VR database in January 2017, January 2018 and July 2018 (https://img.jgi.doe.gov/vr/)11. UpViG, uncultivated provirus.
Analysis and interpretation of standalone genomes present substantial challenges, whether the genomes are eukaryotic, bacterial, archaeal or viral. To address these challenges, MISAG and MIMAG standards were drafted to improve the quality of reporting of microbial genomes derived from single cell or metagenome sequences, which are often incomplete12. Although some aspects of MISAG and MIMAG can be applied to UViGs, the extraordinary diversity of viral genome composition and content, replication strategies, and hosts means that the completeness, quality, taxonomy and ecology of UViGs need to be evaluated via virus-specific metrics.
The Genomic Standards Consortium (http://gensc.org) maintains metadata checklists for MIxS, encompassing genome and metagenome sequences13, marker gene sequences14 and single amplified and metagenome-assembled bacterial and archaeal genomes12. Here we present a set of standards that extend the MIxS checklists to include identification, quality assessment, analysis and reporting of UViGs (Table 1 and Supplementary Tables 1 and 2), together with recommendations on how to perform these analyses. We provide a metadata checklist for database submission and publication of UViGs designed to be flexible enough to accommodate technological and methodological changes over time (Table 1 and Supplementary Table 1). The information gathered through the MIUViG checklist can be directly submitted with new UViG sequences to International Nucleotide Sequence Database Collaboration (INSDC) member databases—the DNA Database of Japan (DDBJ), the European Molecular Biology Laboratory–European Bioinformatics Institute (EMBL-EBI) and US National Center for Biotechnology Information (NCBI)—which will host and display checklist metadata alongside the UViG sequence. These MIUViG standards should also be used along with existing guidelines for virus genome analysis, including those issued by the International Committee on Taxonomy of Viruses (ICTV), which recently endorsed the incorporation of UViGs into the official virus classification scheme15 (https://talk.ictvonline.org). Although MIUViG standards and best practices were designed for genomes of viruses infecting microorganisms, they can also be applied to viruses infecting animals, fungi and plants, and are compatible with standards that are already in place for epidemiological analysis of these viruses16 (Supplementary Table 3).
Recovery of UViGs after virus enrichment
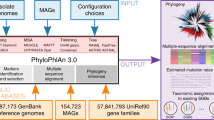
UViGs can be retrieved from datasets enriched for virus genomes, namely viral metagenomes and single-virus genomes (Fig. 2). Viral metagenomes are usually obtained through a combination of filtration steps, DNase or RNase treatments, and RNA or DNA extraction depending on the targeted viruses, then reverse transcription (to find RNA viruses) and shotgun sequencing3,17,18,19. Targeted sequence capture methods can be applied to recover specific virus groups (Fig. 2), and these methods have proven especially useful when viruses are present in small amounts (for example, clinical samples)20. Single-virus methods use flow cytometry to sort individual viral particles before genome amplification and sequencing, to produce viral single amplified genomes (SAGs)9,21,22,23 (Fig. 2). Viral metagenomes and single-virus genomes are usually sequenced with short-read, high-throughput technologies, such as Illumina sequencing, and assembled by algorithms similar to those used for microbial genomes and metagenomes. However, owing to their relatively small genome size (92% of virus genomes in the NCBI Viral RefSeq database are <100 kb)10, short read-based genome assemblies could soon be superseded by long-read sequencing technologies24 (for example, PacBio zero-mode waveguide technology or Oxford Nanopore Technology nanopore sequencing; Fig. 2). Sequencing virus genomes from a single template would notably enable the identification of individual genotypes in mixed populations.
Schematic of methods used to obtain UViGs. Steps that have been adapted from those used to assemble MAGs and SAGs12 or added for UViG are shown for sample preparation (orange) and bioinformatics analysis (blue). Steps specifically required for virus targeting and identification are highlighted in bold. *For viruses with short genomes, long-read technologies can provide complete genomes from shotgun sequencing in a single read, bypassing the assembly step24. **Targeted sequence capture can be used to recover viral genomes from a known virus group. These genomes can be recovered from samples in which they represent a small fraction of the templates (for example, clinical samples20).
The main advantages of datasets produced after enrichment for viruses are good de novo assembly of both abundant and rare viruses, increased confidence that the sequence is of viral origin, and the ability to sequence both active and 'inactive' or 'cryptic' viruses (i.e., viruses that are present in the sample but cannot infect). However, virus-enriched datasets can have over-representation of virulent viruses with high burst size (high number of virus particles released from each infected cell) and under-representation of larger viruses with capsids ≥0.2 μm, such as giant viruses, as a result of the selective filtration steps used25. Furthermore, in silico approaches are often the only option available to determine the host range of UViGs obtained from virus-enriched samples.
Recovery of UViGs without enrichment
Virus sequences are also present in non-virus-enriched datasets, including sorted cells, tissues, or environmental samples collected on 0.2 μm filters4,26,27,28. These sequences could originate from viruses that are replicating in cells, from temperate viruses (proviruses or prophages) that are either integrated into host genomes or present as episomal elements in the host cell, or from free virus particles present in samples.
Analyzing datasets without virus enrichment has several advantages. It can detect lytic, temperate and persistent infection, it overcomes some of the biases arising from the size-based selection of virus particles, and it can be applied to any metagenome. However, UViGs from non-virus-enriched datasets may be biased toward viruses that infect the dominant host cell in the sample, and rare viruses or those infecting rare hosts could be under-represented or absent. Finally, comparisons between virus-enriched and non-virus-enriched datasets suggest that analyzing UViGs across different size fractions and sample types is valuable for exploring the virus genome sequence space29 (Supplementary Fig. 1 and Supplementary Note 1).
Computational identification of viral sequences
Regardless of the type of dataset, the viral origin of UViGs must be validated because even samples enriched for virus particles still contain a substantial amount of cellular DNA30. Contamination can arise either from difficulty in separating virus particles from cellular fractions (for example, ultra-small bacteria31) or from the capture of extracellular DNA in the virus fraction. Cellular sequences can also derive from cell genome fragments that are encased in virus capsids or comparable particles (for example, via transduction), DNA-containing membrane vesicles, or gene transfer agents32,33,34.
Several bioinformatic tools and protocols have been developed to identify sequences from bacteriophages and archaeal viruses35,36,37,38; eukaryotic viruses39; or combinations of bacteriophages, archaeal viruses and large eukaryotic viruses40 (Supplementary Table 4). These approaches rely on a few characteristics, such that a sequence is considered viral if it is significantly similar to known viruses (in terms of gene content or nucleotide usage pattern) or if it is unrelated to any known virus and cellular genome but contains one or more hallmark virus genes. UViGs must therefore be accompanied by a list of virus detection tool(s) and protocol(s) used, together with any thresholds applied (Table 1 and Supplementary Table 1).
Identification of integrated proviruses and their precise boundaries in the host genome is problematic (Box 1). Notably, no high-throughput approach can accurately distinguish active proviruses (still able to replicate and produce virions) from inactive proviral remnants of a past infection28. Thus, although prediction methods are improving, UViGs identified as proviruses should be clearly marked as such, so that these caveats are clear (Table 1 and Supplementary Table 1).
Estimating quality of UViGs
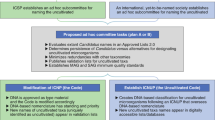
We propose three categories of UViG sequences: genome fragment(s), high-quality draft genomes and finished genomes (Fig. 3 and Table 2). These categories mirror those in MISAG and MIMAG12, and they are matched to categories already proposed for complete-genome sequencing of small viruses in epidemiology and surveillance16 (Supplementary Table 3). UViG quality is more challenging to evaluate than metagenome-assembled genomes (MAGs) or SAGs because most viruses lack conserved sets of single-copy marker genes that can be used to estimate draft genome completeness. However, exceptions exist, such as large eukaryotic dsDNA viruses. To date, researchers have estimated UViG sequence completeness by identifying circular contigs or contigs with inverted terminal repeats as putative complete genomes. For linear contigs, completeness is estimated by comparison to reference genome sequences and typically requires a taxonomic assignment to a (candidate) (sub)family or genus because genome length is relatively homogeneous at these ranks (±10%; Supplementary Fig. 2 and Supplementary Table 5). This assignment can be based on the detection of specific marker genes, such as clade-specific viral orthologous groups (Supplementary Table 6), or based on genome-based classification tools (see “Taxonomy of UViGs”). Estimating completeness is more difficult for segmented genomes, which require either a closely related reference genome or additional in vitro experiments16. A detailed example of how this quality tier classification can be performed on the Global Ocean Virome dataset7 is presented in Supplementary Note 2 and Supplementary Table 7.
“Functional potential” is functional annotation used in gene content analysis. “Host prediction” is the application of different in silico host prediction tools. “Taxonomic classification” is classification of the contig to established groups using marker genes or gene content comparison. “Diversity and distribution” includes vOTU clustering and relative abundance estimation through metagenome read mapping, at the geographical scale or across anatomical sites for host-associated datasets. “New taxonomic groups” concerns the delineation of new proposed groups (for example, families or genera) based exclusively on UViG sequences. “New reference species” refers to the proposal of a new entry in ICTV (https://talk.ictvonline.org/files/taxonomy-proposal-templates/). *Some of these approaches require a minimum contig size—for example, contigs ≥10 kb for taxonomic classification based on gene content59 or diversity estimation47—and will not be applicable to every genome fragment.
Contigs or genome bins representing <90% of the expected genome length, or for which no expected genome length can be determined, would be considered genome fragments. This category might include UViG fragments large enough to be assigned to known virus groups on the basis of gene content and average nucleotide identity. However, high-quality draft or finished genomes are required to establish new taxa (Fig. 3). Sequences from UViG fragments can be used in phylogenetic and diversity studies, either as references for virus operational taxonomic units (see Supplementary Note 4), or through the analysis of virus marker genes encoded in these genome fragments; for example, capsid proteins, terminases, ribonucleotide reductases and DNA- or RNA-dependent RNA polymerases41,42,43,44,45,46. Similarly, UViG fragments can be analyzed to assess the functional gene complement of unknown viruses or link them to potential hosts. Importantly, current methods for automatic virus sequence identification35,36,37,38,39,40 cannot reliably identify short (<10 kb) viral sequences, which should be interpreted with utmost caution.
Contigs or genome bins either predicted as complete or representing ≥90% of the expected genome sequence are high-quality drafts, consistent with standards for microbial genomes12. Repeat regions may lead to erroneous assembly of partial genomes as circular contigs47. Thus, the length of the assembled circular contig should be considered when assessing UViG completeness (Box 1). For UViGs not derived from a consensus assembly, such as single long reads, base calling quality >99% on average (phred score >20) is needed to assign a “high-quality draft” label. Genome sequences assembled into a single contig, or one per segment, with extensive manual review and annotation, can be labeled “finished genomes.” Annotation must include identification of putative gene functions; structural, replication or lysogeny modules; and transcriptional units. The “finished genomes” category is reserved for only the highest quality, manually curated UViGs and is required for the establishment of new virus species (Fig. 3 and Table 2).
Unlike that of SAGs and MAGs12, quality estimation of UViGs does not include a genome contamination threshold. Contamination issues are most prominent in the case of genome bins, whereas most UViGs are represented by a single contig for which in silico simulations have shown that chimeric sequences are rare and present at <2% (ref. 47). In addition, no tools exist to automatically estimate UViG contamination, and thus this information is not included in the current MIUViG checklist. A future updated version of the MIUViG checklist may, however. For include contamination thresholds if such a tool were to be developed. For example, such a tool might exploit single-copy marker genes (once these have been defined for a broader range of viruses) or it might use coverage by metagenome reads, which should in principle be evenly distributed along the genome with no major deviance, except for highly conserved genes.
Annotation of UViGs
Functional annotation of UViGs comprises the following tasks: predicting features in the genome sequence, such as protein-coding genes, tRNAs and integration sites; assigning functions to as many predicted features as possible; and assigning the remaining hypothetical proteins to uncharacterized protein families. Annotation pipelines have been established for different types of viruses48,49, and large differences between viral genome types likely preclude the development of a single tool able to annotate every virus50. Therefore, we recommend that software used to annotate UViGs be reported (Supplementary Table 1).
The choice of methods and reference databases used to annotate predicted proteins should be clearly stated. Homologs of novel virus genes may not be detected with standard methods for pairwise sequence similarity detection, such as BLAST, but instead require the use of more sensitive profile similarity approaches, such as HMMER51, PSI-BLAST52 or HHPred53 (Supplementary Table 8; reviewed in ref. 54). Although sequence profiles for many protein families have been collected, they frequently remain unassociated with any specific function. Therefore, UViG analyses should always report (i) feature prediction method(s), (ii) sequence similarity search method(s), and (iii) database(s) searched (Box 1 and Supplementary Table 1).
Taxonomy of UViGs
Taxonomic classification can provide information on the relationship of a UViG with known viruses. Although the information and criteria used for virus classification have changed over time, virus classification has now converged to genome-based analyses15 (Box 2). The ICTV established specific demarcation criteria for each virus group (Supplementary Table 9) owing to the vast range of viral genomes, mutation rates and evolution. Recently, a consensus has emerged on using whole-genome average nucleotide identity for classification at the species rank, which is used in downstream ecological, evolutionary and functional studies. This consensus was reached through analysis of published population genetics studies55,56 and gene content comparison of NCBI RefSeq10 virus genomes57,58,59 (Supplementary Note 3 and Supplementary Fig. 3). We propose to formalize the use of species-rank virus groups and to name these “virus operational taxonomic units” (vOTUs) to avoid confusion because species groups have been variously named “viral population,” “viral cluster” or “contig cluster” in the literature4,7,60. We suggest standard thresholds of 95% average nucleotide identity over 85% alignment fraction (relative to the shorter sequence) on the basis of a comparison of sequences currently available in NCBI RefSeq10 and IMG/VR11 (Supplementary Note 3 and Supplementary Figs. 3 and 4). Although partial genomes remain challenging to classify, these common thresholds will enable comparative analyses (Supplementary Fig. 5). In addition, vOTU reports should include the clustering method and cutoff, the reference database used (if any), and the genome alignment approach because small differences have been observed between different methods61 (Supplementary Table 1).
For higher taxonomic ranks than species, no consensus has been reached on which approach should be used, although several have been proposed58,59,62,63,64,65,66. Keeping this in mind, UViG reports including taxonomy must clearly indicate the methods and cutoffs applied, and any new taxon must be highlighted as preliminary (for example, “genus-rank cluster,” “putative genus” or “candidate genus,” but not simply “genus,” as this category is reserved for ICTV-recognized groups; Supplementary Table 1). Authors should submit formal taxonomic proposals to the ICTV for consideration (https://talk.ictvonline.org/files/taxonomy-proposal-templates/).
Finally, information about the nature of the genome and mode of expression (i.e., Baltimore classification67) should be included in the UViG description. Similarly, the predicted segmentation state of the genome (segmented or nonsegmented) should be reported, typically derived from taxonomic classification and comparison with the closest references (Supplementary Table 1).
In silico host prediction
Once a new virus genome has been assembled, an important step toward understanding the ecological role of the associated virus is to predict its host(s). In silico approaches are often the only option for UViGs (reviewed in ref. 68; Supplementary Table 10). These can be separated into four main types. First, hosts can be predicted with relatively high precision on the basis of sequence similarity between the UViG and a reference virus genome when a closely related virus is available69,70. Second, hosts can be predicted on the basis of sequence similarities between a UViG and a host genome. These sequence similarities can range from short exact matches (∼20–100 bp), which include CRISPR spacers4,7,68,71, to longer (>100 bp) nucleotide sequence matches, including proviruses integrated into a larger host contig26,68,72,73 (Supplementary Table 10). Host-range predictions based on sequence similarity are the most reliable but require that a closely related host genome has been sequenced68. Third, host taxonomy from domain down to genus rank can be predicted from nucleotide usage signatures reflecting coevolution between virus and host genomes in terms of G+C content, k-mer frequency and codon usage26,74,75. These approaches are usually less specific than sequence similarity–based ones and cannot reliably predict host range below the genus rank, but can provide a predicted host for a larger number of UViGs7 (Supplementary Table 10). Finally, host predictions can be computed from a comparison of abundance profiles of host and virus sequences across spatial or temporal scales, either through abundance correlation25,76,77,78 or through more sophisticated model-based interaction predictors79. Although few datasets are available for robust evaluation of host prediction based on comparison of abundance profiles, we expect this approach to become more powerful and relevant as high-resolution time-series metagenomics becomes more common.
As all these bioinformatic approaches remain predictive, it is crucial that robust false-discovery rate estimations are reported (Supplementary Table 1). Moreover, computational tools do not predict quantitative infection characteristics (for example, infection rate or burst size), which are important for understanding the impacts of viruses on host biology, and thus far only apply to viruses infecting bacteria or archaea. Nevertheless, these predictions are important guides for subsequent in silico, in vitro and in vivo studies, including experimental validation to unequivocally demonstrate a viral infection of a given microbial host. Host predictions should be reported along with details regarding the specific tool(s) used and, importantly, their estimated accuracy as derived either from published benchmarks or from tests conducted in the study (Supplementary Table 1). This information will allow virus–host databases69,80 to progressively incorporate UViGs while still controlling for the sensitivity and accuracy of the predictions provided to users.
Reporting UViGs
We recommend the following best practice for sharing and archiving UViGs and UViG-related data: data publication should center on the data resources of INSDC (http://www.insdc.org/) through one of the member databases, at DDBJ (https://www.ddbj.nig.ac.jp/index-e.html), EMBL-EBI's European Nucleotide Archive (ENA; https://www.ebi.ac.uk/ena) or NCBI (GenBank and the Sequence Read Archive; https://www.ncbi.nlm.nih.gov/nucleotide). If needed, INSDC database curators can be contacted directly for large-scale batch dataset submissions. Where new datasets are generated as part of a UViG study, sequenced samples should be described according to the environment-relevant MIxS checklists and raw read data should be submitted. High-quality and finished UViGs should be submitted as assemblies, the former reported as “draft” accompanied by the required metadata (Table 1). Incomplete assemblies may be submitted, but they must be accompanied by the required metadata (Table 1 and Supplementary Table 1).
Where available, annotation and taxonomic classification should be submitted to INSDC, and occurrence and abundance data reported as 'Analysis' records in the ENA. Reports of abundance data estimated by short-read metagenome mapping should include information about the nucleotide identity and coverage thresholds used, with corresponding estimates of false-positive and false-negative rates either computed de novo or extracted from the literature (for example, from refs. 47, 81; Supplementary Note 4). All INSDC accession codes must be cited in publications. For ICTV classification, only coding-complete genomes (complete high-quality and finished draft UViGs) are currently considered82.
Conclusions
MIUViG standards and best practices for UViG analysis are the virus-specific counterparts to MISAG and MIMAG12. Virus genomics and metagenomics are rapidly expanding and improving as sequencing technologies emerge and mature. At the same time, the development of genome-based virus taxonomy methods as well as unified, comprehensive, and annotated reference databases of virus genomes and/or proteins continues apace. Community adoption of these standards, including through ongoing collaborations with other virus committees (ICTV) and data centers (DDBJ, EMBL-EBI and NCBI), will provide a framework for a systematic exploration of viral genome sequence space and enable the research community to better utilize and report UViGs.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Breitbart, M., Bonnain, C., Malki, K. & Sawaya, N.A. Phage puppet masters of the marine microbial realm. Nat. Microbiol. 3, 754–766 (2018).
Youle, M., Haynes, M. & Rohwer, F. in Viruses: Essential Agents of Life (ed. Witzany, G.) 61–81 (Springer Netherlands, 2012).
Brum, J.R. & Sullivan, M.B. Rising to the challenge: accelerated pace of discovery transforms marine virology. Nat. Rev. Microbiol. 13, 147–159 (2015).
Páez-Espino, D. et al. Uncovering Earth's virome. Nature 536, 425–430 (2016).
Shi, M. et al. Redefining the invertebrate RNA virosphere. Nature 540, 539–543 (2016).
Dayaram, A. et al. Diverse circular replication-associated protein encoding viruses circulating in invertebrates within a lake ecosystem. Infect. Genet. Evol. 39, 304–316 (2016).
Roux, S. et al. Ecogenomics and potential biogeochemical impacts of globally abundant ocean viruses. Nature 537, 689–693 (2016).
Arkhipova, K. et al. Temporal dynamics of uncultured viruses: a new dimension in viral diversity. ISME J. 12, 199–211 (2018).
Wilson, W.H. et al. Genomic exploration of individual giant ocean viruses. ISME J. 11, 1736–1745 (2017).
Brister, J.R., Ako-Adjei, D., Bao, Y. & Blinkova, O. NCBI viral genomes resource. Nucleic Acids Res. 43, D571–D577 (2015).
Páez-Espino, D. et al. IMG/VR: a database of cultured and uncultured DNA viruses and retroviruses. Nucleic Acids Res. 45, D457–D465 (2017).
Bowers, R.M. et al. Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of bacteria and archaea. Nat. Biotechnol. 35, 725–731 (2017).
Field, D. et al. The minimum information about a genome sequence (MIGS) specification. Nat. Biotechnol. 26, 541–547 (2008).
Yilmaz, P. et al. Minimum information about a marker gene sequence (MIMARKS) and minimum information about any (x) sequence (MIxS) specifications. Nat. Biotechnol. 29, 415–420 (2011).
Simmonds, P. et al. Consensus statement: virus taxonomy in the age of metagenomics. Nat. Rev. Microbiol. 15, 161–168 (2017).
Ladner, J.T. et al. Standards for sequencing viral genomes in the era of high-throughput sequencing. MBio 5, e01360–e14 (2014).
Thurber, R.V., Haynes, M., Breitbart, M., Wegley, L. & Rohwer, F. Laboratory procedures to generate viral metagenomes. Nat. Protoc. 4, 470–483 (2009).
Mokili, J.L., Rohwer, F. & Dutilh, B.E. Metagenomics and future perspectives in virus discovery. Curr. Opin. Virol. 2, 63–77 (2012).
Duhaime, M.B., Deng, L., Poulos, B.T. & Sullivan, M.B. Towards quantitative metagenomics of wild viruses and other ultra-low concentration DNA samples: a rigorous assessment and optimization of the linker amplification method. Environ. Microbiol. 14, 2526–2537 (2012).
Wylie, T.N., Wylie, K.M., Herter, B.N. & Storch, G.A. Enhanced virome sequencing using targeted sequence capture. Genome Res. 25, 1910–1920 (2015).
Allen, L.Z. et al. Single virus genomics: a new tool for virus discovery. PLoS One 6, e17722 (2011).
Martinez-Hernandez, F. et al. Single-virus genomics reveals hidden cosmopolitan and abundant viruses. Nat. Commun. 8, 15892 (2017).
Stepanauskas, R. et al. Improved genome recovery and integrated cell-size analyses of individual uncultured microbial cells and viral particles. Nat. Commun. 8, 84 (2017).
Houldcroft, C.J., Beale, M.A. & Breuer, J. Clinical and biological insights from viral genome sequencing. Nat. Rev. Microbiol. 15, 183–192 (2017).
Hingamp, P. et al. Exploring nucleo-cytoplasmic large DNA viruses in Tara Oceans microbial metagenomes. ISME J. 7, 1678–1695 (2013).
Roux, S., Hallam, S.J., Woyke, T. & Sullivan, M.B. Viral dark matter and virus-host interactions resolved from publicly available microbial genomes. Elife 4, e08490 (2015).
Kang, H.S. et al. Prophage genomics reveals patterns in phage genome organization and replication. Preprint at bioRxiv https://www.biorxiv.org/content/early/2017/03/07/114819 (2017).
Casjens, S. Prophages and bacterial genomics: what have we learned so far? Mol. Microbiol. 49, 277–300 (2003).
López-Pérez, M., Haro-Moreno, J.M., Gonzalez-Serrano, R., Parras-Moltó, M. & Rodriguez-Valera, F. Genome diversity of marine phages recovered from Mediterranean metagenomes: size matters. PLoS Genet. 13, e1007018 (2017).
Roux, S., Krupovic, M., Debroas, D., Forterre, P. & Enault, F. Assessment of viral community functional potential from viral metagenomes may be hampered by contamination with cellular sequences. Open Biol. 3, 130160 (2013).
Luef, B. et al. Diverse uncultivated ultra-small bacterial cells in groundwater. Nat. Commun. 6, 6372 (2015).
Frost, L.S., Leplae, R., Summers, A.O. & Toussaint, A. Mobile genetic elements: the agents of open source evolution. Nat. Rev. Microbiol. 3, 722–732 (2005).
Lang, A.S. & Beatty, J.T. Importance of widespread gene transfer agent genes in alpha-proteobacteria. Trends Microbiol. 15, 54–62 (2007).
Biller, S.J. et al. Membrane vesicles in sea water: heterogeneous DNA content and implications for viral abundance estimates. ISME J. 11, 394–404 (2017).
Arndt, D. et al. PHASTER: a better, faster version of the PHAST phage search tool. Nucleic Acids Res. 44, W16–W21 (2016).
Roux, S., Enault, F., Hurwitz, B.L. & Sullivan, M.B. VirSorter: mining viral signal from microbial genomic data. PeerJ 3, e985 (2015).
Amgarten, D., Braga, L.P.P., da Silva, A.M. & Setubal, J.C. MARVEL, a tool for prediction of bacteriophage sequences in metagenomic bins. Front. Genet. 9, 304 (2018).
Ren, J., Ahlgren, N.A., Lu, Y.Y., Fuhrman, J.A. & Sun, F. VirFinder: a novel k-mer based tool for identifying viral sequences from assembled metagenomic data. Microbiome 5, 69 (2017).
Zhao, G. et al. VirusSeeker, a computational pipeline for virus discovery and virome composition analysis. Virology 503, 21–30 (2017).
Páez-Espino, D., Pavlopoulos, G.A., Ivanova, N.N. & Kyrpides, N.C. Nontargeted virus sequence discovery pipeline and virus clustering for metagenomic data. Nat. Protoc. 12, 1673–1682 (2017).
Moniruzzaman, M. et al. Diversity and dynamics of algal Megaviridae members during a harmful brown tide caused by the pelagophyte, Aureococcus anophagefferens. FEMS Microbiol. Ecol. 92, fiw058 (2016).
Sakowski, E.G. et al. Ribonucleotide reductases reveal novel viral diversity and predict biological and ecological features of unknown marine viruses. Proc. Natl. Acad. Sci. USA 111, 15786–15791 (2014).
Marine, R.L., Nasko, D.J., Wray, J., Polson, S.W. & Wommack, K.E. Novel chaperonins are prevalent in the virioplankton and demonstrate links to viral biology and ecology. ISME J. 11, 2479–2491 (2017).
Schmidt, H.F., Sakowski, E.G., Williamson, S.J., Polson, S.W. & Wommack, K.E. Shotgun metagenomics indicates novel family A DNA polymerases predominate within marine virioplankton. ISME J. 8, 103–114 (2014).
Culley, A.I., Lang, A.S. & Suttle, C.A. Metagenomic analysis of coastal RNA virus communities. Science 312, 1795–1798 (2006).
Needham, D.M., Sachdeva, R. & Fuhrman, J.A. Ecological dynamics and co-occurrence among marine phytoplankton, bacteria and myoviruses shows microdiversity matters. ISME J. 11, 1614–1629 (2017).
Roux, S., Emerson, J.B., Eloe-Fadrosh, E.A. & Sullivan, M.B. Benchmarking viromics: an in silico evaluation of metagenome-enabled estimates of viral community composition and diversity. PeerJ 5, e3817 (2017).
Lorenzi, H.A. et al. The viral metagenome annotation pipeline (VMGAP): an automated tool for the functional annotation of viral metagenomic shotgun sequencing data. Stand. Genomic Sci. 4, 418–429 (2011).
McNair, K. et al. Phage genome annotation using the RAST pipeline. Methods Mol. Biol. 1681, 231–238 (2018).
Brister, J.R. et al. Towards viral genome annotation standards, report from the 2010 NCBI Annotation Workshop. Viruses 2, 2258–2268 (2010).
Eddy, S.R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Altschul, S.F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Söding, J. Protein homology detection by HMM-HMM comparison. Bioinformatics 21, 951–960 (2005).
Reyes, A.P., Alves, J.M., Durham, A.M. & Gruber, A. Use of profile hidden Markov models in viral discovery: current insights. Adv. Genomics Genet. 7, 29–45 (2017).
Gregory, A.C. et al. Genomic differentiation among wild cyanophages despite widespread horizontal gene transfer. BMC Genomics 17, 930 (2016).
Duhaime, M.B. et al. Comparative omics and trait analyses of marine Pseudoalteromonas phages advance the phage OTU concept. Front. Microbiol. 8, 1241 (2017).
Mavrich, T.N. & Hatfull, G.F. Bacteriophage evolution differs by host, lifestyle and genome. Nat. Microbiol. 2, 17112 (2017).
Aiewsakun, P. & Simmonds, P. The genomic underpinnings of eukaryotic virus taxonomy: creating a sequence-based framework for family-level virus classification. Microbiome 6, 38 (2018).
Bolduc, B. et al. vConTACT: an iVirus tool to classify double-stranded DNA viruses that infect Archaea and Bacteria. PeerJ 5, e3243 (2017).
Mizuno, C.M., Rodriguez-Valera, F., Kimes, N.E. & Ghai, R. Expanding the marine virosphere using metagenomics. PLoS Genet. 9, e1003987 (2013).
Bào, Y. et al. Implementation of objective PASC-derived taxon demarcation criteria for official classification of filoviruses. Viruses 9, E106 (2017).
Varsani, A. & Krupovic, M. Sequence-based taxonomic framework for the classification of uncultured single-stranded DNA viruses of the family Genomoviridae. Virus Evol. 3, vew037 (2017).
Rohwer, F. & Edwards, R. The phage proteomic tree: a genome-based taxonomy for phage. J. Bacteriol. 184, 4529–4535 (2002).
Lavigne, R. et al. Classification of Myoviridae bacteriophages using protein sequence similarity. BMC Microbiol. 9, 224 (2009).
Nishimura, Y. et al. ViPTree: the viral proteomic tree server. Bioinformatics 33, 2379–2380 (2017).
Meier-Kolthoff, J.P. & Göker, M. VICTOR: genome-based phylogeny and classification of prokaryotic viruses. Bioinformatics 33, 3396–3404 (2017).
Baltimore, D. Expression of animal virus genomes. Bacteriol. Rev. 35, 235–241 (1971).
Edwards, R.A., McNair, K., Faust, K., Raes, J. & Dutilh, B.E. Computational approaches to predict bacteriophage-host relationships. FEMS Microbiol. Rev. 40, 258–272 (2016).
Mihara, T. et al. Linking virus genomes with host taxonomy. Viruses 8, 66 (2016).
Villarroel, J. et al. HostPhinder: a phage host prediction tool. Viruses 8, 116 (2016).
Garcia-Heredia, I. et al. Reconstructing viral genomes from the environment using fosmid clones: the case of haloviruses. PLoS One 7, e33802 (2012).
Roux, S. et al. Ecology and evolution of viruses infecting uncultivated SUP05 bacteria as revealed by single-cell- and meta-genomics. Elife 3, e03125 (2014).
Labonté, J.M. et al. Single-cell genomics-based analysis of virus-host interactions in marine surface bacterioplankton. ISME J. 9, 2386–2399 (2015).
Galiez, C., Siebert, M., Enault, F., Vincent, J. & Söding, J. WIsH: who is the host? Predicting prokaryotic hosts from metagenomic phage contigs. Bioinformatics 33, 3113–3114 (2017).
Ahlgren, N.A., Ren, J., Lu, Y.Y., Fuhrman, J.A. & Sun, F. Alignment-free d2* oligonucleotide frequency dissimilarity measure improves prediction of hosts from metagenomically-derived viral sequences. Nucleic Acids Res. 45, 39–53 (2017).
Reyes, A., Wu, M., McNulty, N.P., Rohwer, F.L. & Gordon, J.I. Gnotobiotic mouse model of phage-bacterial host dynamics in the human gut. Proc. Natl. Acad. Sci. USA 110, 20236–20241 (2013).
Lima-Mendez, G. et al. Determinants of community structure in the global plankton interactome. Science 348, 1262073 (2015).
Dutilh, B.E. et al. A highly abundant bacteriophage discovered in the unknown sequences of human faecal metagenomes. Nat. Commun. 5, 4498 (2014).
Coenen, A.R. & Weitz, J.S. Limitations of correlation-based inference in complex virus-microbe communities. mSystems 3, e00084–18 (2018).
Gao, N.L. et al. MVP: a microbe-phage interaction database. Nucleic Acids Res. 46, D700–D707 (2018).
Aziz, R.K., Dwivedi, B., Akhter, S., Breitbart, M. & Edwards, R.A. Multidimensional metrics for estimating phage abundance, distribution, gene density, and sequence coverage in metagenomes. Front. Microbiol. 6, 381 (2015).
Adams, M.J. et al. 50 years of the International Committee on Taxonomy of Viruses: progress and prospects. Arch. Virol. 162, 1441–1446 (2017).
Reyes, A. et al. Gut DNA viromes of Malawian twins discordant for severe acute malnutrition. Proc. Natl. Acad. Sci. USA 112, 11941–11946 (2015).
Shendure, J. et al. Accurate multiplex polony sequencing of an evolved bacterial genome. Science 309, 1728–1732 (2005).
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005).
Angly, F.E. et al. The marine viromes of four oceanic regions. PLoS Biol. 4, e368 (2006).
Lima-Mendez, G., Van Helden, J., Toussaint, A. & Leplae, R. Prophinder: a computational tool for prophage prediction in prokaryotic genomes. Bioinformatics 24, 863–865 (2008).
Reyes, A. et al. Viruses in the faecal microbiota of monozygotic twins and their mothers. Nature 466, 334–338 (2010).
Yoon, H.S. et al. Single-cell genomics reveals organismal interactions in uncultivated marine protists. Science 332, 714–717 (2011).
Andrewes, C.H. The classification of viruses. J. Gen. Microbiol. 12, 358–361 (1955).
Lwoff, A., Horne, R. & Tournier, P. A system of viruses. Cold Spring Harb. Symp. Quant. Biol. 27, 51–55 (1962).
Lwoff, A. The new provisional committee on nomenclature of viruses. Int. Bull. Bacteriol. Nomencl. Taxon. 14, 53–56 (1964).
King, A.M.Q. et al. Changes to taxonomy and the International Code of Virus Classification and Nomenclature ratified by the International Committee on Taxonomy of Viruses. Arch. Virol. 163, 2601–2631 (2018).
Acknowledgements
This work was supported by the Laboratory Directed Research and Development Program of Lawrence Berkeley National Laboratory under US Department of Energy Contract No. DE-AC02-05CH11231 for S.R.; the Netherlands Organization for Scientific Research (NWO) Vidi grant 864.14.004 for B.E.D.; the Intramural Research Program of the National Library of Medicine, National Institutes of Health for E.V.K., I.K.M., J.R.B. and N.Y.; the Virus-X project (EU Horizon 2020, No. 685778) for F.E. and M.K.; Battelle Memorial Institute's prime contract with the US National Institute of Allergy and Infectious Diseases (NIAID) under Contract No. HHSN272200700016I for J.H.K.; the GOA grant “Bacteriophage Biosystems” from KU Leuven for R.L.; the European Molecular Biology Laboratory for C.A. and G.R.C.; Cairo University Grant 2016-57 for R.K.A.; National Science Foundation award 1456778, National Institutes of Health awards R01 AI132581 and R21 HD086833, and The Vanderbilt Microbiome Initiative award for S.R.B.; National Science Foundation awards DEB-1239976 for M.B. and K.R. and DEB-1555854 for M.B.; the NSF Early Career award DEB-1555854 and NSF Dimensions of Biodiversity #1342701 for K.C.W. and R.A.D.; the Agence Nationale de la Recherche JCJC grant ANR-13-JSV6-0004 and Investissements d'Avenir Méditerranée Infection 10-IAHU-03 for C.D.; the Gordon and Betty Moore Foundation Marine Microbiology Initiative No. 3779 and the Simons Foundation for J.A.F.; the French government “Investissements d'Avenir” program OCEANOMICS ANR-11-BTBR-0008 and European FEDER Fund 1166-39417 for P. Hingamp; Australian Research Council Laureate Fellowship FL150100038 to P. Hugenholtz the National Science Foundation award 1801367 and C-DEBI Research Grant for J.M.L.; the Gordon and Betty Moore Foundation grant 5334 and Ministry of Economy and Competitivity refs. CGL2013-40564-R and SAF2013-49267-EXP for M.M.-G.; the Grant-in-Aid for Scientific Research on Innovative Areas from the Ministry of Education, Culture, Science, Sports, and Technology (MEXT) of Japan No. 16H06429, 16K21723, and 16H06437 for H.O. and T.Y.; National Science Foundation award DBI-1661357 to C.P.; the Ministry of Economy and Competitivity ref CGL2016-76273-P (cofunded with FEDER funds) for F.R.-V.; the Gordon and Betty Moore Foundation awards 3305 and 3790 and NSF Biological Oceanography OCE 1536989 for M.B.S.; the ETH Zurich and Helmut Horten Foundation and the Novartis Foundation for Medical-Biological Research (17B077) for S.S.; a BIOS-SCOPE award from Simons Foundation International and NERC award NE/P008534/1 to B.T.; NSF Biological Oceanography Grant 1635913 for R.V.T.; the Australian Research Council Future Fellowship FT120100480 for N.S.W.; a Gilead Sciences Cystic Fibrosis Research Scholarship for K.L.W.; Gordon and Better Moore Foundation Grant 4971 for S.W.W.; the NSF EPSCoR grant 1736030 for K.E.W.; the National Science Foundation award DEB-4W4596 and National Institutes of Health award R01 GM117361 for M.J.Y.; the Gordon and Betty Moore Foundation No. 7000 and the National Oceanic and Atmospheric Administration (NOAA) under award NA15OAR4320071 for L.Z.A. DDBJ is supported by ROIS and MEXT. The work conducted by the US Department of Energy Joint Genome Institute is supported by the Office of Science of the US Department of Energy under contract no. DE-AC02-05CH11231. The views and conclusions contained in this document are those of the authors and should not be interpreted as necessarily representing the official policies, either expressed or implied, of the US Department of Health and Human Services or of the institutions and companies affiliated with the authors. B.E.D., A.K., M.K., J.H.K., R.L. and A.V. are members of the ICTV Executive Committee, but the views and opinions expressed are those of the authors and not those of the ICTV.
Author information
Authors and Affiliations
Contributions
All authors participated in writing the manuscript and provided critical feedback. S.R. performed the analyses for the supplementary notes and figures.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
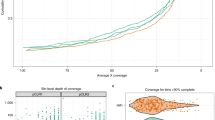
Supplementary Figure 1 Comparison of UViG recovery from microbial (“M”) and viral (“V”) metagenomes originating from the same Tara Oceans samples.
Top panel represents the number of distinct virus contigs ≥ 10kb identified in each dataset. The bottom panel depicts the ratio of “shared”, i.e., detected in both viral and microbial fraction of the sample, and “unique”, i.e., detected only in one fraction, contigs in each microbial and viral fraction. Datasets were originally analyzed in refs. 1,2. SRF: surface, DCM: deep chlorophyll maximum.
Supplementary Figure 2 Genome size variation for different types of viruses and different taxonomic levels.
Genome length of virus genomes from NCBI RefSeq were compared at different taxonomic ranks and are presented separately for four main types of viruses (dsDNA, ssDNA, RNA and reverse-transcribing RNA, viroids and satellites). Genome length variation was calculated as a coefficient of variation, i.e. standard deviation of genome length in the group divided by average genome length in the grouop (for groups with >1 genome). Underlying data are available in Supplementary Table 5. Boxplots lower and upper hinges correspond to the first and third quartiles (the 25th and 75th percentiles), while whisker extend from the nearest hinge to the smallest/largest value no further than 1.5 * IQR from the hinge (where IQR is the inter-quartile range, or distance between the first and third quartiles). dsDNA: double-stranded DNA; ssDNA: single-stranded DNA.
Supplementary Figure 3 Pairwise average nucleotide identity (ANI) and alignment fraction (AF) for NCBI Viral RefSeq genomes and IMG/VR.
Only genome pairs with ANI >60% and AF >20% were considered. ANI and AF were binned in 1% intervals, and are represented here as a heatmap (i.e. cell coloring represents the number of pairwise comparisons at the corresponding ANI and AF intervals). On the top right corner (i.e., AF and ANI close to 100%), three main groups of genome pairs are delineated with black dashed circles, and the proposed standard cutoff is highlighted in dark red. Note that for this clustering, the cutoff was applied as follows: pairs of genomes with ≥ 85% AF were first selected, and whole genome (wg) ANI was then calculated by multiplying the observed ANI by the observed AF. This wgANI was then compared to the corresponding whole genome ANI cutoff (i.e. 95% ANI * 85% AF = 80.75% wgANI). This allows for hits with ≤ 95% ANI but ≥ 85 % AF to be considered as well, i.e. a pair of genomes with 90% ANI on 100% AF would be considered as “passing” the cutoff. Examples of genome comparisons for each group are presented in Supplementary Fig. 4.
Supplementary Figure 4 Examples of pairwise genome comparisons from the three groups of genome pairs highlighted in Supplementary Figure 3.
For each example, nucleotide similarity (blastn) and amino acid similarity (tblastx) are displayed, alongside the ANI, AF, and wgANI (i.e. ANI over the whole length of the shorter genome). AF, alignment fraction; ANI, average nucleotide identity; wgANI, whole-genome average nucleotide identity.
Supplementary Figure 5 Estimation of whole genome ANI from fragmented genomes.
To evaluate the impact of genome fragmentation on whole-genome average nucleotide identity (wgANI) estimation, pairs of genomes from NCBI RefSeq with wgANI ≥ 70% and ≥ 20kb were selected, random fragments were generated (from 1 to 45kb) from one of the two genomes, and then compared to the other complete genome. The resulting estimated wgANI between the fragment and complete genome was then compared with the original values estimated from the two complete genomes (y-axis). Boxplots lower and upper hinges correspond to the first and third quartiles (the 25th and 75th percentiles), while whisker extend from the nearest hinge to the smallest/largest value no further than 1.5 * IQR from the hinge (where IQR is the inter-quartile range, or distance between the first and third quartiles).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 (PDF 1122 kb)
Supplementary Notes
Supplementary Notes 1–4 (PDF 196 kb)
Supplementary Table 1
List of mandatory and optional metadata for UViGs (XLSX 9 kb)
Supplementary Table 2
List of metadata from previous standards relevant for UViGs21 (XLSX 17 kb)
Supplementary Table 3
Comparison between UViGs categories and the quality categories proposed for small DNA/RNA virus whole-genome sequencing for epidemiology and surveillance by Ladner et al.22 (XLSX 5 kb)
Supplementary Table 4
List and characteristics of tools used to identify virus sequences in mixed datasets published or updated since 201223–31 (XLSX 6 kb)
Supplementary Table 5
Variation in genome length for virus families and genera with two or more genomes, from NCBI RefSeq v83. (XLSX 25 kb)
Supplementary Table 6
List of potential marker genes for virus orders, families or genera, based on the VOGdb v83 (http://vogdb.org/) (XLSX 85 kb)
Supplementary Table 7
List of UViGs from the GOV dataset4 considered as high-quality drafts or finished genomes (XLSX 38 kb)
Supplementary Table 8
List of databases providing collections of HMM profiles for virus protein families32–35 (XLSX 6 kb)
Supplementary Table 9
Current species demarcation criteria from ICTV ninth and tenth reports. (XLSX 46 kb)
Supplementary Table 10
Approaches available for in silico host prediction18,37–42 (XLSX 6 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International (CC BY 4.0) licence. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons licence, users will need to obtain permission from the licence holder to reproduce the material. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Roux, S., Adriaenssens, E., Dutilh, B. et al. Minimum Information about an Uncultivated Virus Genome (MIUViG). Nat Biotechnol 37, 29–37 (2019). https://doi.org/10.1038/nbt.4306
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.4306
This article is cited by
-
Benchmarking bioinformatic virus identification tools using real-world metagenomic data across biomes
Genome Biology (2024)
-
Hot springs viruses at Yellowstone National Park have ancient origins and are adapted to thermophilic hosts
Communications Biology (2024)
-
A metagenomic catalog of the early-life human gut virome
Nature Communications (2024)
-
Discovery and description of novel phage genomes from urban microbiomes sampled by the MetaSUB consortium
Scientific Reports (2024)
-
Biogeographic patterns and drivers of soil viromes
Nature Ecology & Evolution (2024)