Abstract
The promiscuous nature of T-cell receptors (TCRs) allows T cells to recognize a large variety of pathogens, but makes it challenging to understand and control T-cell recognition1. Existing technologies provide limited information about the key requirements for T-cell recognition and the ability of TCRs to cross-recognize structurally related elements2,3. Here we present a 'one-pot' strategy for determining the interactions that govern TCR recognition of peptide–major histocompatibility complex (pMHC). We measured the relative affinities of TCRs to libraries of barcoded peptide–MHC variants and applied this knowledge to understand the recognition motif, here termed the TCR fingerprint. The TCR fingerprints of 16 different TCRs were identified and used to predict and validate cross-recognized peptides from the human proteome. The identified fingerprints differed among TCRs recognizing the same epitope, demonstrating the value of this strategy for understanding T-cell interactions and assessing potential cross-recognition before selection of TCRs for clinical development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Sewell, A.K. Why must T cells be cross-reactive? Nat. Rev. Immunol. 12, 669–677 (2012).
Obenaus, M. et al. Identification of human T-cell receptors with optimal affinity to cancer antigens using antigen-negative humanized mice. Nat. Biotechnol. 33, 402–407 (2015).
Cameron, B.J. et al. Identification of a Titin-derived HLA-A1-presented peptide as a cross-reactive target for engineered MAGE A3-directed T cells. Sci. Transl. Med. 5, 197ra103 (2013).
Davis, M.M. & Bjorkman, P.J. T-cell antigen receptor genes and T-cell recognition. Nature 334, 395–402 (1988).
Mason, D. A very high level of crossreactivity is an essential feature of the T-cell receptor. Immunol. Today 19, 395–404 (1998).
Arstila, T.P. et al. A direct estimate of the human αβ T cell receptor diversity. Science 286, 958–961 (1999).
Robins, H.S. et al. Comprehensive assessment of T-cell receptor β-chain diversity in αβ T cells. Blood 114, 4099–4107 (2009).
Cornberg, M. & Wedemeyer, H. Hepatitis C virus infection from the perspective of heterologous immunity. Curr. Opin. Virol. 16, 41–48 (2016).
Wooldridge, L. et al. A single autoimmune T cell receptor recognizes more than a million different peptides. J. Biol. Chem. 287, 1168–1177 (2012).
Birnbaum, M.E. et al. Deconstructing the peptide-MHC specificity of T cell recognition. Cell 157, 1073–1087 (2014).
Adams, J.J. et al. Structural interplay between germline interactions and adaptive recognition determines the bandwidth of TCR-peptide-MHC cross-reactivity. Nat. Immunol. 17, 87–94 (2016).
Gee, M.H. et al. Antigen identification for orphan T cell receptors expressed on tumor-infiltrating lymphocytes. Cell 172, 549–563.e16 (2018).
Sibener, L.V. et al. Isolation of a structural mechanism for uncoupling T cell receptor signaling from peptide-MHC binding. Cell 174, 672–687.e27 (2018).
Wooldridge, L. et al. CD8 controls T cell cross-reactivity. J. Immunol. 185, 4625–4632 (2010).
Schaubert, K.L. et al. Generation of robust CD8+ T-cell responses against subdominant epitopes in conserved regions of HIV-1 by repertoire mining with mimotopes. Eur. J. Immunol. 40, 1950–1962 (2010).
Bentzen, A.K. et al. Large-scale detection of antigen-specific T cells using peptide-MHC-I multimers labeled with DNA barcodes. Nat. Biotechnol. 34, 1037–1045 (2016).
Bentzen, A.K. & Hadrup, S.R. Evolution of MHC-based technologies used for detection of antigen-responsive T cells. Cancer Immunol. Immunother. 66, 657–666 (2017).
Lyngaa, R. et al. T-cell responses to oncogenic Merkel cell polyomavirus proteins distinguish patients with Merkel cell carcinoma from healthy donors. Clin. Cancer Res. 20, 1768–1778 (2014).
Rodenko, B. et al. Generation of peptide-MHC class I complexes through UV-mediated ligand exchange. Nat. Protoc. 1, 1120–1132 (2006).
Toebes, M. et al. Design and use of conditional MHC class I ligands. Nat. Med. 12, 246–251 (2006).
Enouz, S., Carrié, L., Merkler, D., Bevan, M.J. & Zehn, D. Autoreactive T cells bypass negative selection and respond to self-antigen stimulation during infection. J. Exp. Med. 209, 1769–1779 (2012).
Miller, N.J. et al. Tumor-infiltrating Merkel cell polyomavirus-specific t cells are diverse and associated with improved patient survival. Cancer Immunol. Res. 5, 137–147 (2017).
Grant, C.E., Bailey, T.L. & Noble, W.S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Lee, P.P. et al. Characterization of circulating T cells specific for tumor-associated antigens in melanoma patients. Nat. Med. 5, 677–685 (1999).
Albert, L.J. & Inman, R.D. Molecular mimicry and autoimmunity. N. Engl. J. Med. 341, 2068–2074 (1999).
van den Berg, J.H. et al. Case report of a fatal serious adverse event upon administration of T cells transduced with a MART-1-specific T-cell receptor. Mol. Ther. 23, 1541–1550 (2015).
Fowler, D.M. & Fields, S. Deep mutational scanning: a new style of protein science. Nat. Methods 11, 801–807 (2014).
Harris, D.T. et al. Deep mutational scans as a guide to engineering high affinity t cell receptor interactions with peptide-bound major histocompatibility complex. J. Biol. Chem. 291, 24566–24578 (2016).
Linette, G.P. et al. Cardiovascular toxicity and titin cross-reactivity of affinity-enhanced T cells in myeloma and melanoma. Blood 122, 863–871 (2013).
Morgan, R.A. et al. Cancer regression and neurological toxicity following anti-MAGE-A3 TCR gene therapy. J. Immunother. 36, 133–151 (2013).
Xu, Q., Schlabach, M.R., Hannon, G.J. & Elledge, S.J. Design of 240,000 orthogonal 25mer DNA barcode probes. Proc. Natl. Acad. Sci. USA 106, 2289–2294 (2009).
Kivioja, T. et al. Counting absolute numbers of molecules using unique molecular identifiers. Nat. Methods 9, 72–74 (2011).
Chang, C.X.L. et al. Conditional ligands for Asian HLA variants facilitate the definition of CD8+ T-cell responses in acute and chronic viral diseases. Eur. J. Immunol. 43, 1109–1120 (2013).
Hadrup, S.R. et al. High-throughput T-cell epitope discovery through MHC peptide exchange. Methods Mol. Biol. 524, 383–405 (2009).
Frøsig, T.M. et al. Design and validation of conditional ligands for HLA-B*08:01, HLA-B*15:01, HLA-B*35:01, and HLA-B*44:05. Cytometry A 87, 967–975 (2015).
Hadrup, S.R. et al. Parallel detection of antigen-specific T-cell responses by multidimensional encoding of MHC multimers. Nat. Methods 6, 520–526 (2009).
Andersen, R.S. et al. Parallel detection of antigen-specific T cell responses by combinatorial encoding of MHC multimers. Nat. Protoc. 7, 891–902 (2012).
Linnemann, C. et al. High-throughput identification of antigen-specific TCRs by TCR gene capture. Nat. Med. 19, 1534–1541 (2013).
Bolotin, D.A. et al. Next generation sequencing for TCR repertoire profiling: platform-specific features and correction algorithms. Eur. J. Immunol. 42, 3073–3083 (2012).
Han, A., Glanville, J., Hansmann, L. & Davis, M.M. Linking T-cell receptor sequence to functional phenotype at the single-cell level. Nat. Biotechnol. 32, 684–692 (2014).
Kühlcke, K. et al. Retroviral transduction of T lymphocytes for suicide gene therapy in allogeneic stem cell transplantation. Bone Marrow Transplant. 25 (Suppl. 2), S96–S98 (2000).
Nielsen, M. & Andreatta, M. NetMHCpan-3.0; improved prediction of binding to MHC class I molecules integrating information from multiple receptor and peptide length datasets. Genome Med. 8, 33 (2016).
Fiser, A. & Sali, A. ModLoop: automated modeling of loops in protein structures. Bioinformatics 19, 2500–2501 (2003).
Stein, A. & Kortemme, T. Improvements to robotics-inspired conformational sampling in Rosetta. PLoS One 8, e63090 (2013).
Kaufmann, K.W., Lemmon, G.H., Deluca, S.L., Sheehan, J.H. & Meiler, J. Practically useful: what the Rosetta protein modeling suite can do for you. Biochemistry 49, 2987–2998 (2010).
Schymkowitz, J. et al. The FoldX web server: an online force field. Nucleic Acids Res. 33, W382–W388 (2005).
Wagih, O. ggseqlogo: a versatile R package for drawing sequence logos. Bioinformatics 33, 3645–3647 (2017).
Thomsen, M.C.F. & Nielsen, M. Seq2Logo: a method for construction and visualization of amino acid binding motifs and sequence profiles including sequence weighting, pseudo counts and two-sided representation of amino acid enrichment and depletion. Nucleic Acids Res. 40, W281–W287 (2012).
Jurtz, V. et al. NetMHCpan-4.0: improved peptide-MHC class I interaction predictions integrating eluted ligand and peptide binding affinity data. J. Immunol. 199, 3360–3368 (2017).
Robinson, M.D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
Acknowledgements
We would like to thank all Merkel cell carcinoma patients and healthy donors contributing material to this study; T. Joeris, W. Agace and K. Lahl (Lund University, Lund, Sweden) for sharing of splenocytes from OT-1 and OT-3 transgenic mice; B. Rotbøl and T. Tamhane for excellent technical assistance; and S.R.H.'s group for scientific discussions. This research was funded in part through the European Research Council, StG 677268 NextDART, Lundbeck Foundation Fellowship R190-2014-4178 (to S.R.H.), the Independent Research Fund Denmark (DFF 4004-00422 and DFF 7014-00055, to S.R.H. and M.N.), NIH NCI Cancer Center Support Grant P30 CA015704, K24-CA-139052 and R01 CA176841, Kelsey Dickson Team Science Courage Research Team Award, Prostate Cancer Foundation Award 15CHAS04, and the UW MCC Patient Gift Fund (to P.N.).
Author information
Authors and Affiliations
Contributions
A.K.B. and L.S. conceived the idea, designed and performed experiments, analyzed data, made figures and wrote the manuscript; K.K.J., A.M.M. and L.E.J. conceived the bioinformatic processing, analyzed data, made figures and revised the manuscript; N.J.M. and C.D.C. identified and established T cell clones; R.L. identified and characterized TCRs and TCR-transduced T cells; D.M.K., J.C.B. and P.N. provided patient material for TCR identification and supervised the studies; C.L. and T.N.M.S. sequenced and expressed TCRs and revised the manuscript; P.M. and M.N. conceived the bioinformatic processing, supervised data analyses and revised the manuscript; S.R.H. conceived the idea, supervised the study, designed experiments, analyzed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
A.K.B. and S.R.H. are co-inventors on a patent covering the use of DNA barcode-labeled MHC multimers (WO2015185067 and WO2015188839). N.J.M., C.D.C., D.M.K. and P.N. are co-inventors on a patent application filed by their employer, University of Washington, concerning the HLA-A*0201-restricted TCR sequences.
Integrated supplementary information
Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints
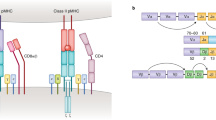
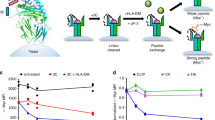
(a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide is substituted with all naturally occurring amino acids. The resultant peptide variants are assembled with MHC molecules, and individually multiplexed on a PE-labeled dextran backbone together with individual DNA barcodes. Thus, a given DNA barcode forms a tag for the corresponding pMHC variant. T cells recognizing the original pMHC are incubated with all MHC multimer variants simultaneously. The MHC multimer binding cells are isolated based on the shared PE-label and their associated DNA barcodes are amplified and sequenced. The hierarchy of pMHC interactions is revealed from the relative number of DNA-barcode reads specifically associated with a given MHC multimer. DNA barcode reads are typically transformed to log2FC values (data in the example corresponds to results from Fig. 1a). (b) A clonal TCR will bind a pool of pMHC variants in a hierarchical manner that is governed by the avidity of the given pMHC and the TCR. (c) The hierarchy of TCR-pMHC interactions is used to determine the recognition pattern of that given TCR, visualized as a Shannon plot (here determined ‘TCR fingerprint’).
Supplementary Figure 2 Feasibility of using DNA barcode-labeled multimers to determine the binding hierarchy of multiple pMHCs in a ‘one-pot’ screen
Bar plots of the relative log2FC values from the DNA barcode-based analyses of PBMCs transduced with (a-c) a HLA-B*0702APN engaging TCR or (d) a HLA-A*2402EWW engaging TCR. Each bar represents a pMHC variant with an amino acid substitution at the given peptide position. Bars of the same color indicate peptides that were included in a parallel screening. The dotted line at zero indicates the normalized log2FC of the original pMHC. (a-b) The MHC multimer libraries applied in the analyses included all possible 9- and 10-mer variants of APNCYGNIPL generated from substituting with alanine. (a) Shows a series of three analyses where # indicates a peptide variant that was excluded in a given screening. Data is representative of triplicate analyses. (b) Bar plot from a series of analyses using the same MHC multimer library of APNCYGNIPL variants. In these analyses a varying amount of DNA barcode-labeled MHC multimers were used, i.e. a titration from 23 nM to 0.92 nM in respect to each pMHC (corresponding to 100%, 20% or 4% of the amount originally applied, n=1 for each variable). (c) Bar plot from a series of analyses where a range of 10-mer peptides produced from substituting with alanine, lysine or glutamic acid were applied to generate three different libraries of DNA barcode-labeled MHC multimers that were used to stain the HLA-B*0702APN engaging TCR. (d) The MHC multimer libraries applied in these analyses included all possible 9- and 10-mer variants of EWWRSGGFSF generated from substituting with alanine. The series of analyses are equivalent to that shown in (b) only conducted on a different TCR and at lower MHC multimer concentrations, i.e. a titration from 0.92 nM to 0.036 nM in respect to each pMHC (corresponding to 4%, 0.8% or 0.16% of the amount originally applied, n=1 for each variable).
Supplementary Figure 3 Replicate screening of the HLA-B*0702APN-engaging TCR showing read counts and data from an irrelevant sample
Results obtained from the DNA barcode-based analysis of T cells transduced with a TCR recognizing the HLA-B*0702 restricted peptide, APNCYGNIPL (replicate of the data shown in Fig. 1a-c). The analysis was performed with all possible variations of peptides created by single-position amino acid substitutions. The hierarchy of pMHC interactions expressed as (a) clonality reduced read counts or (b) log2FC relative to a triplicate baseline sample (see Supplementary Note 1). A healthy donor PBMC sample (BC-D62) was screened with the same MHC multimer panel in parallel. The plotted order of read counts or log2FC of each pMHC-associated DNA barcode is determined by the hierarchy obtained from screening the HLA-B*0702APN responsive TCR. (c-f) Heat map showing the clonality reduced read counts or log2FC of (c,d) the HLA-B*0702APN responsive TCR or (e,f) the healthy donor BC-D62, respectively. Each row represents a given amino acid and each column a position in the peptide sequence. The amino acids of the original peptide target are marked with black borders. c and e showing read counts, and d and f showing Log2FC are colored according to the same keys respectively. (g) The replicate recognition pattern of the HLA-B*0702APN interacting TCR, here visualized as a sequence logo based on the data from a-d.
Supplementary Figure 4 Replicate screening of the HLA-A*2402EWW-engaging TCR showing read counts and data from an irrelevant sample
Results obtained from the DNA barcode-based analysis of T cells transduced with a TCR recognizing the HLA-A*2402 restricted peptide, EWWRSGGFSF (replicate of the data shown in Fig. 1d-f). The analysis was performed with all possible variations of peptides created by single-position amino acid substitutions. The hierarchy of pMHC interactions expressed as (a) clonality reduced read counts or (b) log2FC relative to a triplicate baseline sample (see Supplementary Note). A healthy donor PBMC sample (BC-D42) was screened with the same MHC multimer panel in parallel. The plotted order of read counts or log2FC of each pMHC-associated DNA barcode is determined by the hierarchy obtained from screening the HLA-A*2402EWW responsive TCR. (c-f) Heat map showing the clonality reduced read counts or log2FC of (c,d) the HLA-A*2402EWW responsive TCR or (e,f) the healthy donor BC-D42, respectively. Each row represents a given amino acid and each column a position in the peptide sequence. The amino acids of the original peptide target are marked with black borders. c and e showing read counts, and d and f showing Log2FC are colored according to the same keys respectively. (g) The replicate recognition pattern of the HLA-A*2402EWW interacting TCR, here visualized as a sequence logo based on the data from a-d.
Supplementary Figure 5 The effect of peptide–MHC interaction on analyzing the recognition pattern of the HLA-B*0702APN-engaging TCR
(a) Sequence logo showing the amino acid preferences for peptide binding to HLA-B*0702 constructed with NetMHCpan-4.049 by finding the binding cores of the top 1% strongest predicted binders among 100,000 natural random 10-mer peptides and visualizing these using R with the Shannon method in Seq2Logo48. (b) Scatter plot of the experimentally determined peptide binding to HLA-B*0702 (MHC ELISA, Supplementary Note 3), given as the relative quantity of a pMHC variant (only alanine substitutions) after UV-exchange, compared to that of the original peptide, in correlation to the experimentally obtained recognition properties of the given pHLA-B*0702 interacting with the TCR. The MHC ELISA data is average of 5 experiments (n=2). Each dot in b represents one peptide-MHC variation. The coloring indicates the position of a given amino acid substitution and the asterisks indicates the original peptide. (c) Dot plots from staining the HLA-B*0702APN engaging TCR with fluorescently labeled MHC multimers carrying one of six variations of APNCYGNIPL and an irrelevant peptide (p*, the UV conditional peptide). 1-3 are all examples of peptides predicted as strong binders to HLA-B*0702 (%Rank<0.5), here only 1 and 2 are recognized by the TCR. 4-6 are examples of peptides that are predicted as poor binders to MHC (%Rank>2), but the TCR is still able to recognize 4 and 5. The respective peptide sequences (substitutions in red), %Rank, MFI and percentages out of total CD8 T cells are indicated within the contour plots. The fluorescent based multimer stainings were performed twice.
Supplementary Figure 6 The effect of peptide–MHC interaction on analyzing the recognition pattern of the HLA-A*2402EWW-engaging TCR
(a) Sequence logo showing the amino acid preferences for peptide binding to HLA-A*2402 constructed with NetMHCpan-4.049 by finding the binding cores of the top 1% strongest predicted binders among 100,000 natural random 10-mer peptides and visualizing these using R with the Shannon method in Seq2Logo48. (b) Shows the sequencing output expressed in log2FC after enrichment of only the correctly folded pMHC amongst the full DNA barcode-labeled MHC multimer library comprising all 191 single substitution EWWRSGGFSF variants (Supplementary Data 4 and Supplementary Note 3). Correctly folded pMHC were captured on W6/32 beads that were subsequently sorted. The pMHC associated DNA barcodes were amplified and sequenced. This was performed at T0, which was just after collecting and reducing the volume of the MHC multimer panel, T4, after four hours incubation at RT, and at T20, after 20 hours incubation at RT. Each dot represents one peptide-MHC variation. The coloring indicates the position of a given amino acid substitution and the asterisks indicates the original peptide. (c) Dot plots from staining the HLA-A*2402EWW engaging TCR with fluorescently labeled MHC multimers carrying one of five variations of EWWRSGGFSF and an irrelevant peptide (p*, the UV conditional peptide). 1 and 2 are examples of peptides predicted as strong binders to HLA-A*2402 MHC (%Rank<0.5), only peptide 1 is recognized by the TCR. 3-5 are examples of peptides that are predicted as poor binders to MHC (%Rank>2), but the TCR is still able to recognize all three peptides. The respective peptide sequences (substitutions in red), %Rank, MFI and percentages out of total CD8 T cells are indicated within the contour plots. The experiments were performed twice.
Supplementary Figure 7 In silico modeling of the APNCYGNIPL peptide conformation when bound to HLA-B*0702
(a) Structural model of the original peptide, APNCYGNIPL, binding to HLA-B*0702. Color scaling indicates the importance of a given amino acid for retaining the conformation of the original peptide when bound to the MHC (the information content is listed in Supplementary Data 5). (b) Bar plot of the FoldX derived ΔΔG energy between the original peptide (APNCYGNIPL) and HLA-B*0702, as well as all the peptide variants created by substituting each peptide position (x-axis) with all naturally occurring amino acids (n=19). If the ΔΔG is larger than zero it indicates that the given amino acid substitution has destabilized the peptide-MHC interaction while a ΔΔG smaller than zero indicates that the substitution has stabilized the peptide-MHC interaction. (c) Sequence logo showing the structurally predicted peptide-MHC binding preference for HLA-B*0702APN based on the energy change (ΔΔ) calculated in b.
Supplementary Figure 8 In silico modeling of the EWWRSGGFSF peptide conformation when bound to HLA-A*2402
(a) Structural model of original peptide, EWWRSGGFSF, binding to HLA-A*2402. Color scaling indicates the importance of a given amino acid for retaining the conformation of the peptide when bound to the MHC (the information content is listed in Supplementary Data 6). (b) Bar plot of the FoldX derived ΔΔG energy between the original peptide (EWWRSGGFSF) and HLA-A*2402, as well as all the peptide variants created by substituting each peptide position (x-axis) with all naturally occurring amino acids (n=19). If the ΔΔG is larger than zero it indicates that the given amino acid substitution has destabilized the peptide-MHC interaction while a ΔΔG smaller than zero indicates that the substitution has stabilized the peptide-MHC interaction. (c) Sequence logo showing the structurally predicted peptide-MHC binding preference for HLA-A*2402EWW based on the energy change (ΔΔ) calculated in b.
Supplementary Figure 9 The HLA-A*2402EWW-engaging TCRs recognize peptides with multiple amino acid substitutions
(a) Heat map showing the amino acid preferences of the TCR recognizing the HLA-A*2402 restricted peptide, EWWRSGGFSF, expressed as log2FC of read counts relative to a triplicate baseline sample (see Supplementary Note). The TCR transduced clones were stained with a MHC multimer library composed of peptides with all single amino acid substitutions of the original EWWRSGGFSF peptide, as well as double amino acid substitutions covering 12 naturally occurring amino acids, where positions 4-8 were substituted two amino acids at a time. The heat map includes only double amino acid substitution variants (the effect of single amino acid substitutions can be seen in Fig. 1 and Supplementary Fig. 4). Double substitution variants resulting in peptides with a %Rank>0.5 were excluded from the analysis (total number of double substitution peptide variants, n=776, Supplementary Data 7). The data largely confirm the findings when screening only with the single substitution library, that there is some flexibility of the amino acids at positions 4 and 5, while almost no other double substitutions are tolerated. The barcode-based analysis of double substitutions is representative of duplicate analyses. (b) Dot plots from staining the HLA-A*2402EWW engaging TCR with fluorescently labeled MHC multimers carrying one of 14 variations of EWWRSGGFSF and an irrelevant peptide (p*, the UV conditional peptide). The variants are comprised of peptides with 3 to 8 amino acid substitutions compared to the original peptide sequence (1). The substitutions were either at positions that had no preference for certain amino acids for TCR recognition (positions 1, 3 or 9, as visualized in the TCR fingerprint, Fig. 1f) or substituted with an amino acid that was prominent in the TCR fingerprint (positions 2, 4, 5, 6, 8 and 10). The MHC anchor at positions 2 were in some cases instead substituted with tyrosine (Y), which is the preferred amino acid for the peptide-MHC interaction (peptides 8, 9, 10, 11, 13, see Supplementary Fig. 6). We saw that the TCR were able to recognize peptide variants with up to 6 amino acids substituted compared to the original sequence (peptide 4 and 6) when the substituted amino acids were represented in the TCR fingerprint. The respective peptide sequences (substitutions in red), %Rank, MFI and percentages out of total CD8 T cells are indicated within the contour plots. The fluorescent-based MHC multimer stainings were performed once.
Supplementary Figure 10 The DNA barcode-based MHC multimer analysis of OT-1 and OT-3 T cells, showing read counts, log2FC and data from an irrelevant sample
The underlying data from the DNA barcode-based analysis of splenocytes derived from OT-1 or OT-3 transgenic mice or T cells from wild-type C57BL/6 mice, respectively, used for Fig. 2a-d. The analysis was performed with all possible variations created by single-position amino acid substitutions of the H-2Kb restricted peptide, SIINFEKL. (a,b) Shows the clonality reduced read counts of each pMHC-associated DNA barcode plotted based on the hierarchy of counts derived from (a) screening the OT-1 T cells or (b) screening the OT-3 T cells. The obtained read counts from screening splenocytes from wild-type C57BL/6 mice are also plotted. (c-h) Heat map showing the read counts or log2FC derived from screening of (c,d) the OT-1 T cells, (e,f) the OT-3 T cells or (g,h) the wild-type C57BL/6 splenocytes respectively. Each row represents a given amino acid and each column a position in the peptide sequence. The amino acids of the original peptide target are marked with black borders. The analysis was performed once.
Supplementary Figure 11 Fluorescence-based MHC multimer analyses of OT-1 and OT-3 T cells
(a) Dot plots from the fluorescent-based MHC multimer staining of splenocytes derived from OT-1 or OT-3 mice. MHC tetramers carrying one of seven variations of SIINFEKL were applied in single color MHC multimer analyses. When comparing the staining of OT-1 and OT-3 in relation to their binding to the original peptide (1), the most variation between the TCRs is seen when staining with MHC multimers carrying peptide 3, 6 or 7. Peptide 3, which has a K to R substitution at position 7, leads to a higher MFI of the OT-1 T cells but a lower MFI of the OT-3 T cells. The opposite is seen for peptides 6 and 7, with N to P or S to M substitutions at position 4 and 1 respectively. When staining with the corresponding tetramers, the OT-1 T cells has a marked decreased in MFI, whereas the MFI of the OT-3 T cells is retained. These binding properties reflects the amino acid preferences evident from the individual TCR fingerprints (Fig. 2c,d). The respective peptide sequences, MFI and percentages out of total CD8+ T cells are indicated adjacent to or within the contour plots. All peptide variants are predicted to be strong binders of H-2Kb (%Rank<0.5). b) Dot plots from staining OT-1, OT-3 or wild-type C57BL/6 splenocytes with the full panel of 153 DNA barcode-labeled MHC multimer variants, a single irrelevant MHC multimer carrying the UV-conditional peptide (p*) or the original peptide SIINFEKL. The MFI and percentages out of total CD8 T cells are indicated within the contour plots. The MHC multimer stainings were performed once.
Supplementary Figure 12 Detailed information on the 12 HLA-A*0201KLL-engaging TCRs
(a) Principal component analysis (PCA) to visualize the diversity of the 12 TCR fingerprints of the MCC-derived T cells from Fig. 2e. Each dot represents one T cell clone (n=12 individual T cell clones). (b) Bar plots of the cytokine secretion of clone 2 and clone 5, w876, after stimulating the clonal T cells with peptides containing alanine substitutions at the indicated positions. The bars correspond to the individual stainings of the data shown in Fig. 2f. The grey bars indicate the original peptide that has an alanine at position 6. Cytokine secretion was determined once. (c) A correlation matrix showing the interrelationship between each of the 1000 peptides (predicted from the human proteome) that match best the individual TCR fingerprints of the 12 MCC clones (n=12,000) of Fig. 2e. Correlation was determined using Pearson’s correlation coefficient. Similarity is depicted on both color grading and size of dots. (d) Screening for T cell recognition of 75 peptides that are potentially cross-recognized by one or more of the 12 clonal T cells that has the HLA-A*A0201 restricted KLLEIAPNC peptide as original target. A library including 75 potentially cross-recognized peptides were applied in a DNA barcode-based MHC multimer analysis (Fig. 3a and Supplementary Data 12). The clonality reduced read counts resulting from the DNA barcode-based screen of all 75 pMHC multimers and all 12 clones are plotted (y-axis) according to %Rank score (x-axis). Dotted line at x=2 marks the recommended cut-off of peptides that are considered as binders to MHC. The closed symbol indicates a response that were also stained with fluorescently labeled MHC tetramers. The plotted data is mean of duplicate analyses and error bars indicate range of duplicates.
Supplementary Figure 13 General gating strategy
(a) Shows an example of the initial gating of CD8+ T cells. (b) Shows an example of the multimer positive population of total CD8+ T cells, which was sorted in a DNA barcode-based analysis. This gating strategy was applied for all MHC multimer analyses.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13 (PDF 3170 kb)
Supplementary Information
Supplementary Data 1, 2, 5, 6, 9, 11 and 13 and Supplementary Notes 1–4 (PDF 610 kb)
Supplementary Data 3
APNCYGNIPL peptide variants applied in Figure 1a–c,g and Supplementary Figures 3 and 5 (XLSX 12 kb)
Supplementary Data 4
EWWRSGGFSF peptide variants applied in Figure 1d–f,h–k and Supplementary Figures 4 and 6 (XLSX 12 kb)
Supplementary Data 7
EWWRSGGFSF double substitution variants applied in Figure 1i–k and Supplementary Figure 5. The MHC multimer panel comprised these 776 peptides and the 191 single substitution peptides listed in Supplementary Data 2. (XLSX 26 kb)
Supplementary Data 8
SIINFEKL peptide variants applied in Figure 2a–d and Supplementary Figures 10 and 11 (XLSX 11 kb)
Supplementary Data 10
KLLEIAPNC peptide variants applied in Figure 2 and Supplementary Figure 12 (XLSX 12 kb)
Supplementary Data 12
The 75 peptides comprising the top ten of the FIMO-based priority list that best match each of the TCR fingerprints of the 12 HLA-A*0201 MCCKLL-engaging TCRs, including the original KLLEIAPNC target. Related to Figure 3. (XLSX 619 kb)
Rights and permissions
About this article
Cite this article
Bentzen, A., Such, L., Jensen, K. et al. T cell receptor fingerprinting enables in-depth characterization of the interactions governing recognition of peptide–MHC complexes. Nat Biotechnol 36, 1191–1196 (2018). https://doi.org/10.1038/nbt.4303
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.4303
This article is cited by
-
Novel insights into TCR-T cell therapy in solid neoplasms: optimizing adoptive immunotherapy
Experimental Hematology & Oncology (2024)
-
A bead-based method for high-throughput mapping of the sequence- and force-dependence of T cell activation
Nature Methods (2022)
-
Identification of neoantigens for individualized therapeutic cancer vaccines
Nature Reviews Drug Discovery (2022)
-
T cells targeted to TdT kill leukemic lymphoblasts while sparing normal lymphocytes
Nature Biotechnology (2022)
-
T-cell receptor-based therapy: an innovative therapeutic approach for solid tumors
Journal of Hematology & Oncology (2021)