Abstract
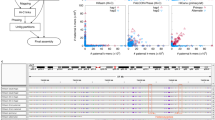
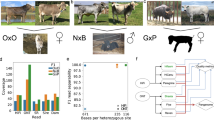
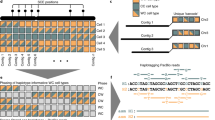
Complex allelic variation hampers the assembly of haplotype-resolved sequences from diploid genomes. We developed trio binning, an approach that simplifies haplotype assembly by resolving allelic variation before assembly. In contrast with prior approaches, the effectiveness of our method improved with increasing heterozygosity. Trio binning uses short reads from two parental genomes to first partition long reads from an offspring into haplotype-specific sets. Each haplotype is then assembled independently, resulting in a complete diploid reconstruction. We used trio binning to recover both haplotypes of a diploid human genome and identified complex structural variants missed by alternative approaches. We sequenced an F1 cross between the cattle subspecies Bos taurus taurus and Bos taurus indicus and completely assembled both parental haplotypes with NG50 haplotig sizes of >20 Mb and 99.998% accuracy, surpassing the quality of current cattle reference genomes. We suggest that trio binning improves diploid genome assembly and will facilitate new studies of haplotype variation and inheritance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Phillippy, A.M. New advances in sequence assembly. Genome Res. 27, xi–xiii (2017).
Koren, S. et al. Reducing assembly complexity of microbial genomes with single-molecule sequencing. Genome Biol. 14, R101 (2013).
Korlach, J. et al. De novo PacBio long-read and phased avian genome assemblies correct and add to reference genes generated with intermediate and short reads. Gigascience 6, 1–16 (2017).
Myers, E.W. et al. A whole-genome assembly of Drosophila. Science 287, 2196–2204 (2000).
Mouse Genome Sequencing Consortium. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002).
International Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Cao, H. et al. De novo assembly of a haplotype-resolved human genome. Nat. Biotechnol. 33, 617–622 (2015).
Steinberg, K.M. et al. Single haplotype assembly of the human genome from a hydatidiform mole. Genome Res. 24, 2066–2076 (2014).
Schneider, V.A. et al. Evaluation of GRCh38 and de novo haploid genome assemblies demonstrates the enduring quality of the reference assembly. Genome Res. 27, 849–864 (2017).
Chaisson, M.J. et al. Resolving the complexity of the human genome using single-molecule sequencing. Nature 517, 608–611 (2015).
Batzoglou, S. et al. ARACHNE: a whole-genome shotgun assembler. Genome Res. 12, 177–189 (2002).
Vinson, J.P. et al. Assembly of polymorphic genomes: algorithms and application to Ciona savignyi. Genome Res. 15, 1127–1135 (2005).
Levy, S. et al. The diploid genome sequence of an individual human. PLoS Biol. 5, e254 (2007).
Patterson, M. et al. WhatsHap: weighted haplotype assembly for future-generation sequencing reads. J. Comput. Biol. 22, 498–509 (2015).
Edge, P., Bafna, V. & Bansal, V. HapCUT2: robust and accurate haplotype assembly for diverse sequencing technologies. Genome Res. 27, 801–812 (2017).
Larkin, D.M. et al. Whole-genome resequencing of two elite sires for the detection of haplotypes under selection in dairy cattle. Proc. Natl. Acad. Sci. USA 109, 7693–7698 (2012).
Yang, H., Chen, X. & Wong, W.H. Completely phased genome sequencing through chromosome sorting. Proc. Natl. Acad. Sci. USA 108, 12–17 (2011).
Falconer, E. & Lansdorp, P.M. Strand-seq: a unifying tool for studies of chromosome segregation. Semin. Cell Dev. Biol. 24, 643–652 (2013).
Selvaraj, S., R Dixon, J., Bansal, V. & Ren, B. Whole-genome haplotype reconstruction using proximity-ligation and shotgun sequencing. Nat. Biotechnol. 31, 1111–1118 (2013).
Chin, C.S. et al. Phased diploid genome assembly with single-molecule real-time sequencing. Nat. Methods 13, 1050–1054 (2016).
Weisenfeld, N.I., Kumar, V., Shah, P., Church, D.M. & Jaffe, D.B. Direct determination of diploid genome sequences. Genome Res. 27, 757–767 (2017).
Seo, J.S. et al. De novo assembly and phasing of a Korean human genome. Nature 538, 243–247 (2016).
Porubsky, D. et al. Dense and accurate whole-chromosome haplotyping of individual genomes. Nat. Commun. 8, 1293 (2017).
Matthews, B.J. et al. Improved Aedes aegypti mosquito reference genome assembly enables biological discovery and vector control. Nature (in the press).
International HapMap Consortium. The International HapMap Project. Nature 426, 789–796 (2003).
The 1000 Genomes Project Consortium. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
Eberle, M.A. et al. A reference data set of 5.4 million phased human variants validated by genetic inheritance from sequencing a three-generation 17-member pedigree. Genome Res. 27, 157–164 (2017).
Malinsky, M., Simpson, J.T. & Durbin, R. trio-sga: facilitating de novo assemblyof highly heterozygous genomes with parent–child trios. bioRxiv Preprint at https://www.biorxiv.org/content/early/2016/05/03/051516 (2016).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Vurture, G.W. et al. GenomeScope: fast reference-free genome profiling from short reads. Bioinformatics 33, 2202–2204 (2017).
Waterhouse, R.M. et al. BUSCO applications from quality assessments to gene prediction and phylogenomics. Mol. Biol. Evol. https://doi.org/10.1093/molbev/msx319 (2017).
Salzberg, S.L. et al. GAGE: a critical evaluation of genome assemblies and assembly algorithms. Genome Res. 22, 557–567 (2012).
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome Biol. 5, R12 (2004).
Nattestad, M. & Schatz, M.C. Assemblytics: a web analytics tool for the detection of variants from an assembly. Bioinformatics 32, 3021–3023 (2016).
Lamesch, P. et al. The Arabidopsis Information Resource (TAIR): improved gene annotation and new tools. Nucleic Acids Res. 40, D1202–D1210 (2012).
Dilthey, A.T. et al. High-accuracy HLA type inference from whole-genome sequencing data using population reference graphs. PLoS Comput. Biol. 12, e1005151 (2016).
Zimin, A.V. et al. A whole-genome assembly of the domestic cow, Bos taurus. Genome Biol. 10, R42 (2009).
Canavez, F.C. et al. Genome sequence and assembly of Bos indicus. J. Hered. 103, 342–348 (2012).
Zhou, Y. et al. Genome-wide CNV analysis reveals variants associated with growth traits in Bos indicus. BMC Genomics 17, 419 (2016).
Sedlazeck, F.J. et al. Accurate detection of complex structural variations using single-molecule sequencing. Nat. Methods 15, 461–468 (2018).
Jain, M. et al. Nanopore sequencing and assembly of a human genome with ultra-long reads. Nat. Biotechnol. 36, 338–345 (2018).
Li, H. Minimap and miniasm: fast mapping and de novo assembly for noisy long sequences. Bioinformatics 32, 2103–2110 (2016).
Kolmogorov, M., Yuan, J., Lin, Y. & Pevzner, P. Assembly of long error-prone reads using repeat graphs. bioRxiv Preprint at https://www.biorxiv.org/content/early2018/01/12/247148 (2018).
Chaisson, M.J., Mukherjee, S., Kannan, S. & Eichler, E.E. Duplications de novo using polyploid phasing. in International Conference on Research in Computational Molecular Biology (ed. Sahinalp S.) 117–133 (Springer, 2017).
English, A.C. et al. Mind the gap: upgrading genomes with Pacific Biosciences RS long-read sequencing technology. PLoS One 7, e47768 (2012).
Chin, C.S. et al. Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat. Methods 10, 563–569 (2013).
Loman, N.J., Quick, J. & Simpson, J.T. A complete bacterial genome assembled de novo using only nanopore sequencing data. Nat. Methods 12, 733–735 (2015).
Ma, B., Tromp, J. & Li, M. PatternHunter: faster and more sensitive homology search. Bioinformatics 18, 440–445 (2002).
Nattestad, M., Chin, C.-S. & Schatz, M.C. Ribbon: visualizing complex genome alignments and structural variation. bioRxiv Preprint at https://www.biorxiv.org/content/early/2016/10/20/082123 (2016).
Mapleson, D., Garcia Accinelli, G., Kettleborough, G., Wright, J. & Clavijo, B.J. KAT: a K-mer analysis toolkit to quality control NGS datasets and genome assemblies. Bioinformatics 33, 574–576 (2017).
Kajitani, R. et al. Efficient de novo assembly of highly heterozygous genomes from whole-genome shotgun short reads. Genome Res. 24, 1384–1395 (2014).
Fofanov, Y. et al. How independent are the appearances of n-mers in different genomes? Bioinformatics 20, 2421–2428 (2004).
Dilthey, A., Cox, C., Iqbal, Z., Nelson, M.R. & McVean, G. Improved genome inference in the MHC using a population reference graph. Nat. Genet. 47, 682–688 (2015).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Garrison, E. & Marth, G. Haplotype-based variant detection from short-read sequencing. Preprint at https://arxiv.org/abs/1207.3907 (2012).
Casper, J. et al. The UCSC Genome Browser database: 2018 update. Nucleic Acids Res. 46, D762–D769 (2018).
Acknowledgements
We thank W, Thompson, K. Kuhn, K. McClure and R. Lee for technical assistance, and T. Graves-Lindsay and Washington University in St. Louis for public release of the PacBio NA12878 data. S.K., A.R., B.P.W. and A.M.P. were supported by the Intramural Research Program of the National Human Genome Research Institute, US National Institutes of Health. S.H. and J.L.W. were funded from the JS Davies bequest to the University of Adelaide. T.P.L.S. was supported by USDA-ARS Project 3040-31000-100-00D. D.M.B. was supported by USDA-ARS Project 5090-31000-026-00-D. This research was also supported by a grant of the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (grant number: HI17C2098). This work used the computational resources of the NIH HPC Biowulf cluster (https://hpc.nih.gov).
Author information
Authors and Affiliations
Contributions
A.M.P. and T.P.L.S. conceived and coordinated the project. S.K. and A.R. designed the trio-binning method. S.K., A.R. and B.P.W. implemented the software. S.K., A.R., B.P.W., A.T.D., D.M.B., S.B.K. and A.M.P. performed analyses. S.H. designed and performed breeding experiments and sample collections. J.L.W. contributed to development of the concept and provision of samples. T.P.L.S. performed sequencing. S.K., A.R., T.P.L.S., J.L.W. and A.M.P. wrote the manuscript. All of the authors approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
S.B.K. is a current employee of Pacific Biosciences.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1-21, Supplementary Tables 1-10, and Supplementary Note 1 (PDF 3748 kb)
Rights and permissions
About this article
Cite this article
Koren, S., Rhie, A., Walenz, B. et al. De novo assembly of haplotype-resolved genomes with trio binning. Nat Biotechnol 36, 1174–1182 (2018). https://doi.org/10.1038/nbt.4277
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.4277
This article is cited by
-
Haplotype-resolved assembly of a tetraploid potato genome using long reads and low-depth offspring data
Genome Biology (2024)
-
Genome assembly in the telomere-to-telomere era
Nature Reviews Genetics (2024)
-
Genome assembly provides insights into the genome evolution of Baccaurea ramiflora Lour.
Scientific Reports (2024)
-
Origin and evolution of the triploid cultivated banana genome
Nature Genetics (2024)
-
De novo diploid genome assembly using long noisy reads
Nature Communications (2024)