Abstract
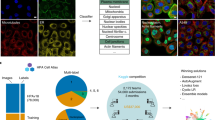
Pattern recognition and classification of images are key challenges throughout the life sciences. We combined two approaches for large-scale classification of fluorescence microscopy images. First, using the publicly available data set from the Cell Atlas of the Human Protein Atlas (HPA), we integrated an image-classification task into a mainstream video game (EVE Online) as a mini-game, named Project Discovery. Participation by 322,006 gamers over 1 year provided nearly 33 million classifications of subcellular localization patterns, including patterns that were not previously annotated by the HPA. Second, we used deep learning to build an automated Localization Cellular Annotation Tool (Loc-CAT). This tool classifies proteins into 29 subcellular localization patterns and can deal efficiently with multi-localization proteins, performing robustly across different cell types. Combining the annotations of gamers and deep learning, we applied transfer learning to create a boosted learner that can characterize subcellular protein distribution with F1 score of 0.72. We found that engaging players of commercial computer games provided data that augmented deep learning and enabled scalable and readily improved image classification.
This is a preview of subscription content, access via your institution
Access options






Similar content being viewed by others
References
Bouwer, J. et al. Petabyte data management and automated data workflow in neuroscience: delivering data from the instruments to the researcher's fingertips. Microsc. Microanal. 17, 276–277 (2011).
Ferrucci, D. et al. Building Watson: an overview of the DeepQA project. AI Magazine 31, 59–79 (2010).
Larrañaga, P. et al. Machine learning in bioinformatics. Brief. Bioinform. 7, 86–112 (2006).
Silver, D. et al. Mastering the game of Go with deep neural networks and tree search. Nature 529, 484–489 (2016).
Litjens, G. et al. A survey on deep learning in medical image analysis. Med. Image Anal. 42, 60–88 (2017).
Cohn, J.P. Citizen science: can volunteers do real research? Bioscience 58, 192–197 (2008).
Uhlen, M. et al. Towards a knowledge-based Human Protein Atlas. Nat. Biotechnol. 28, 1248–1250 (2010).
Thul, P.J. et al. A subcellular map of the human proteome. Science 356, eaai3321 (2017).
Boland, M.V. & Murphy, R.F. A neural network classifier capable of recognizing the patterns of all major subcellular structures in fluorescence microscope images of HeLa cells. Bioinformatics 17, 1213–1223 (2001).
Huang, K. & Murphy, R.F. Boosting accuracy of automated classification of fluorescence microscope images for location proteomics. BMC Bioinformatics 5, 78 (2004).
Newberg, J.Y. et al. Automated analysis of Human Protein Atlas immunofluorescence images. Proc. IEEE Int. Symp. Biomed. Imaging 5193229, 1023–1026 (2009).
Li, J., Newberg, J.Y., Uhlén, M., Lundberg, E. & Murphy, R.F. Automated analysis and reannotation of subcellular locations in confocal images from the Human Protein Atlas. PLoS One 7, e50514 (2012).
Li, J., Xiong, L., Schneider, J. & Murphy, R.F. Protein subcellular location pattern classification in cellular images using latent discriminative models. Bioinformatics 28, i32–i39 (2012).
Coelho, L.P. et al. Determining the subcellular location of new proteins from microscope images using local features. Bioinformatics 29, 2343–2349 (2013).
Chebira, A. et al. A multiresolution approach to automated classification of protein subcellular location images. BMC Bioinformatics 8, 210 (2007).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Pärnamaa, T. & Parts, L. Accurate classification of protein subcellular localization from high-throughput microscopy images using deep learning. G3 (Bethesda) 7, 1385–1392 (2017).
Kraus, O.Z., Ba, J.L. & Frey, B.J. Classifying and segmenting microscopy images with deep multiple instance learning. Bioinformatics 32, i52–i59 (2016).
Nathalie Japkowicz, S.S. The class imbalance problem: A systematic study. Intell. Data Anal. 6, 429–449 (2002).
Coelho, L.P., Peng, T. & Murphy, R.F. Quantifying the distribution of probes between subcellular locations using unsupervised pattern unmixing. Bioinformatics 26, i7–i12 (2010).
Zhao, T., Velliste, M., Boland, M.V. & Murphy, R.F. Object type recognition for automated analysis of protein subcellular location. IEEE Trans. Image Process. 14, 1351–1359 (2005).
Shen, Y.-Y.X.L.-X.Y.H.-B. Bioimage-based protein subcellular location prediction: a comprehensive review. Front. Comput. Sci. 12, 26–39 (2018).
Khatib, F. et al. Algorithm discovery by protein folding game players. Proc. Natl. Acad. Sci. USA 108, 18949–18953 (2011).
Khatib, F. et al. Crystal structure of a monomeric retroviral protease solved by protein folding game players. Nat. Struct. Mol. Biol. 18, 1175–1177 (2011).
Chris, J. et al. Galaxy Zoo: 'Hanny's Voorwerp', a quasar light echo? Mon. Not. R. Astron. Soc. 399, 129–140 (2009).
Clery, D. Galaxy evolution. Galaxy zoo volunteers share pain and glory of research. Science 333, 173–175 (2011).
Raddick, M.J. et al. Galaxy Zoo: exploring the motivations of citizen science volunteers. Astron. Educ. Rev. 9, 18 (2010).
Lee, J. et al. RNA design rules from a massive open laboratory. Proc. Natl. Acad. Sci. USA 111, 2122–2127 (2014).
Sørensen, J.J. et al. Exploring the quantum speed limit with computer games. Nature 532, 210–213 (2016).
Hughes, A. et al. Quantius: Generic, high-fidelity human annotation of scientific images at 105-clicks-per-hour. Preprint at https://doi.org/www.biorxiv.org/content/early/2017/07/15/164087 (2017).
Danielle, N., Shapiro, J.C. & Mueller, P.A. Using mechanical turk to study clinical populations. Clin. Pyschol. Sci. 1, 213–220 (2013).
Cox, J. et al. How is success defined and measured in online citizen science? A case study of Zooniverse projects. Comput. Sci. Eng. 17, 28–41 (2015).
Feng, W., Brandt, D. & Shah, D. A long-term study of a popular MMORPG. Proceedings of the 6th ACM SIGCOMM Workshop on Network and System Support for Games 19–24 (2007).
Warfield, S.K., Zou, K.H. & Wells, W.M. Simultaneous truth and performance level estimation (STAPLE): an algorithm for the validation of image segmentation. IEEE Trans. Med. Imaging 23, 903–921 (2004).
Snow, R., O'Connor, B., Jurafsky, D. & Ng, A. Cheap and fast, but is it good? Evaluating non-expert annotations for natural language tasks. Conference on Empirical Methods in Natural Language Processing 254–263 (2008).
Calise, S.J. et al. Glutamine deprivation initiates reversible assembly of mammalian rods and rings. Cell. Mol. Life Sci. 71, 2963–2973 (2014).
Carcamo, W.C. et al. Induction of cytoplasmic rods and rings structures by inhibition of the CTP and GTP synthetic pathway in mammalian cells. PLoS One 6, e29690 (2011).
Handfield, L.F., Chong, Y.T., Simmons, J., Andrews, B.J. & Moses, A.M. Unsupervised clustering of subcellular protein expression patterns in high-throughput microscopy images reveals protein complexes and functional relationships between proteins. PLOS Comput. Biol. 9, e1003085 (2013).
Hasanpour, S., Rouhani, M., Fayyaz, M. & Sabokrou, M. Lets keep it simple, Using simple architectures to outperform deeper and more complex architectures. Preprint at https://doi.org/arxiv.org/abs/1608.06037 (2016).
Acknowledgements
We acknowledge the staff of the Human Protein Atlas program for valuable contributions. We acknowledge the EVE Development team, the University of Reykjavik and the University of Iceland for assistance with the game implementation. We acknowledge MMOS Sarl for serving images and managing response collection and CCP hf and MMOS Sarl for financially supporting the image storage and serving throughout Project Discovery. Funding to E.L. was provided by the Knut and Alice Wallenberg Foundation.
Author information
Authors and Affiliations
Contributions
A.S., B.R., B.F., A.N. and E.L. conceived the study. M.H., A.S., B.F., E.L., D.P.S. and C.F.W. developed the methodology for the study. A.S. and B.R. developed the citizen science engine. L.C., H.L., S.R. and B.F. developed the game narrative and implementation. Project Discovery was played by thousands of players of EVE Online. D.P.S., L.Å., M.W., R.S. and E.L. provided game support. C.F.W., K.S. and D.P.S. developed the machine learning. D.P.S., C.F.W. and E.L. carried out data analysis and investigation. D.P.S., C.F.W. and E.L. wrote the manuscript. D.P.S. and C.F.W. created the figures. E.L. supervised and administered the project and acquired funding.
Corresponding author
Ethics declarations
Competing interests
A.S. and B.R. are founders of MMOS Sarl.
Integrated supplementary information
Supplementary Figure 1 Thirty-day retention for each month of Project Discovery.
Rows represent the month players joined Project Discovery, and columns represent the number of months the corresponding user group has been playing for.
Supplementary Figure 2 Individual player performance in Project Discovery
(a) Individual player accuracies (dots) for players with a minimum of 10 image evaluations show that player accuracy generally increases as players evaluate more samples (contour). Despite ~10% of players perform worse than naively guessing the most common class (Cytoplasm, blue dots), the consensus accuracy (black line) remains remarkably higher than the player average. Though a large number of poor players drop off after 100 samples or so, player performance remains remarkably unimproved over samples analyzed. (b) Player performance vs time spent per task (seconds) shows no discernable trend. This measure is confounded with time which players spent on other in-game actions with the interface open.
Supplementary Figure 3 Project Discovery performance relative to HPA v14.
(a) Gamer over-represented co-annotations with solution classes from the HPA Cell Atlas v14 (p<1e-2, one-tailed Binomial test, Bonferroni corrected by row, sample size indicated in parenthesis on each row/column) of gamer predicted labels (columns, blue), with expected co-localization frequencies from HPA v14 (rows, red). Columns with large numbers of significant over co-annotations represent generally over annotated classifications by the gamers (Nucleus, Cytoplasm, Aggresome, Microtubule ends). (b) Proportion of co-annotation in Project Discovery from gamer labels (columns, blue) with HPA Cell Atlas v14 labels (rows, red). Note particularly that novel classes (e.g. nucleoli rim) are co-annotated with their logical parent class (nucleoli) indicating successful refinement of labels.
Supplementary Figure 4 Schematic outline of how the different methods presented in this paper generate their annotations
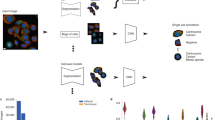
(a) Project Discovery (PD) let citizen scientists use a game interface to annotate images, taken from the Human Protein Atlas (HPA), into one or more of 29 different classes. (b) Localization Cellular Annotation Tool (Loc-CAT) is a neural network model which, using image derived features, annotates HPA images into one or more of 23 different classes. (c) Gamer Augmented Loc-CAT (GA Loc-CAT) uses image derived features in conjunction with player votes from PD to classify images from the HPA into one or more of 23 different classes. The votes from the gamers are presented as a p-value vector which is concatenated to the image features and fed to the Loc-CAT architecture. (d) Loc-CAT+ uses a separate neural network trained to estimate what players from PD would have voted for (“pseudo gamer”) together with the image features to classify images from the HPA into one or more of 23 different classes. The output from the “pseudo gamer” is concatenated to the feature vector and used as input to the Loc-CAT architecture.
Supplementary Figure 5 Overrepresented co-annotations in Loc-CAT+
Loc-CAT+ over-represented co-annotations with solution classes from the HPA Cell Atlas v14 (p<1e-2, one-tailed Binomial test, Bonferroni corrected by row, sample size indicated in parenthesis on each row/column) of Loc-CAT+ predicted labels (columns, blue), with expected co-localization frequencies from HPA v14 (rows, red). Columns with large numbers of significant over co-annotations (n>5) represent generally over annotated classifications by Loc-CAT+.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 (PDF 1623 kb)
Supplementary Table 1
Comparison of protein subcellular localization methods from fluorescent microscopy images (XLSX 15 kb)
Supplementary Table 2
Project Discovery optimized per-class cutoffs (XLS 19 kb)
Supplementary Table 3
Rods & Rings localized proteins found by Project Discovery (XLSX 10 kb)
Supplementary Table 4
Loc-CAT optimized per-class cutoffs (XLS 19 kb)
Supplementary Data Set 1
HPA version 14 “gold standard” annotations (ZIP 480 kb)
Supplementary Data Set 2
SLF feature names used in Loc-CAT DNN (XLSX 25 kb)
Supplementary Data Set 3
Expert reannotation results (TXT 9 kb)
Rights and permissions
About this article
Cite this article
Sullivan, D., Winsnes, C., Åkesson, L. et al. Deep learning is combined with massive-scale citizen science to improve large-scale image classification. Nat Biotechnol 36, 820–828 (2018). https://doi.org/10.1038/nbt.4225
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.4225
This article is cited by
-
PIFiA: self-supervised approach for protein functional annotation from single-cell imaging data
Molecular Systems Biology (2024)
-
Improving microbial phylogeny with citizen science within a mass-market video game
Nature Biotechnology (2024)
-
Labelling instructions matter in biomedical image analysis
Nature Machine Intelligence (2023)
-
Single-cell subcellular protein localisation using novel ensembles of diverse deep architectures
Communications Biology (2023)
-
Online citizen science with the Zooniverse for analysis of biological volumetric data
Histochemistry and Cell Biology (2023)