Abstract
Clinical translation of in vivo genome editing to treat human genetic diseases requires thorough preclinical studies in relevant animal models to assess safety and efficacy. A promising approach to treat hypercholesterolemia is inactivating the secreted protein PCSK9, an antagonist of the LDL receptor. Here we show that single infusions in six non-human primates of adeno-associated virus vector expressing an engineered meganuclease targeting PCSK9 results in dose-dependent disruption of PCSK9 in liver, as well as a stable reduction in circulating PCSK9 and serum cholesterol. Animals experienced transient, asymptomatic elevations of serum transaminases owing to the formation of T cells against the transgene product. Vector DNA and meganuclease expression declined rapidly, leaving stable populations of genome-edited hepatocytes. A second-generation PCSK9-specific meganuclease showed reduced off-target cleavage. These studies demonstrate efficient, physiologically relevant in vivo editing in non-human primates, and highlight safety considerations for clinical translation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Li, H. et al. In vivo genome editing restores haemostasis in a mouse model of haemophilia. Nature 475, 217–221 (2011).
Barzel, A. et al. Promoterless gene targeting without nucleases ameliorates haemophilia B in mice. Nature 517, 360–364 (2015).
Ran, F.A. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015).
Nelson, C.E. et al. In vivo genome editing improves muscle function in a mouse model of Duchenne muscular dystrophy. Science 351, 403–407 (2016).
Long, C. et al. Postnatal genome editing partially restores dystrophin expression in a mouse model of muscular dystrophy. Science 351, 400–403 (2016).
Tabebordbar, M. et al. In vivo gene editing in dystrophic mouse muscle and muscle stem cells. Science 351, 407–411 (2016).
Yin, H. et al. Structure-guided chemical modification of guide RNA enables potent non-viral in vivo genome editing. Nat. Biotechnol. 35, 1179–1187 (2017).
Nathwani, A.C. et al. Long-term safety and efficacy of factor IX gene therapy in hemophilia B. N. Engl. J. Med. 371, 1994–2004 (2014).
Nathwani, A.C. et al. Adenovirus-associated virus vector-mediated gene transfer in hemophilia B. N. Engl. J. Med. 365, 2357–2365 (2011).
Horton, J.D., Cohen, J.C. & Hobbs, H.H. Molecular biology of PCSK9: its role in LDL metabolism. Trends Biochem. Sci. 32, 71–77 (2007).
Fitzgerald, K. et al. Effect of an RNA interference drug on the synthesis of proprotein convertase subtilisin/kexin type 9 (PCSK9) and the concentration of serum LDL cholesterol in healthy volunteers: a randomised, single-blind, placebo-controlled, phase 1 trial. Lancet 383, 60–68 (2014).
Sijbrands, E.J. Inhibition of PCSK9 in familial hypercholesterolaemia. Lancet 380, 6–7 (2012).
Raal, F.J. et al. PCSK9 inhibition with evolocumab (AMG 145) in heterozygous familial hypercholesterolaemia (RUTHERFORD-2): a randomised, double-blind, placebo-controlled trial. Lancet 385, 331–340 (2015).
Horton, J.D., Cohen, J.C. & Hobbs, H.H. PCSK9: a convertase that coordinates LDL catabolism. J. Lipid Res. 50 (Suppl), S172–S177 (2009).
Ding, Q. et al. Permanent alteration of PCSK9 with in vivo CRISPR-Cas9 genome editing. Circ. Res. 115, 488–492 (2014).
Chadwick, A.C., Wang, X. & Musunuru, K. In vivo base editing of PCSK9 (proprotein convertase subtilisin/kexin type 9) as a therapeutic alternative to genome editing. Arterioscler. Thromb. Vasc. Biol. 37, 1741–1747 (2017).
Arnould, S. et al. The I-CreI meganuclease and its engineered derivatives: applications from cell modification to gene therapy. PEDS 24, 27–31 (2011).
Epinat, J.C. et al. A novel engineered meganuclease induces homologous recombination in yeast and mammalian cells. Nucleic Acids Res. 31, 2952–2962 (2003).
Antunes, M.S., Smith, J.J., Jantz, D. & Medford, J.I. Targeted DNA excision in Arabidopsis by a re-engineered homing endonuclease. BMC Biotechnol. 12, 86 (2012).
Honig, A. et al. Transient expression of virally delivered meganuclease in planta generates inherited genomic deletions. Mol. Plant 8, 1292–1294 (2015).
Arnould, S. et al. Engineered I-CreI derivatives cleaving sequences from the human XPC gene can induce highly efficient gene correction in mammalian cells. J. Mol. Biol. 371, 49–65 (2007).
Grizot, S. et al. Efficient targeting of a SCID gene by an engineered single-chain homing endonuclease. Nucleic Acids Res. 37, 5405–5419 (2009).
He, C. et al. Lentiviral protein delivery of meganucleases in human cells mediates gene targeting and alleviates toxicity. Gene Ther. 21, 759–766 (2014).
Grizot, S. et al. Generation of redesigned homing endonucleases comprising DNA-binding domains derived from two different scaffolds. Nucleic Acids Res. 38, 2006–2018 (2010).
Ménoret, S. et al. Generation of Rag1-knockout immunodeficient rats and mice using engineered meganucleases. FASEB J. 27, 703–711 (2013).
Redondo, P. et al. Molecular basis of xeroderma pigmentosum group C DNA recognition by engineered meganucleases. Nature 456, 107–111 (2008).
Zheng, Z. et al. Anchored multiplex PCR for targeted next-generation sequencing. Nat. Med. 20, 1479–1484 (2014).
Gao, B., Jeong, W.I. & Tian, Z. Liver: An organ with predominant innate immunity. Hepatology 47, 729–736 (2008).
Tsai, S.Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Kleinstiver, B.P. et al. Genome-wide specificities of CRISPR-Cas Cpf1 nucleases in human cells. Nat. Biotechnol. 34, 869–874 (2016).
D'Amour, K.A. et al. Production of pancreatic hormone-expressing endocrine cells from human embryonic stem cells. Nat. Biotechnol. 24, 1392–1401 (2006).
Cheng, X. et al. Self-renewing endodermal progenitor lines generated from human pluripotent stem cells. Cell Stem Cell 10, 371–384 (2012).
Covington, K.R. & Fuqua, S.A. Role of MTA2 in human cancer. Cancer Metastasis Rev. 33, 921–928 (2014).
Scartezini, M. et al. The PCSK9 gene R46L variant is associated with lower plasma lipid levels and cardiovascular risk in healthy U.K. men. Clin. Sci. (Lond.) 113, 435–441 (2007).
Lakoski, S.G., Lagace, T.A., Cohen, J.C., Horton, J.D. & Hobbs, H.H. Genetic and metabolic determinants of plasma PCSK9 levels. J. Clin. Endocrinol. Metab. 94, 2537–2543 (2009).
Mendell, J.R. et al. Single-dose gene-replacement therapy for spinal muscular atrophy. N. Engl. J. Med. 377, 1713–1722 (2017).
Rangarajan, S. et al. AAV5-factor VIII gene transfer in severe hemophilia A. N. Engl. J. Med. 377, 2519–2530 (2017).
Greig, J.A. et al. Non-clinical study examining AAV8.TBG.hLDLR vector-associated toxicity in chow-fed wild-type and LDLR+/− rhesus macaques. Hum. Gene Ther. Clin. Dev. 28, 39–50 (2017).
Davidoff, A.M. et al. Comparison of the ability of adeno-associated viral vectors pseudotyped with serotype 2, 5, and 8 capsid proteins to mediate efficient transduction of the liver in murine and nonhuman primate models. Mol. Ther. 11, 875–888 (2005).
Nietupski, J.B. et al. Systemic administration of AAV8-α-galactosidase A induces humoral tolerance in nonhuman primates despite low hepatic expression. Mol. Ther. 19, 1999–2011 (2011).
Chew, W.L. et al. A multifunctional AAV-CRISPR-Cas9 and its host response. Nat. Methods 13, 868–874 (2016).
Gao, G. et al. Adeno-associated virus-mediated gene transfer to nonhuman primate liver can elicit destructive transgene-specific T cell responses. Hum. Gene Ther. 20, 930–942 (2009).
Calcedo, R. et al. Immune responses in 101HEMB01, a phase 1/2 open-label, single ascending dose-finding trial of DTX101 (AAVrh10FIX) in patients with severe hemophilia B. Blood 130, 3333 (2017).
Guseva, N.V. et al. The NAB2-STAT6 gene fusion in solitary fibrous tumor can be reliably detected by anchored multiplexed PCR for targeted next-generation sequencing. Cancer Genet. 209, 303–312 (2016).
Amatu, A. et al. Novel CAD-ALK gene rearrangement is drugable by entrectinib in colorectal cancer. Br. J. Cancer 113, 1730–1734 (2015).
Lock, M. et al. Rapid, simple, and versatile manufacturing of recombinant adeno-associated viral vectors at scale. Hum. Gene Ther. 21, 1259–1271 (2010).
Lock, M., Alvira, M.R., Chen, S.J. & Wilson, J.M. Absolute determination of single-stranded and self-complementary adeno-associated viral vector genome titers by droplet digital PCR. Hum. Gene Ther. Methods 25, 115–125 (2014).
Calcedo, R., Vandenberghe, L.H., Gao, G., Lin, J. & Wilson, J.M. Worldwide epidemiology of neutralizing antibodies to adeno-associated viruses. J. Infect. Dis. 199, 381–390 (2009).
Bell, P. et al. Analysis of tumors arising in male B6C3F1 mice with and without AAV vector delivery to liver. Mol. Ther. 14, 34–44 (2006).
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30, 614–620 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Girardot, C., Scholtalbers, J., Sauer, S., Su, S.Y. & Furlong, E.E. Je, a versatile suite to handle multiplexed NGS libraries with unique molecular identifiers. BMC Bioinformatics 17, 419 (2016).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Tsai, S.Q., Topkar, V.V., Joung, J.K. & Aryee, M.J. Open-source guideseq software for analysis of GUIDE-seq data. Nat. Biotechnol. 34, 483 (2016).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Crooks, G.E., Hon, G., Chandonia, J.M. & Brenner, S.E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Schneider, T.D. & Stephens, R.M. Sequence logos: a new way to display consensus sequences. Nucleic Acids Res. 18, 6097–6100 (1990).
Calcedo, R. et al. Host immune responses to chronic adenovirus infections in human and nonhuman primates. J. Virol. 83, 2623–2631 (2009).
Calcedo, R. et al. Class I-restricted T-cell responses to a polymorphic peptide in a gene therapy clinical trial for α-1-antitrypsin deficiency. Proc. Natl. Acad. Sci. USA 114, 1655–1659 (2017).
Somers, A. et al. Generation of transgene-free lung disease-specific human induced pluripotent stem cells using a single excisable lentiviral stem cell cassette. Stem Cells 28, 1728–1740 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate - a practical and powerful approach to multiple testing. J. R. Stat. Soc. Series B Stat. Methodol. 57, 289–300 (1995).
Acknowledgements
We thank the Penn Vector Core for supplying AAV vectors, the Program in Comparative Medicine for animal care and procedures, the Nucleic Acid Technologies Core for assistance with deep sequencing, the Immunology Core for assistance with immunology assays, the CHOP Human Pluripotent Stem Cell Core for providing CHOPWT4, H. Zhang for technical assistance, Y. Zhu for assistance on histology analyses, and C. Lee for WebLogo generation. We also thank J. Stewart for editorial assistance with the manuscript. This work was supported by Penn Medicine and Precision Biosciences.
Author information
Authors and Affiliations
Contributions
L.W., D.J., and J.M.W. conceived this study. L.W., J.S., C.B., J.Z., L.Y., R.C., A.S., and J.M.W. designed the experiments. C.B., L.Y., J. L., P.B., E.L.B., A.S., V.V.B., Z.H., and J.W. performed the experiments. P.C., Y.C., J.L., A.S., and M.L. conducted the bioinformatics analysis of the deep sequencing data. M.L. conducted statistical analyses. J.M.W., L.W., C.B., P.C., J.Z., L.Y., M.L., P.B., R.C., and D.J. wrote and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
J.M.W. is an advisor to, holds equity in, and has a sponsored research agreement with REGENXBIO; he also has a sponsored research agreement with Ultragenyx, Biogen, and Janssen, which are licensees of Penn technology. In addition, he has sponsored research agreements with Precision Biosciences and Moderna Therapeutics. J.M.W. holds equity in Solid Bio. J.M.W. and L.W. are inventors on patents that have been licensed to various biopharmaceutical companies. J.S., J.L., V.V.B., and D.J. are employees of, and hold equity in, Precision Biosciences.
Integrated supplementary information
Supplementary Figure 1 The M1PCSK9 and M2PCSK9 meganucleases.
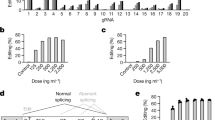
(a) Structure of wild-type I-CreI. The two I-CreI monomers comprising the functional homodimer are shown in gray. The primary amino acids involved in conferring target site specificity are colored red in the structure. The recognition sequence for I-CreI is shown beneath the structure with base pairs that are directly contacted by the enzyme shown in red. (b, c) Models of the single-chain (b) M1PCSK9 and (c) M2PCSK9 meganucleases and their intended target site in the PCSK9 gene are shown. The N- and C-terminal subunits of each meganuclease are shown in gray. Amino acids comprising the DNA-binding surface are colored teal and identified with numbering consistent with wild-type I-CreI. The PCSK9 target sequence is shown with base pairs that are believed to be contacted directly by the engineered meganucleases colored in teal. The three amino acids that differ between M1PCSK9 and M2PCSK9 are colored blue, as are the two base pairs in the target sequence that these amino acids are predicted to contact. (d) M1PCSK9 and M2PCSK9 introduce mutations at the intended site in PCSK9 in HEK-293 cells. Cells were mock transfected or electroporated with mRNA encoding either M1PCSK9 or M2PCSK9. Seventy-two hours post transfection, genomic DNA was isolated and amplified using PCSK9-specific PCR primers. PCR products were digested with T7-endonuclease and visualized on an agarose gel. The PCR products from meganuclease-transfected cells yielded smaller bands (indicated by black arrows) consistent with a high frequency of indel mutations at the intended target site. Deep sequencing of the PCR products revealed indel frequencies of 54.0% and 56.5% in cells transfected with M1PCSK9 and M2PCSK9, respectively. Similar results were obtained in three experiments. (e, f) In vivo evaluation of editing efficiency of M1PCSK9 and M2PCSK9 on episomal hPCSK9 cDNA delivered by AAV9.hPCSK9 vector in Rag1 KO mice. Adult male Rag1 KO mice first received an intravenous injection of AAV9.hPCSK9 vector (3.5x1010 GC), and 14 days later received a second vector injection of (e) AAV8-M1PCSK9 or (f) AAV8-M2PCSK9 at three doses (5.0x1011, 1.0x1011, or 2.0x1010 GC). PCSK9 levels were measured in mouse serum samples collected on days 7, 14, 21, 28, and 56 and presented as percentage of levels on day 14. Data from each individual mouse are shown (n=3 mice per cohort).
Supplementary Figure 2 Serum lipid profiles of macaques in this study.
Serum samples collected from each macaque before and after AAV8 vector treatment were assayed for total cholesterol, HDL, LDL, and triglyceride (TRIG) levels.
Supplementary Figure 3 Indel analysis on the rhPCSK9 targeted locus by deep sequencing of PCR amplicons.
DNA isolated from liver biopsy sample was used as template for PCR (251 bp amplicon indicated as 1; 417 bp amplicons indicated as 2) and deep sequencing analysis. DNA isolated from each animal’s PBMCs (collected before study) served as a control.
Supplementary Figure 4 Detection of potential M1PCSK9- or M2PCSK9-mediated off-target sites in the rhesus monkey genome by GUIDE-seq.
LLC-MK2 cells, which are rhesus monkey kidney cells, were co-transfected with plasmids expressing M1PCSK9 or M2PCSK9 and 16 pmol or 50 pmol of blunt or sticky-ended dsODN (4 nt overhang at the 3’end). Cells were harvested five days later and genomic DNA was isolated for GUIDE-seq analysis. (a) On-target and the top 45 off-target reads by M1PCSK9 co-transfected with 50 pmol sticky-ended dsODN are shown. (b) On-target and the top 45 off-target reads by M2PCSK9 co-transfected with 50 pmol sticky-ended dsODN are shown. (c – f) A WebLogo representation of the off-target reads identified in each GUIDE-seq experiment are shown. Please see Supplementary Data 1 for the complete data set.
Supplementary Figure 5 Liver function test results of the macaques in this study.
Liver functional tests including alanine transaminase (ALT), aspartate transaminase (AST), alkaline phosphatase (ALP), and gamma-glutamyl transpeptidase (GGTP) were performed on serum samples collected before and after AAV8 vector treatment from each macaque.
Supplementary Figure 6 Examination of liver histopathology on biopsy samples collected at two time points following vector administration.
(a) Representative pictures of hematoxylin and eosin staining. One section per sample was examined. Scale bar, 50 μm. (b) Summary of liver histology findings.
Supplementary Figure 7 T-cell responses to the peptide libraries of M1PCSK9 (pool A, B), M2PCSK9 (M2PCSK9-S), or the AAV8 capsid library (pools AAV8 A, B, C) by IFN-γ ELISpot following AAV8 vector administration.
The red asterisk indicates a positive T-cell response against a particular peptide library, which is arbitrarily defined as a spot-forming unit (SFU) per million PBMCs > 55 and above three-fold of SFU on the medium control. Each PBMC sample obtained at each time point was assayed once.
Supplementary Figure 8 Mapping of T-cell dominant epitopes in M1PCSK9.
Mapping of the dominant epitopes in peptide pools of M1PCSK9 or M2PCSK9 was performed on PBMCs from RA1866, RA1829, and RA2125 by IFN-γ ELISpot using the pool-array method. Protein sequences of (a) M1PCSK9 and (b) M2PCSK9 are shown. Mapped potential T-cell epitopes are shown in bold and underlined. The three amino acids that are different from M1PCSK9 and M2PCSK9 are indicated in green. Red brackets indicate the boundary of pool A and B. (c) A list of identified potential T-cell epitopes is shown.
Supplementary Figure 9 Flow cytometry gate strategy.
(a) iPSC-derived hepatocytes stained with isotype control were used as a negative control. (b) iPSC-derived hepatocytes were stained with AAT. An ALB plot was gated according to the isotype control.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 (PDF 1965 kb)
Supplementary Dataset 1
Potential M1PCSK9- or M2PCSK9-mediated off-target sites in the rhesus macaque genome identified by GUIDE-seq in LLCMK2 cells. N = 5 experiments for each nuclease. (XLSX 48 kb)
Supplementary Dataset 2
Potential M2PCSK9-mediated off-target sites in the human genome identified by GUIDE-seq in iPSC-derived hepatocytes. N = 3 experiment conditions. (XLSX 281 kb)
Supplemental Tables
Supplementary Tables 1–4 (PDF 538 kb)
Rights and permissions
About this article
Cite this article
Wang, L., Smith, J., Breton, C. et al. Meganuclease targeting of PCSK9 in macaque liver leads to stable reduction in serum cholesterol. Nat Biotechnol 36, 717–725 (2018). https://doi.org/10.1038/nbt.4182
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.4182
This article is cited by
-
Targeting proprotein convertase subtilisin/kexin type 9 (PCSK9): from bench to bedside
Signal Transduction and Targeted Therapy (2024)
-
In situ combinatorial synthesis of degradable branched lipidoids for systemic delivery of mRNA therapeutics and gene editors
Nature Communications (2024)
-
Gene Editing for the Treatment of Hypercholesterolemia
Current Atherosclerosis Reports (2024)
-
Efficient elimination of MELAS-associated m.3243G mutant mitochondrial DNA by an engineered mitoARCUS nuclease
Nature Metabolism (2023)
-
Mitochondrial gene editing
Nature Reviews Methods Primers (2023)