Abstract
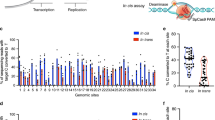
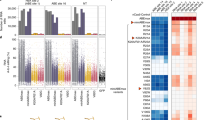
Base editors enable targeted single-nucleotide conversions in genomic DNA. Here we show that expression levels are a bottleneck in base-editing efficiency. We optimize cytidine (BE4) and adenine (ABE7.10) base editors by modification of nuclear localization signals (NLS) and codon usage, and ancestral reconstruction of the deaminase component. The resulting BE4max, AncBE4max, and ABEmax editors correct pathogenic SNPs with substantially increased efficiency in a variety of mammalian cell types.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Landrum, M.J. et al. Nucleic Acids Res. 44, D862–D868 (2016).
Komor, A.C., Kim, Y.B., Packer, M.S., Zuris, J.A. & Liu, D.R. Nature 533, 420–424 (2016).
Gaudelli, N.M. et al. Nature 551, 464–471 (2017).
Komor, A.C. et al. Sci. Adv. 3, eaao4774 (2017).
Li, G. et al. Protein Cell 8, 776–779 (2017).
Liang, P. et al. Protein Cell 8, 811–822 (2017).
Ryu, S.-M. et al. Nat. Biotechnol. https://doi.org/dx.doi.org/10.1038/nbt.4148 (2018).
Hess, G.T., Tycko, J., Yao, D. & Bassik, M.C. Mol. Cell 68, 26–43 (2017).
Kim, J.H. et al. PLoS One 6, e18556 (2011).
Suzuki, K. et al. Nature 540, 144–149 (2016).
Hanson, G. & Coller, J. Nat. Rev. Mol. Cell Biol. 19, 20–30 (2018).
Harms, M.J. & Thornton, J.W. Nat. Rev. Genet. 14, 559–571 (2013).
Wheeler, L.C., Lim, S.A., Marqusee, S. & Harms, M.J. Curr. Opin. Struct. Biol. 38, 37–43 (2016).
Risso, V.A., Gavira, J.A., Mejia-Carmona, D.F., Gaucher, E.A. & Sanchez-Ruiz, J.M. J. Am. Chem. Soc. 135, 2899–2902 (2013).
Krokan, H.E., Drabløs, F. & Slupphaug, G. Oncogene 21, 8935–8948 (2002).
Schenk, B. et al. J. Clin. Invest. 108, 1687–1695 (2001).
Bennett, D.L. & Woods, C.G. Lancet Neurol. 13, 587–599 (2014).
Liu, N. et al. Cell 173, 430–442 e417 (2018).
Amato, A. et al. Int. J. Lab. Hematol. 36, 13–19 (2014).
Badran, A.H. et al. Nature 533, 58–63 (2016).
Gibson, D.G. et al. Nat. Methods 6, 343–345 (2009).
Kim, Y.B. et al. Nat. Biotechnol. 35, 371–376 (2017).
Li, H. & Durbin, R. Bioinformatics 25, 1754–1760 (2009).
Hu, J.H. et al. Nature 556, 57–63 (2018).
UniProt Consortium Nucleic Acids Res. 46, 2699 (2018).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. J. Mol. Biol. 215, 403–410 (1990).
Katoh, K. & Standley, D.M. Mol. Biol. Evol. 30, 772–780 (2013).
Kalyaanamoorthy, S., Minh, B.Q., Wong, T.K.F., von Haeseler, A. & Jermiin, L.S. Nat. Methods 14, 587–589 (2017).
Nguyen, L.T., Schmidt, H.A., von Haeseler, A. & Minh, B.Q. Mol. Biol. Evol. 32, 268–274 (2015).
Hoang, D.T., Chernomor, O., von Haeseler, A., Minh, B.Q. & Vinh, L.S. Mol. Biol. Evol. 35, 518–522 (2018).
Yang, Z. Mol. Biol. Evol. 24, 1586–1591 (2007).
Acknowledgements
This work was supported by the Ono Pharma Foundation, DARPA HR0011-17-2-0049, US NIH RM1 HG009490, R01 EB022376, and R35 GM118062, and HHMI. Flow cytometry was supported by NCI P30CCA14051. L.W.K. is an NSF Graduate Research Fellow and was supported by NIH Training Grant T32 GM095450. J.L.D. gratefully acknowledges graduate fellowship support from the NSF and Hertz Foundation. We thank J. Coller, G. Hansen, M. Weiss, and A. Sharma for helpful discussions.
Author information
Authors and Affiliations
Contributions
L.W.K., J.L.D., C.W., J.M.L., T.T., G.A.N., and J.P.M. generated reagents and conducted experiments. C.W. and A.R. performed computational analyses. D.R.L. supervised the research. All authors contributed to writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
D.R.L. is a consultant and co-founder of Editas Medicine, Pairwise Plants, and Beam Therapeutics, companies that use genome editing. L.W.K., J.L.D., C.W., and D.R.L. have filed patent applications on aspects on this work. The authors declare no competing non-financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–17, Supplementary Note 1, Supplementary Sequences 1–5 (PDF 2354 kb)
Supplementary Data 1
MAFFT alignment APOBEC homologs in FASTA format. (TXT 112 kb)
Supplementary Data 2
Flow cytometry gating examples for all cell types used (PDF 3674 kb)
Supplementary Data 3
APOBEC tree in Newick format (TXT 32 kb)
Rights and permissions
About this article
Cite this article
Koblan, L., Doman, J., Wilson, C. et al. Improving cytidine and adenine base editors by expression optimization and ancestral reconstruction. Nat Biotechnol 36, 843–846 (2018). https://doi.org/10.1038/nbt.4172
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.4172