Abstract
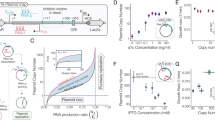
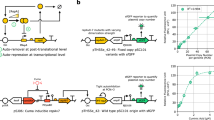
The internal environment of growing cells is variable and dynamic, making it difficult to introduce reliable parts, such as promoters, for genetic engineering. Here, we applied control-theoretic ideas to design promoters that maintained constant levels of expression at any copy number. Theory predicts that independence to copy number can be achieved by using an incoherent feedforward loop (iFFL) if the negative regulation is perfectly non-cooperative. We engineered iFFLs into Escherichia coli promoters using transcription-activator-like effectors (TALEs). These promoters had near-identical expression in different genome locations and plasmids, even when their copy number was perturbed by genomic mutations or changes in growth medium composition. We applied the stabilized promoters to show that a three-gene metabolic pathway to produce deoxychromoviridans could retain function without re-tuning when the stabilized-promoter-driven genes were moved from a plasmid into the genome.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
07 April 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41587-022-01300-7
References
Keasling, J.D. Synthetic biology and the development of tools for metabolic engineering. Metab. Eng.14, 189–195 (2012).
Pfleger, B.F., Pitera, D.J., Smolke, C.D. & Keasling, J.D. Combinatorial engineering of intergenic regions in operons tunes expression of multiple genes. Nat. Biotechnol.24, 1027–1032 (2006).
Torella, J.P. et al. Rapid construction of insulated genetic circuits via synthetic sequence-guided isothermal assembly. Nucleic Acids Res.42, 681–689 (2014).
Li, G.-W., Burkhardt, D., Gross, C. & Weissman, J.S. Quantifying absolute protein synthesis rates reveals principles underlying allocation of cellular resources. Cell157, 624–635 (2014).
Haseltine, E.L. & Arnold, F.H. Synthetic gene circuits: design with directed evolution. Annu. Rev. Biophys. Biomol. Struct.36, 1–19 (2007).
Smanski, M.J. et al. Functional optimization of gene clusters by combinatorial design and assembly. Nat. Biotechnol.32, 1241–1249 (2014).
Kosuri, S. et al. Composability of regulatory sequences controlling transcription and translation in Escherichia coli. Proc. Natl. Acad. Sci. USA110, 14024–14029 (2013).
Nielsen, A.A., Segall-Shapiro, T.H. & Voigt, C.A. Advances in genetic circuit design: novel biochemistries, deep part mining, and precision gene expression. Curr. Opin. Chem. Biol.17, 878–892 (2013).
Farasat, I. et al. Efficient search, mapping and optimization of multiprotein genetic systems in diverse bacteria. Mol. Syst. Biol.10, 731 (2014).
Kittleson, J.T., Wu, G.C. & Anderson, J.C. Successes and failures in modular genetic engineering. Curr. Opin. Chem. Biol.16, 329–336 (2012).
Cardinale, S. & Arkin, A.P. Contextualizing context for synthetic biology—identifying causes of failure of synthetic biological systems. Biotechnol. J.7, 856–866 (2012).
Arkin, A.P. A wise consistency: engineering biology for conformity, reliability, predictability. Curr. Opin. Chem. Biol.17, 893–901 (2013).
Carrier, T., Jones, K.L. & Keasling, J.D. mRNA stability and plasmid copy-number effects on gene expression from an inducible promoter system. Biotechnol. Bioeng.59, 666–672 (1998).
Wong Ng, J., Chatenay, D., Robert, J. & Poirier, M.G. Plasmid copy-number noise in monoclonal populations of bacteria. Phys. Rev. E81, 011909 (2010).
Lopilato, J., Bortner, S. & Beckwith, J. Mutations in a new chromosomal gene of Escherichia coli K-12, pcnB, reduce plasmid copy number of pBR322 and its derivatives. Mol. Gen. Genet.205, 285–290.
Węgrzyn, G. Replication of plasmids during bacterial response to amino acid starvation. Plasmid41, 1–16 (1999).
Lin-Chao, S. & Bremer, H. Effect of the bacterial growth rate on replication control of plasmid pBR322 in. Escherichia coli. Mol. Gen. Genet.203, 143–149.
Lin-Chao, S., Chen, W.-T. & Wong, T.-T. High copy number of the pUC plasmid results from a Rom–Rop-suppressible point mutation in RNA II. Mol. Microbiol.6, 3385–3393 (1992).
Stueber, D. & Bujard, H. Transcription from efficient promoters can interfere with plasmid replication and diminish expression of plasmid-specified genes. EMBO J.1, 1399–1404 (1982).
Cheah, U.E., Weigand, W.A. & Stark, B.C. Effects of recombinant plasmid size on cellular processes in Escherichia coli. Plasmid18, 127–134 (1987).
Corchero, J.L. & Villaverde, A. Plasmid maintenance in Escherichia coli recombinant cultures is dramatically, steadily and specifically influenced by features of the encoded proteins. Biotechnol. Bioeng.58, 625–632 (1998).
Chandler, M.G. & Pritchard, R.H. The effect of gene concentration and relative gene dosage on gene output in Escherichia coli. Mol. Gen. Genet.138, 127–141 (1975).
Block, D.H.S., Hussein, R., Liang, L.W. & Lim, H.N. Regulatory consequences of gene translocation in bacteria. Nucleic Acids Res.40, 8979–8992 (2012).
Kittleson, J.T., Cheung, S. & Anderson, J.C. Rapid optimization of gene dosage in E. coli using DIAL strains. J. Biol. Eng.5, 10 (2011).
Ma, W., Trusina, A., El-Samad, H., Lim, W.A. & Tang, C. Defining network topologies that can achieve biochemical adaptation. Cell138, 760–773 (2009).
Bleris, L. et al. Synthetic incoherent feedforward circuits show adaptation to the amount of their genetic template. Mol. Syst. Biol.7, 519 (2011).
Ang, J., Bagh, S., Ingalls, B.P. & McMillen, D.R. Considerations for using integral feedback control to construct a perfectly adapting synthetic gene network. J. Theor. Biol.266, 723–738 (2010).
Shen-Orr, S.S., Milo, R., Mangan, S. & Alon, U. Network motifs in the transcriptional regulation network of Escherichia coli. Nat. Genet.31, 64–68 (2002).
Mangan, S. & Alon, U. Structure and function of the feedforward-loop network motif. Proc. Natl. Acad. Sci. USA100, 11980–11985 (2003).
Klumpp, S., Zhang, Z. & Hwa, T. Growth-rate-dependent global effects on gene expression in bacteria. Cell139, 1366–1375 (2009).
Rogers, J.M. et al. Context influences on TALE–DNA binding revealed by quantitative profiling. Nat. Commun.6, 7440 (2015).
Copeland, M.F., Politz, M.C., Johnson, C.B., Markley, A.L. & Pfleger, B.F. A transcription-activator-like effector (TALE) induction system mediated by proteolysis. Nat. Chem. Biol.12, 254–260 (2016).
Zhang, F. et al. Efficient construction of sequence-specific TAL effectors for modulating mammalian transcription. Nat. Biotechnol.29, 149–153 (2011).
Jack, B.R. et al. Predicting the genetic stability of engineered DNA sequences with the EFM calculator. ACS Synth. Biol.4, 939–943 (2015).
Peterson, J. & Phillips, G.J. New pSC101-derivative cloning vectors with elevated copy numbers. Plasmid59, 193–201 (2008).
Ho, T.Q., Zhong, Z., Aung, S. & Pogliano, J. Compatible bacterial plasmids are targeted to independent cellular locations in Escherichia coli. EMBO J.21, 1864–1872 (2002).
Martínez-García, E., Aparicio, T., de Lorenzo, V. & Nikel, P.I. New transposon tools tailored for metabolic engineering of Gram-negative microbial cell factories. Front. Bioeng. Biotechnol.2, 46 (2014).
Fernandez-Rodriguez, J., Yang, L., Gorochowski, T.E., Gordon, D.B. & Voigt, C.A. Memory and combinatorial logic based on DNA inversions: dynamics and evolutionary stability. ACS Synth. Biol.4, 1361–1372 (2015).
Pósfai, G. et al. Emergent properties of reduced-genome Escherichia coli. Science312, 1044–1046 (2006).
Schmidt, F.R. Optimization and scale up of industrial fermentation processes. Appl. Microbiol. Biotechnol.68, 425–435 (2005).
Moser, F. et al. Genetic circuit performance under conditions relevant for industrial bioreactors. ACS Synth. Biol.1, 555–564 (2012).
Sontag, E.D. Mathematical Control Theory. Deterministic Finite Dimensional Systems 2nd ed. (Springer-Verlag, New York, 1998).
Voigt, C.A. Genetic parts to program bacteria. Curr. Opin. Biotechnol.17, 548–557 (2006).
Stanton, B.C. et al. Genomic mining of prokaryotic repressors for orthogonal logic gates. Nat. Chem. Biol.10, 99–105 (2014).
Goentoro, L., Shoval, O., Kirschner, M.W. & Alon, U. The incoherent feedforward loop can provide fold-change detection in gene regulation. Mol. Cell36, 894–899 (2009).
Mangan, S., Zaslaver, A. & Alon, U. The coherent feedforward loop serves as a sign-sensitive delay element in transcription networks. J. Mol. Biol.334, 197–204 (2003).
Tsang, J., Zhu, J. & van Oudenaarden, A. MicroRNA-mediated feedback and feedforward loops are recurrent network motifs in mammals. Mol. Cell26, 753–767 (2007).
Sakuma, T. et al. Repeating pattern of non-RVD variations in DNA-binding modules enhances TALEN activity. Sci. Rep.3, 3379 (2013).
Lee, C., Kim, J., Shin, S.G. & Hwang, S. Absolute and relative qPCR quantification of plasmid copy number in Escherichia coli. J. Biotechnol.123, 273–280 (2006).
Das, S., Noe, J.C., Paik, S. & Kitten, T. An improved arbitrary primed-PCR method for rapid characterization of transposon insertion sites. J. Microbiol. Methods63, 89–94 (2005).
Acknowledgements
This work was supported by US Office of Naval Research Multidisciplinary University Research Initiative grant no. N00014-16-1-2388 (T.H.S.-S., E.D.S., and C.A.V.), US National Institutes of Health National Institute of General Medical Sciences Center for Integrated Synthetic Biology grant no. P50-GM098792 (T.H.S.-S. and C.A.V.), US National Institutes of Standards and Technology grant no. 70-NANB16H164 (T.H.S.-S. and C.A.V.), US National Science Foundation Synthetic Biology Engineering Research Center grant no. EEC-0540879 (C.A.V.), and a Fannie and John Hertz Fellowship (T.H.S.-S.).
Author information
Authors and Affiliations
Contributions
T.H.S.-S., E.D.S. and C.A.V. conceived the study and designed the experiments; T.H.S.-S. performed the experiments and analyzed the data; and T.H.S.-S., E.D.S. and C.A.V. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–22, Supplementary Tables 1–3, and Supplementary Notes 1 and 2 (PDF 6864 kb)
Rights and permissions
About this article
Cite this article
Segall-Shapiro, T., Sontag, E. & Voigt, C. Engineered promoters enable constant gene expression at any copy number in bacteria. Nat Biotechnol 36, 352–358 (2018). https://doi.org/10.1038/nbt.4111
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.4111
This article is cited by
-
Dynamic monitoring of oscillatory enzyme activity of individual live bacteria via nanoplasmonic optical antennas
Nature Photonics (2023)
-
Microbial production of nutraceuticals: Metabolic engineering interventions in phenolic compounds, poly unsaturated fatty acids and carotenoids synthesis
Journal of Food Science and Technology (2023)
-
Advances in microbial engineering for the production of value-added products in a biorefinery
Systems Microbiology and Biomanufacturing (2023)
-
Redesigning regulatory components of quorum-sensing system for diverse metabolic control
Nature Communications (2022)
-
Inducible plasmid copy number control for synthetic biology in commonly used E. coli strains
Nature Communications (2022)