Abstract
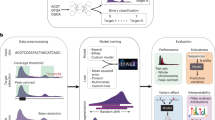
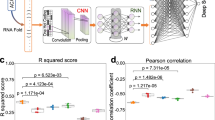
We present two algorithms to predict the activity of AsCpf1 guide RNAs. Indel frequencies for 15,000 target sequences were used in a deep-learning framework based on a convolutional neural network to train Seq-deepCpf1. We then incorporated chromatin accessibility information to create the better-performing DeepCpf1 algorithm for cell lines for which such information is available and show that both algorithms outperform previous machine learning algorithms on our own and published data sets.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
References
Zetsche, B. et al. Cell 163, 759–771 (2015).
Zetsche, B. et al. Nat. Biotechnol. 35, 31–34 (2017).
Hur, J.K. et al. Nat. Biotechnol. 34, 807–808 (2016).
Kim, Y. et al. Nat. Biotechnol. 34, 808–810 (2016).
Xu, R. et al. Plant Biotechnol. J. 15, 713–717 (2017).
Kim, D. et al. Nat. Biotechnol. 34, 863–868 (2016).
Kleinstiver, B.P. et al. Nat. Biotechnol. 34, 869–874 (2016).
Kim, H.K. et al. Nat. Methods 14, 153–159 (2017).
Doench, J.G. et al. Nat. Biotechnol. 34, 184–191 (2016).
Lee, C.M., Davis, T.H. & Bao, G. Exp. Physiol. doi:10.1113/EP086043 (2017).
Encode Project Consortium. Nature 489, 57–74 (2012).
Chari, R., Yeo, N.C., Chavez, A. & Church, G.M. ACS Synth. Biol. 6, 902–904 (2017).
Haeussler, M. et al. Genome Biol. 17, 148 (2016).
Yamano, T. et al. Cell 165, 949–962 (2016).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Genome Biol. 10, R25 (2009).
LeCun, Y., Bengio, Y. & Hinton, G. Nature 521, 436–444 (2015).
Goodfellow, I., Bengio, Y. & Courville, A. Deep Learning (MIT Press, 2016).
Min, S., Lee, B. & Yoon, S. Brief. Bioinform. 18, 851–869 (2017).
Alipanahi, B., Delong, A., Weirauch, M.T. & Frey, B.J. Nat. Biotechnol. 33, 831–838 (2015).
Kelley, D.R., Snoek, J. & Rinn, J.L. Genome Res. 26, 990–999 (2016).
Doench, J.G. et al. Nat. Biotechnol. 32, 1262–1267 (2014).
Wang, T., Wei, J.J., Sabatini, D.M. & Lander, E.S. Science 343, 80–84 (2014).
Chari, R., Mali, P., Moosburner, M. & Church, G.M. Nat. Methods 12, 823–826 (2015).
Moreno-Mateos, M.A. et al. Nat. Methods 12, 982–988 (2015).
Xu, H. et al. Genome Res. 25, 1147–1157 (2015).
Wong, N., Liu, W. & Wang, X. Genome Biol. 16, 218 (2015).
Bergstra, J. et al. in. Proc. 9th Python Sci. Conf. 3–10 (2010).
Kingma, D.P. & Ba, J. Preprint at https://arxiv.org/abs/1412.6980 (2014).
Srivastava, N., Hinton, G.E., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. J. Mach. Learn. Res. 15, 1929–1958 (2014).
Acknowledgements
The authors thank E.-S. Lee for proofreading and R. Gopalappa, N. Kim, S. Park, and J. Park for assisting in sample preparation. This work was supported in part by the National Research Foundation of Korea (grants 2017R1A2B3004198 (H.K.), 2017M3A9B4062403 (H.K.), 2013M3A9B4076544 (H.K.), 2014M3C9A3063541 (S.Y.)), Brain Korea 21 Plus Project (Yonsei University College of Medicine), Brain Korea 21 Plus Project (SNU ECE) in 2017, Institute for Basic Science (IBS; IBS-R026-D1), and the Korean Health Technology R&D Project, Ministry of Health and Welfare, Republic of Korea (grants HI17C0676 (H.K.), and HI16C1012 (H.K.)).
Author information
Authors and Affiliations
Contributions
H.K.K., M.S., and S.J. performed experiments to build data sets of AsCpf1 indel frequencies. S.M. and S.Y. developed the framework, and carried out the model training and computational validation. J.W.C. performed bioinformatic analyses. Y.K. and S.L. made substantial contributions to the performance of the experiments including cell culture and deep-sequencing. H.H.K. conceived and designed the study. H.K.K., S.M., S.Y., and H.H.K. analyzed the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Yonsei University and Seoul National University have filed a patent based on this work, in which H.K.K., S.M., M.S., S.J., S.Y., and H.K. are co-inventors.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14 and Supplementary Note (PDF 2816 kb)
Supplementary Tables
All tables that are included together, Supplementary tables 2, 4, and 6 (PDF 521 kb)
Supplementary Table 1
Source data used for this study. (XLSX 2463 kb)
Supplementary Table 3
Model selection results of Seq-deepCpf1 (XLSX 19 kb)
Supplementary Table 5
Oligonucleotides used in this study (XLSX 40 kb)
Supplementary Table 7
Confidence intervals for the result values (XLSX 15 kb)
Supplementary Code
The source code of Seq-deepCpf1 and DeepCpf1 (ZIP 750 kb)
Rights and permissions
About this article
Cite this article
Kim, H., Min, S., Song, M. et al. Deep learning improves prediction of CRISPR–Cpf1 guide RNA activity. Nat Biotechnol 36, 239–241 (2018). https://doi.org/10.1038/nbt.4061
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.4061
This article is cited by
-
Deep learning models to predict the editing efficiencies and outcomes of diverse base editors
Nature Biotechnology (2024)
-
Machine learning for antimicrobial peptide identification and design
Nature Reviews Bioengineering (2024)
-
Rapid screening of point mutations by mismatch amplification mutation assay PCR
Applied Microbiology and Biotechnology (2024)
-
Potential therapeutic strategies for osteoarthritis via CRISPR/Cas9 mediated gene editing
Reviews in Endocrine and Metabolic Disorders (2024)
-
CRISPR/Cas9: a powerful tool in colorectal cancer research
Journal of Experimental & Clinical Cancer Research (2023)