Abstract
Bacterial cell envelope protein (CEP) complexes mediate a range of processes, including membrane assembly, antibiotic resistance and metabolic coordination. However, only limited characterization of relevant macromolecules has been reported to date. Here we present a proteomic survey of 1,347 CEPs encompassing 90% inner- and outer-membrane and periplasmic proteins of Escherichia coli. After extraction with non-denaturing detergents, we affinity-purified 785 endogenously tagged CEPs and identified stably associated polypeptides by precision mass spectrometry. The resulting high-quality physical interaction network, comprising 77% of targeted CEPs, revealed many previously uncharacterized heteromeric complexes. We found that the secretion of autotransporters requires translocation and the assembly module TamB to nucleate proper folding from periplasm to cell surface through a cooperative mechanism involving the β-barrel assembly machinery. We also establish that an ABC transporter of unknown function, YadH, together with the Mla system preserves outer membrane lipid asymmetry. This E. coli CEP 'interactome' provides insights into the functional landscape governing CE systems essential to bacterial growth, metabolism and drug resistance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Babu, M. et al. Interaction landscape of membrane-protein complexes in Saccharomyces cerevisiae . Nature 489, 585–589 (2012).
Typas, A. & Sourjik, V. Bacterial protein networks: properties and functions. Nat. Rev. Microbiol. 13, 559–572 (2015).
Hurdle, J.G., O'Neill, A.J., Chopra, I. & Lee, R.E. Targeting bacterial membrane function: an underexploited mechanism for treating persistent infections. Nat. Rev. Microbiol. 9, 62–75 (2011).
Hagan, C.L., Silhavy, T.J. & Kahne, D. β-Barrel membrane protein assembly by the Bam complex. Annu. Rev. Biochem. 80, 189–210 (2011).
Whitfield, C. & Trent, M.S. Biosynthesis and export of bacterial lipopolysaccharides. Annu. Rev. Biochem. 83, 99–128 (2014).
Díaz-Mejía, J.J., Babu, M. & Emili, A. Computational and experimental approaches to chart the Escherichia coli cell-envelope-associated proteome and interactome. FEMS Microbiol. Rev. 33, 66–97 (2009).
Butland, G. et al. Interaction network containing conserved and essential protein complexes in Escherichia coli . Nature 433, 531–537 (2005).
Hu, P. et al. Global functional atlas of Escherichia coli encompassing previously uncharacterized proteins. PLoS Biol. 7, e96 (2009).
Nichols, R.J. et al. Phenotypic landscape of a bacterial cell. Cell 144, 143–156 (2011).
Babu, M. et al. Genetic interaction maps in Escherichia coli reveal functional crosstalk among cell envelope biogenesis pathways. PLoS Genet. 7, e1002377 (2011).
Armean, I.M., Lilley, K.S. & Trotter, M.W. Popular computational methods to assess multiprotein complexes derived from label-free affinity purification and mass spectrometry (AP-MS) experiments. Mol. Cell. Proteomics 12, 1–13 (2013).
Varjosalo, M. et al. Interlaboratory reproducibility of large-scale human protein-complex analysis by standardized AP-MS. Nat. Methods 10, 307–314 (2013).
Kim, S. & Pevzner, P.A.M.S.-G.F. MS-GF+ makes progress towards a universal database search tool for proteomics. Nat. Commun. 5, 5277 (2014).
Lee, I., Date, S.V., Adai, A.T. & Marcotte, E.M. A probabilistic functional network of yeast genes. Science 306, 1555–1558 (2004).
Sowa, M.E., Bennett, E.J., Gygi, S.P. & Harper, J.W. Defining the human deubiquitinating enzyme interaction landscape. Cell 138, 389–403 (2009).
Guruharsha, K.G. et al. A protein complex network of Drosophila melanogaster . Cell 147, 690–703 (2011).
Orchard, S. et al. The MIntAct project--IntAct as a common curation platform for 11 molecular interaction databases. Nucleic Acids Res. 42, D358–D363 (2014).
Arifuzzaman, M. et al. Large-scale identification of protein-protein interaction of Escherichia coli K-12. Genome Res. 16, 686–691 (2006).
Rajagopala, S.V. et al. The binary protein-protein interaction landscape of Escherichia coli . Nat. Biotechnol. 32, 285–290 (2014).
Szklarczyk, D. et al. STRING v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447–D452 (2015).
Vlasblom, J. et al. Novel function discovery with GeneMANIA: a new integrated resource for gene function prediction in Escherichia coli . Bioinformatics 31, 306–310 (2015).
Rodionova, I.A. et al. The phosphocarrier protein HPr of the bacterial phosphotransferase system globally regulates energy metabolism by directly interacting with multiple enzymes in Escherichia coli . J. Biol. Chem. 292, 14250–14257 (2017).
Wang, J.Z., Du, Z., Payattakool, R., Yu, P.S. & Chen, C.F. A new method to measure the semantic similarity of GO terms. Bioinformatics 23, 1274–1281 (2007).
Babu, M. et al. Quantitative genome-wide genetic interaction screens reveal global epistatic relationships of protein complexes in Escherichia coli . PLoS Genet. 10, e1004120 (2014).
Kumar, A. et al. Conditional epistatic interaction maps reveal global functional rewiring of genome integrity pathways in Escherichia coli . Cell Reports 14, 648–661 (2016).
Wan, C. et al. Panorama of ancient metazoan macromolecular complexes. Nature 525, 339–344 (2015).
Havugimana, P.C. et al. A census of human soluble protein complexes. Cell 150, 1068–1081 (2012).
Mogk, A., Huber, D. & Bukau, B. Integrating protein homeostasis strategies in prokaryotes. Cold Spring Harb. Perspect. Biol. 3, a004366 (2011).
Pinske, C. et al. Physiology and bioenergetics of [NiFe]-hydrogenase 2-catalyzed H2-consuming and H2-producing reactions in Escherichia coli . J. Bacteriol. 197, 296–306 (2015).
Wang, Y. et al. A supercomplex spanning the inner and outer membranes mediates the biogenesis of β-barrel outer membrane proteins in bacteria. J. Biol. Chem. 291, 16720–16729 (2016).
Leung, H.C., Xiang, Q., Yiu, S.M. & Chin, F.Y. Predicting protein complexes from PPI data: a core-attachment approach. J. Comput. Biol. 16, 133–144 (2009).
Moraes, T.F. & Reithmeier, R.A. Membrane transport metabolons. Biochim. Biophys. Acta 1818, 2687–2706 (2012).
Charbit, A., Reizer, J. & Saier, M.H. Jr. Function of the duplicated IIB domain and oligomeric structure of the fructose permease of Escherichia coli . J. Biol. Chem. 271, 9997–10003 (1996).
Kim, H.M., Park, Y.H., Yoon, C.K. & Seok, Y.J. Histidine phosphocarrier protein regulates pyruvate kinase A activity in response to glucose in Vibrio vulnificus. Mol. Microbiol. 96, 293–305 (2015).
Létoffé, S., Delepelaire, P. & Wandersman, C. The housekeeping dipeptide permease is the Escherichia coli heme transporter and functions with two optional peptide binding proteins. Proc. Natl. Acad. Sci. USA 103, 12891–12896 (2006).
Petrarca, P., Ammendola, S., Pasquali, P. & Battistoni, A. The Zur-regulated ZinT protein is an auxiliary component of the high-affinity ZnuABC zinc transporter that facilitates metal recruitment during severe zinc shortage. J. Bacteriol. 192, 1553–1564 (2010).
Pandey, R. et al. MntR(Rv2788): a transcriptional regulator that controls manganese homeostasis in Mycobacterium tuberculosis. Mol. Microbiol. 98, 1168–1183 (2015).
Horazdovsky, B.F. & Hogg, R.W. Genetic reconstitution of the high-affinity L-arabinose transport system. J. Bacteriol. 171, 3053–3059 (1989).
Brown, E.D. & Wright, G.D. Antibacterial drug discovery in the resistance era. Nature 529, 336–343 (2016).
Selkrig, J. et al. Discovery of an archetypal protein transport system in bacterial outer membranes. Nat. Struct. Mol. Biol. 19, 506–510, S1 (2012).
Gruss, F. et al. The structural basis of autotransporter translocation by TamA. Nat. Struct. Mol. Biol. 20, 1318–1320 (2013).
Heinz, E., Selkrig, J., Belousoff, M.J. & Lithgow, T. Evolution of the Translocation and Assembly Module (TAM). Genome Biol. Evol. 7, 1628–1643 (2015).
Henderson, I.R. & Owen, P. The major phase-variable outer membrane protein of Escherichia coli structurally resembles the immunoglobulin A1 protease class of exported protein and is regulated by a novel mechanism involving Dam and oxyR. J. Bacteriol. 181, 2132–2141 (1999).
Malinverni, J.C. & Silhavy, T.J. An ABC transport system that maintains lipid asymmetry in the gram-negative outer membrane. Proc. Natl. Acad. Sci. USA 106, 8009–8014 (2009).
Dong, H. et al. Structural insights into cardiolipin transfer from the Inner membrane to the outer membrane by PbgA in Gram-negative bacteria. Sci. Rep. 6, 30815 (2016).
Caufield, J.H., Abreu, M., Wimble, C. & Uetz, P. Protein complexes in bacteria. PLOS Comput. Biol. 11, e1004107 (2015).
Ostlund, G. et al. InParanoid 7: new algorithms and tools for eukaryotic orthology analysis. Nucleic Acids Res. 38, D196–D203 (2010).
Lan, G., Schulmeister, S., Sourjik, V. & Tu, Y. Adapt locally and act globally: strategy to maintain high chemoreceptor sensitivity in complex environments. Mol. Syst. Biol. 7, 475 (2011).
Cuthbertson, L., Mainprize, I.L., Naismith, J.H. & Whitfield, C. Pivotal roles of the outer membrane polysaccharide export and polysaccharide copolymerase protein families in export of extracellular polysaccharides in gram-negative bacteria. Microbiol. Mol. Biol. Rev. 73, 155–177 (2009).
Nadler, C. et al. Cycling of Etk and Etp phosphorylation states is involved in formation of group 4 capsule by Escherichia coli . PLoS One 7, e37984 (2012).
Käll, L., Krogh, A. & Sonnhammer, E.L. A combined transmembrane topology and signal peptide prediction method. J. Mol. Biol. 338, 1027–1036 (2004).
Hayat, S., Peters, C., Shu, N., Tsirigos, K.D. & Elofsson, A. Inclusion of dyad-repeat pattern improves topology prediction of transmembrane β-barrel proteins. Bioinformatics 32, 1571–1573 (2016).
Petersen, T.N., Brunak, S., von Heijne, G. & Nielsen, H. SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat. Methods 8, 785–786 (2011).
Orfanoudaki, G. & Economou, A. Proteome-wide subcellular topologies of E. coli polypeptides database (STEPdb). Mol. Cell. Proteomics 13, 3674–3687 (2014).
Horler, R.S., Butcher, A., Papangelopoulos, N., Ashton, P.D. & Thomas, G.H. EchoLOCATION: an in silico analysis of the subcellular locations of Escherichia coli proteins and comparison with experimentally derived locations. Bioinformatics 25, 163–166 (2009).
Babu, M. et al. Sequential peptide affinity purification system for the systematic isolation and identification of protein complexes from Escherichia coli . Methods Mol. Biol. 564, 373–400 (2009).
Butland, G. et al. eSGA: E. coli synthetic genetic array analysis. Nat. Methods 5, 789–795 (2008).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 0008 (2006).
Datsenko, K.A. & Wanner, B.L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 97, 6640–6645 (2000).
Zhang, Z. et al. Functional characterization of the heterooligomeric EbrAB multidrug efflux transporter of Bacillus subtilis . Biochemistry 46, 5218–5225 (2007).
Lutz, R. & Bujard, H. Independent and tight regulation of transcriptional units in Escherichia coli via the LacR/O, the TetR/O and AraC/I1-I2 regulatory elements. Nucleic Acids Res. 25, 1203–1210 (1997).
Klumpp, S., Zhang, Z. & Hwa, T. Growth rate-dependent global effects on gene expression in bacteria. Cell 139, 1366–1375 (2009).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Xu, Z. & Hao, B. CVTree update: a newly designed phylogenetic study platform using composition vectors and whole genomes. Nucleic Acids Res. 37, W174–W178 (2009).
Boc, A., Diallo, A.B. & Makarenkov, V. T-REX: a web server for inferring, validating and visualizing phylogenetic trees and networks. Nucleic Acids Res. 40, W573–W579 (2012).
Remm, M., Storm, C.E. & Sonnhammer, E.L. Automatic clustering of orthologs and in-paralogs from pairwise species comparisons. J. Mol. Biol. 314, 1041–1052 (2001).
Kitagawa, M. et al. Complete set of ORF clones of Escherichia coli ASKA library (a complete set of E. coli K-12 ORF archive): unique resources for biological research. DNA Res. 12, 291–299 (2005).
Bligh, E.G. & Dyer, W.J. A rapid method of total lipid extraction and purification. Can. J. Biochem. Physiol. 37, 911–917 (1959).
Oursel, D. et al. Lipid composition of membranes of Escherichia coli by liquid chromatography/tandem mass spectrometry using negative electrospray ionization. Rapid Commun. Mass Spectrom. 21, 1721–1728 (2007).
Garrett, T.A., O'Neill, A.C. & Hopson, M.L. Quantification of cardiolipin molecular species in Escherichia coli lipid extracts using liquid chromatography/electrospray ionization mass spectrometry. Rapid Commun. Mass Spectrom. 26, 2267–2274 (2012).
Tyurina, Y.Y. et al. Characterization of cardiolipins and their oxidation products by LC-MS analysis. Chem. Phys. Lipids 179, 3–10 (2014).
Hsu, F.F. & Turk, J. Characterization of cardiolipin as the sodiated ions by positive-ion electrospray ionization with multiple stage quadrupole ion-trap mass spectrometry. J. Am. Soc. Mass Spectrom. 17, 1146–1157 (2006).
van den Ent, F. & Löwe, J. RF cloning: a restriction-free method for inserting target genes into plasmids. J. Biochem. Biophys. Methods 67, 67–74 (2006).
Charbonneau, M.E., Berthiaume, F. & Mourez, M. Proteolytic processing is not essential for multiple functions of the Escherichia coli autotransporter adhesin involved in diffuse adherence (AIDA-I). J. Bacteriol. 188, 8504–8512 (2006).
Côté, J.P., Charbonneau, M.E. & Mourez, M. Glycosylation of the Escherichia coli TibA self-associating autotransporter influences the conformation and the functionality of the protein. PLoS One 8, e80739 (2013).
Heise, T. & Dersch, P. Identification of a domain in Yersinia virulence factor YadA that is crucial for extracellular matrix-specific cell adhesion and uptake. Proc. Natl. Acad. Sci. USA 103, 3375–3380 (2006).
Acknowledgements
We thank T. Silhavy, C. Whitfield, J.P. Côté, M. Mourez and P. Dersch for generously providing plasmids and reagents. We are also grateful to R. Reithmeier and M. Jessulat (Babu Lab) for advice. This work was supported by grants from the Canadian Institutes of Health Research to J.F.G., J.P. and A.E. (MOP-106449), J.P., A.E. and M.B. (PJT -148831) and T.F.M. (MOP-115182); the Ontario Ministry of Education and Innovation to T.F.M. and A.E.; the National Institutes of Health to P.U., M.H.S. and A.E. (GM109895); the Natural Sciences and Engineering Research Council of Canada to T.F.M. (DG-40197), A.Go. (DG-315735), J.P. (DG-06664), and M.B. (DG-20234); and the Canada Foundation for Innovation to T.F.M., M.B. and A.E.
Author information
Authors and Affiliations
Contributions
M.B. and A.E. designed and together supervised the project. Y.P. and O.P. constructed tagged strains. O.K., S.K. and V.D. performed protein purifications, while H.G. and Z.M. performed MS analyses. S.P. performed database searches and curation, and created the web portal. C.C., S.K., J.M., V.K., I.R., A.Ga., H.A., E.H., Z.Z., A.V., D.B., M.H., C.C.L., M.E.B., Y.H., M.S. and S.V.R. performed the follow-up and validation experiments. C.B.-T., S.P., Q.Z., Y.J., A.K., J.H.C. and S.W. performed the scoring and data analysis. M.B. and A.E. wrote the manuscript, with input from C.B.-T., C.C., J.V., A.Go., J.F.G., M.S., P.U., T.F.M. and J.P. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Evaluation and recovery of endogenous SPA-tagged E. coli cell envelope proteins (CEPs).
(a) Detergent solubilization of SPA-tagged CEPs. Immunoblots showing representative set of inner membrane proteins (IMPs), outer membrane proteins (OMPs), and periplasmic proteins (PEPs) in various indicated detergents. Detergent extraction efficiency was estimated from the immunoblot results by comparing total recovery (anti-FLAG antibody used to detect tagged bait CEPs) of SPA-tagged CEP with (i) and without (w/o; ii) detergent, and the relative band signal intensity was reported in terms of percentile (iii). Triton, C12E8 and DDM highlighted by a dashed rectangular box (i) was deemed to be effective for the CEPs tested in a pilot optimization study. Detailed description of the detergents are shown in Supplementary Table 5 and Supplementary Note 1. (b) Silver-stained SDS polyacrylamide gels (i) portraying bacterial proteins identified by mass spectrometry (MS) after solubilization and affinity purification in the presence of detergents indicated for a TonB energy transducing (ExbB) SPA-tagged CEP. Distinct gel bands representing putative polypeptide subunits of the purified ExbB protein were analyzed by in-gel trypsin digestion followed by MALDI-TOF MS; protein identities are indicated with arrows. Asterisk indicates tagged CEP bait recovery (as detected by MALDI-TOF MS) in the presence of 3 different detergents is shown along with a control, emphasizing that the ExbB bait was not recovered in the absence of detergent. Known interactions of ExbB, including ExbD, FecA, FepE and FhuA that were captured in at least one of the 3 detergents are shown. Additional previously known ExbB interactions missed by MALDI-TOF MS but identified in our LC-MS/MS (ii) are shown with their corresponding spectral count (SPC) from SEQUEST and MS ID probability score (%) from STATQUEST algorithms. (c) Immunoprecipitations (IP) on detergent-solubilized extracts isolated from the indicated SPA-tagged CEPs. Anti-FLAG antibody was used to detect the tagged bait CEPs. Untagged or wild-type (WT) DY330 parental strain (1,2) served as negative control; wash, protein extract not bound to the beads after 3rd washing with and w/o the detergent; and EB, protein bound to the beads after elution. Asterisk indicates the bait recovery with strong signal intensity in the presence of DDM solubilized extracts. Molecular masses (kDa) of the marker (M) proteins are indicated. (d) Tagged CEPs (annotated and orphan) vs all E. coli soluble proteins (SPs) showing links to drug sensitivity from the large-scale phenomics screen (Nichols et al., 2011).
Supplementary Figure 2 Data quality and cell envelope proteins (CEPs) recovery (coverage).
(a) Affinity-purified 290 SPA-tagged CEPs (shown in % out of 785 baits tested) from multiple biological replicates (right) of late log-phase E. coli cultures grown in rich media, and 495 baits that were subjected to AP/MS once (left) in the presence of 1 to 3 different detergents. (b-e) Number of tagged and successfully purified CEPs (based on bait recovery from MS) is plotted per compartment (log10 scale; b), as well as against protein abundance (expressed in log10 ppm; c), transmembrane α-helix (TMH; d-i) or ß-barrel (d-ii) and Pfam domain (e). IMPs, inner membrane proteins; OMPs, outer membrane proteins; IM, inner membrane; OM, outer membrane; PE, periplasm; LPI, inner lipoprotein; LPO, outer lipoprotein; EC, extracellular; MR, membrane-related.
Supplementary Figure 3 Assessing quality and benchmarking cell envelope protein-protein interactions (cePPIs) in the network.
(a) Proportion of PPIs from each bait CEP filtered at different peptide identification probability score cut-off from SEQUEST/STATQUEST algorithm is plotted against the known interactions curated in EcoCyc. (b) Coverage and accuracy in benchmarking cePPI against reference EcoCyc PPI data set; p-value (ΣLLS vs other scores) computed by Mann-Whitney test and corrected for multiple hypotheses testing using Benjamin and Hochberg. (c) Fraction of interaction partners from E. coli CEPs vs non-CEPs; p-value by Kolmogorov-Smirnov test. (d) Average gene ontology (GO) semantic similarity of the putatively interacting CEPs from this study (CE network) vs other published studies. P-value significance (computed by Student's t-test) between CE network vs other studies for biological processes or cellular component GO annotations are shown with asterisk. (e) Box plots showing the Pearson correlation coefficients of the genetic interaction (GI) profiles for pairs of cePPIs compiled from the indicated E. coli studies vs randomly (Rand) drawn protein pairs; p-value by Student's t-test. (f) Physically interacting CEPs hypersensitive to drugs targeting the CE or different processes; p-value by Student's t-test. (g) Assessment of performance measures by comparing the area-under-the curve (AUC) for interactions that are deemed high confidence (HC) and/or medium confidence (MC). “All” refers to HC+MC cePPIs, as well as from previously reported large-scale (Butland et al., 2005; Hu et al., 2009, and Rajagopala et al., 2014) E. coli studies (using 5-fold cross validation), in benchmarking against cePPIs from curated EcoCyc protein complexes.
Supplementary Figure 4 Biochemical fractionation coupled to mass spectrometry (BF/MS) strategy to identify E. coli cePPIs.
(a) Schematic showing the identification of PPIs from BF/MS and selection criteria used to filter the dataset to verify the reliability of the predicted cePPIs from AP/MS. (b) Detergent solubilized CEPs extracted from E. coli using 0.020% Triton and, separately, 0.05% DDM (below the critical micelle concentration for each detergent) were fractionated into 84 biochemical fractions using size-exclusion chromatography (SEC-HPLC), proteolytically digested and the resulting peptides analyzed, in duplicate, by precision tandem MS. Hierarchical clustering of co-eluting proteins from SEC-HPLC identified by LC-MS/MS. Shading indicates spectral counts (SPC) recorded by LC-MS/MS. (c) Performance (true positive rate, TPR, vs false positive rate, FPR) of cePPI scoring algorithms (co-apex; weighted cross-correlation, WCC; Pearson correlation coefficient, PCC) that calculates correlation measures between all possible pairs of proteins to capture their tendency to co-elute was used to select a threshold cut-off for each score based on capture of curated cePPIs derived from EcoCyc (area-under-the curve; AUC). (d) Pearson correlation (r = 0.8) profiles between Triton (red) and DDM (blue) detergent solubilized proteins (extracted from cultured E. coli membranes) identified by tandem MS is plotted using normalized spectral counts. (e) Independent testing of cePPI pairs by (i) yeast (Y2H) or bacterial (B2H) two-hybrid assays, and (ii) B2H-based β-galactosidase activity (miller units represented as mean ± SD from at least three biological replicates); dashed line indicates background signal, p-value according to Student's t-test.
Supplementary Figure 5 Properties of cell envelope protein (CEP) interaction network.
(a) Number of essential CEPs and/or those targeted by drugs are annotated to various subcellular compartments. (b) Degree connectivity of essential vs non-essential CEPs; p-value by Student's t-test. (c) Fraction of CEPs conserved to various degrees across 412 proteobacterial species. (d) Mean degree distribution of CEPs annotated to various subcellular compartments. (e) Co-localization (i) and distribution (ii) of CEP interacting partners within the same or different sub-cellular compartments; p-value by Student's t-test. IM, inner membrane; OM, outer membrane; PE, periplasm; MR, membrane-related; LPI, inner lipoprotein; LPO, outer lipoprotein; EC, extracellular space; CY, cytoplasm.
Supplementary Figure 6 Assessment of multi-protein complexes (MPCs) predicted via core-attachment algorithm.
(a) Size distribution of the predicted MPCs. (b) Number of predicted complexes containing a CEP that overlap with an EcoCyc complex. Each MPC is binned according to the maximum overlap of the predicted complex with an EcoCyc complex. (c) Binary interactions tested by yeast two-hybrid (Y2H) and/or bacterial two-hybrid (B2H) assays (bridged by one or more intermediates in the protein complexes derived from AP/MS experiments. We show mapping of 88 binary PPIs (purple dotted edges) tested to 103 putative MPCs, where 26 PPIs bridged within 25 complexes, and 75 between 85 complexes.
Supplementary Figure 7 Membrane-or transport-associated metabolons.
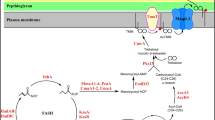
(a) Allosteric activation of PykF by HPr. Steady-state kinetics of PykF was determined as a function of the phosphoenolpyruvate (PEP; 0.5 to 8 mM) in the absence (circles; i) and presence (triangles; i) of 1uM HPr. The resultant kinetic parameters are presented in Table 1 (ii). The effect of varying concentrations of dephospho-HPr (triangles; iii) on PykF activity, P-HPr (circles) using a concentration of PEP 0.4 mM (1.5 mM). (b) Overview of PPIs observed within the indicated ABC transport-associated metabolon system. Each transport metabolon system (a-k) is designated with a three letter designation, followed by preferred substrate in parentheses. The two vertical parallel lines separating the inner membrane (IM) with cytoplasm (CY) or periplasm (PE) is shown schematically. The letter presented in each subcellular compartment indicates the last letter of the protein subunit name. For example, ArtM, ArtP and ArtQ (panel a) are indicated as M, P and Q, respectively; and the letter 'S' in PE denotes binding solute. In panel j, MFP refers to the membrane fusion protein, and in panel k, only the presence of core ABC exporter is shown. The ΣLLS score (≥ 5.27) between two interconnected subunits is represented in each rectangle. (c) Interactions among the housekeeping dipeptide permease (Dpp) subunits. (d) Interactions within and between the two symmetrical Zinc (ZinT, ZnuABC) and the arabinose (AraFGH) ABC transporter systems. (e) Polycistronic bacterial operons encompassing transport proteins and enzymes that metabolize the imported substrates. Structures are shown for potential GlpF (PDB ID: 1LDA) transporter and GlpX (PDB ID: 1NI9) enzyme complexes, while for others no structural information available.
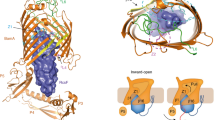
Supplementary Figure 8 TamA/BamA structural homology and TamB production of mature/functional Ag43 on the cell surface.
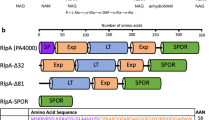
(a) Crystal structures of the insertase domains of BamA (pdb entry 4C4V, cyan) and TamA (pdb entry 4C00, green) solved from E. coli. The extracellular loop 6 is drawn in magenta, and the conserved VRGY motif required for the function of the translocases from Omp85 family is shown in stick representation. BamA and TamA belong to the Omp85 family and share a similar fold mechanism. Their β-barrel domains reveal an unusual cracked-barrel architecture created by the lack of H-bonds between strands 1 and 16 that are loosely associated to create access from the barrel lumen to the hydrophobic lipid phase. As well, their cracked-barrel architecture is suggested to promote a lateral opening of the β-barrel to insert OMP within the membrane (Noinaj et al., Nature, 501, 385-390, 2013). (b) (i) Transmission-electron micrograph showing immunogold-labelling of His-tagged Ag43 autotransporters on the surface (dots are demarcated in arrows; zoom-in of the dot shown in top right panel) of wild-type (WT), ΔtamA and ΔtamB E. coli cells previously transformed with an empty or Ag43 expressing plasmids; scale bar, 100 nm. (ii) Full electron micrographs of the indicated strains observed at a magnification of 40,000 x 1.5; scale bar, 300 nm. (c, d) Release of α-Ag43 fragment from the bacterial cell surface under thermal denaturation. The supernatant preparations from heat-treated bacterium (incubated at 25 or 55 °C for 5 min) were loaded onto SDS-PAGE (c) and analyzed using MALDI-TOF mass spectrometry (observed masses listed in d; n.d. for not detected). (e) Mass determination of the heat-released Ag43 passenger domain from the bacterial surface. Supernatant fractions following a 5 min incubation at 55 °C of WT (top and middle) and ΔtamB (bottom) E. coli cells analyzed using MALDI-TOF to detect Ag43 α-domain release from the bacterial cell surface under thermal denaturation.
Supplementary Figure 9 Multi-protein complex (MPC) conservation.
(a) Phylogenetic tree (i) based on the comparison of the complete proteomes of 20 different bacteria that were closely related to E. coli. Evolutionary conservation of PPIs (ii) based on the co-occurrence of CEP (or those that are essential) orthologs across diverse bacterial species; bars scaled relative to actual species CEP counts; *p-value by Student's t-test. (b) Fraction of conserved subunits of the 540 predicted MPCs across diverse classes of bacteria, with orthologs of E. coli subunits of the sulfonate-sulfur utilization (black dashed line in the heatmap) complex (ID 107) shown as an example.
Supplementary Figure 10 Overlap of cePPIs between paralogs.
(a) Distribution of the fraction of shared interaction partners between paralog (blue) vs random (red) pairs. (b) Average connectivity of paralogs with shared interaction partners vs those without shared partners; p-value by Student's t-test. (c, d) Scatterplot showing the shared interaction partners between paralogs based on the sequence similarity (c) and normalized fold difference of protein abundance (d). (e) Sub-networks showing the shared (dotted line) or unique (thick line) interaction partners with paralogs of the (i) methyl-accepting chemotaxis receptors, (ii) RND-family pumps, as well as (iii) OM capsule polysaccharide exporter and inner membrane (IM) tyrosine protein kinases.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 (PDF 2287 kb)
Supplementary Table 1
Classification of the E. coli proteome and target cell envelope protein selection for AP/MS screens (XLSX 1320 kb)
Supplementary Table 2
Co-purifying protein pairs compiled from reference Ecoyc database and from this study (XLSX 3336 kb)
Supplementary Table 3
The physiological relevance and quality of PPIs by anecdotal supporting evidence, drug sensitivity profiles, and two-hybrid screens (XLSX 632 kb)
Supplementary Table 4
Putative (or novel) CEP complexes and their subunits identified by the core-attachment based clustering algorithm are indicated with their respective topology model (XLSX 451 kb)
Supplementary Table 5
Antibiotic susceptibility, evolutionary conservation, and paralogous analyses on CEPs or complexes, and bacterial strains/plasmids used in this study (XLSX 4185 kb)
Rights and permissions
About this article
Cite this article
Babu, M., Bundalovic-Torma, C., Calmettes, C. et al. Global landscape of cell envelope protein complexes in Escherichia coli. Nat Biotechnol 36, 103–112 (2018). https://doi.org/10.1038/nbt.4024
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.4024
This article is cited by
-
KdpD is a tandem serine histidine kinase that controls K+ pump KdpFABC transcriptionally and post-translationally
Nature Communications (2024)
-
Enhancing coevolutionary signals in protein–protein interaction prediction through clade-wise alignment integration
Scientific Reports (2024)
-
E. coli allantoinase is activated by the downstream metabolic enzyme, glycerate kinase, and stabilizes the putative allantoin transporter by direct binding
Scientific Reports (2023)
-
Co-fractionation–mass spectrometry to characterize native mitochondrial protein assemblies in mammalian neurons and brain
Nature Protocols (2023)
-
Treatment of wastewater for reuse using advanced oxidation process: a bacterial inactivation mechanism approach
International Journal of Environmental Science and Technology (2023)