Abstract
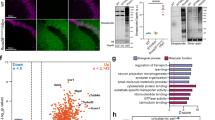
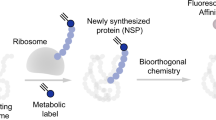
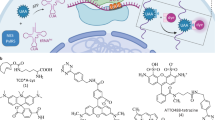
Although advances in protein labeling methods have made it possible to measure the proteome of mixed cell populations, it has not been possible to isolate cell-type-specific proteomes in vivo. This is because the existing methods for metabolic protein labeling in vivo access all cell types. We report the development of a transgenic mouse line where Cre-recombinase-induced expression of a mutant methionyl-tRNA synthetase (L274G) enables the cell-type-specific labeling of nascent proteins with a non-canonical amino-acid and click chemistry. Using immunoblotting, imaging and mass spectrometry, we use our transgenic mouse to label and analyze proteins in excitatory principal neurons and Purkinje neurons in vitro (brain slices) and in vivo. We discover more than 200 proteins that are differentially regulated in hippocampal excitatory neurons by exposing mice to an environment with enriched sensory cues. Our approach can be used to isolate, analyze and quantitate cell-type-specific proteomes and their dynamics in healthy and diseased tissues.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Laughlin, S.T., Baskin, J.M., Amacher, S.L. & Bertozzi, C.R. In vivo imaging of membrane-associated glycans in developing zebrafish. Science 320, 664–667 (2008).
Dieterich, D.C., Link, A.J., Graumann, J., Tirrell, D.A. & Schuman, E.M. Selective identification of newly synthesized proteins in mammalian cells using bioorthogonal noncanonical amino acid tagging (BONCAT). Proc. Natl. Acad. Sci. USA 103, 9482–9487 (2006).
Elsässer, S.J., Ernst, R.J., Walker, O.S. & Chin, J.W. Genetic code expansion in stable cell lines enables encoded chromatin modification. Nat. Methods 13, 158–164 (2016).
Mahdavi, A. et al. Engineered aminoacyl-tRNA synthetase for cell-selective analysis of mammalian protein synthesis. J. Am. Chem. Soc. 138, 4278–4281 (2016).
Ngo, J.T. et al. Cell-selective metabolic labeling of proteins. Nat. Chem. Biol. 5, 715–717 (2009).
Erdmann, I. et al. Cell-selective labelling of proteomes in Drosophila melanogaster. Nat. Commun. 6, 7521 (2015).
Yuet, K.P. et al. Cell-specific proteomic analysis in Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 112, 2705–2710 (2015).
Link, A.J. et al. Discovery of aminoacyl-tRNA synthetase activity through cell-surface display of noncanonical amino acids. Proc. Natl. Acad. Sci. USA 103, 10180–10185 (2006).
Miyazaki, J. et al. Expression vector system based on the chicken beta-actin promoter directs efficient production of interleukin-5. Gene 79, 269–277 (1989).
Zambrowicz, B.P. et al. Disruption of overlapping transcripts in the ROSA beta geo 26 gene trap strain leads to widespread expression of beta-galactosidase in mouse embryos and hematopoietic cells. Proc. Natl. Acad. Sci. USA 94, 3789–3794 (1997).
Tsien, J.Z. et al. Subregion- and cell type-restricted gene knockout in mouse brain. Cell 87, 1317–1326 (1996).
Taniguchi, H. et al. A resource of Cre driver lines for genetic targeting of GABAergic neurons in cerebral cortex. Neuron 71, 995–1013 (2011).
Dieterich, D.C. et al. In situ visualization and dynamics of newly synthesized proteins in rat hippocampal neurons. Nat. Neurosci. 13, 897–905 (2010).
tom Dieck, S. et al. Direct visualization of newly synthesized target proteins in situ. Nat. Methods 12, 411–414 (2015).
Howarth, C., Peppiatt-Wildman, C.M. & Attwell, D. The energy use associated with neural computation in the cerebellum. J. Cereb. Blood Flow Metab. 30, 403–414 (2010).
Cho, K.O., Hunt, C.A. & Kennedy, M.B. The rat brain postsynaptic density fraction contains a homolog of the Drosophila discs-large tumor suppressor protein. Neuron 9, 929–942 (1992).
Innocenti, M. et al. Abi1 is essential for the formation and activation of a WAVE2 signalling complex. Nat. Cell Biol. 6, 319–327 (2004).
Dateki, M. et al. Neurochondrin negatively regulates CaMKII phosphorylation, and nervous system-specific gene disruption results in epileptic seizure. J. Biol. Chem. 280, 20503–20508 (2005).
Han, K. et al. Fragile X-like behaviors and abnormal cortical dendritic spines in cytoplasmic FMR1-interacting protein 2-mutant mice. Hum. Mol. Genet. 24, 1813–1823 (2015).
Weedon, M.N. et al. Exome sequencing identifies a DYNC1H1 mutation in a large pedigree with dominant axonal Charcot-Marie-Tooth disease. Am. J. Hum. Genet. 89, 308–312 (2011).
Novak, M.J. et al. An ITPR1 gene deletion causes spinocerebellar ataxia 15/16: a genetic, clinical and radiological description. Mov. Disord. 25, 2176–2182 (2010).
Bennett, E.L., Diamond, M.C., Krech, D. & Rosenzweig, M.R. Chemical and anatomical plasticity brain. Science 146, 610–619 (1964).
Shih, Y.T. & Hsueh, Y.P. VCP and ATL1 regulate endoplasmic reticulum and protein synthesis for dendritic spine formation. Nat. Commun. 7, 11020 (2016).
Ong, S.E. et al. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol. Cell. Proteomics 1, 376–386 (2002).
Monetti, M., Nagaraj, N., Sharma, K. & Mann, M. Large-scale phosphosite quantification in tissues by a spike-in SILAC method. Nat. Methods 8, 655–658 (2011).
Eichelbaum, K., Winter, M., Berriel Diaz, M., Herzig, S. & Krijgsveld, J. Selective enrichment of newly synthesized proteins for quantitative secretome analysis. Nat. Biotechnol. 30, 984–990 (2012).
Liu, R. et al. Quantitative non-canonical amino acid tagging (QuaNCAT) proteomics identifies distinct patterns of protein synthesis rapidly induced by hypertrophic agents in cardiomyocytes, revealing new aspects of metabolic remodeling. Mol. Cell. Proteomics 15, 3170–3189 (2016).
McShane, E. et al. Kinetic analysis of protein stability reveals age-dependent degradation. Cell 167, 803–815 (2016).
Alvarez-Castelao, B. & Schuman, E.M. The regulation of synaptic protein turnover. J. Biol. Chem. 290, 28623–28630 (2015).
McNair, K., Broad, J., Riedel, G., Davies, C.H. & Cobb, S.R. Global changes in the hippocampal proteome following exposure to an enriched environment. Neuroscience 145, 413–422 (2007).
Moser, E.I., Kropff, E. & Moser, M.B. Place cells, grid cells, and the brain's spatial representation system. Annu. Rev. Neurosci. 31, 69–89 (2008).
O'Keefe, J. & Dostrovsky, J. The hippocampus as a spatial map. Preliminary evidence from unit activity in the freely-moving rat. Brain Res. 34, 171–175 (1971).
Achim, K. et al. High-throughput spatial mapping of single-cell RNA-seq data to tissue of origin. Nat. Biotechnol. 33, 503–509 (2015).
Cembrowski, M.S., Wang, L., Sugino, K., Shields, B.C. & Spruston, N. Hipposeq: a comprehensive RNA-seq database of gene expression in hippocampal principal neurons. eLife 5, e14997 (2016).
de Felipe, P. et al. E unum pluribus: multiple proteins from a self-processing polyprotein. Trends Biotechnol. 24, 68–75 (2006).
Stoppini, L., Buchs, P.A. & Muller, D. A simple method for organotypic cultures of nervous tissue. J. Neurosci. Methods 37, 173–182 (1991).
Branda, C.S. & Dymecki, S.M. Talking about a revolution: The impact of site-specific recombinases on genetic analyses in mice. Dev. Cell 6, 7–28 (2004).
Bachmanov, A.A., Tordoff, M.G. & Beauchamp, G.K. Sweetener preference of C57BL/6ByJ and 129P3/J mice. Chem. Senses 26, 905–913 (2001).
Dieterich, D.C. et al. Labeling, detection and identification of newly synthesized proteomes with bioorthogonal non-canonical amino-acid tagging. Nat. Protoc. 2, 532–540 (2007).
Szychowski, J. et al. Cleavable biotin probes for labeling of biomolecules via azide-alkyne cycloaddition. J. Am. Chem. Soc. 132, 18351–18360 (2010).
Wis´niewski, J.R., Zougman, A., Nagaraj, N. & Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 6, 359–362 (2009).
Schanzenbächer, C.T., Sambandan, S., Langer, J.D. & Schuman, E.M. Nascent proteome remodeling following homeostatic scaling at hippocampal synapses. Neuron 92, 358–371 (2016).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Vizcaíno, J.A. et al. ProteomeXchange provides globally coordinated proteomics data submission and dissemination. Nat. Biotechnol. 32, 223–226 (2014).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Sutton, M.A. et al. Miniature neurotransmission stabilizes synaptic function via tonic suppression of local dendritic protein synthesis. Cell 125, 785–799 (2006).
Eden, E., Navon, R., Steinfeld, I., Lipson, D. & Yakhini, Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics 10, 48 (2009).
Sharma, K. et al. Cell type- and brain region-resolved mouse brain proteome. Nat. Neurosci. 18, 1819–1831 (2015).
Zeisel, A. et al. Brain structure. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015).
Kirsch, L., Liscovitch, N. & Chechik, G. Localizing genes to cerebellar layers by classifying ISH images. PLoS Comput. Biol. 8, e1002790 (2012).
Acknowledgements
We thank H. Geptin, D. Vogel, N. Fuerst, I. Wüllenweber and F. Rupprecht for excellent technical assistance. We thank E. Noll for the synthesis of ANL and P. Landgraf for the synthesis of the SH-alkyne. We thank S. Garg for her help with FUNCAT and M. Heumueller for his help with some of the experiments. We thank R. Pieaud and S. Junek for their assistance with imaging. We thank F. Kretschmer for his help with the open field analysis. We thank E. Northrup, S. Zeissler, S. Gil Mast and the animal facility of MPI for Brain Research for their excellent support. We thank J. Sanes and J. Chakkalakal for the early generation of a Thy-1 MetRS* mouse. Work in the laboratory of E.M.S. is supported by the Max Planck Society, the European Research Council, DFG CRC 902 and 1080, and the DFG Cluster of Excellence for Macromolecular Complexes. B.A. was supported by a Marie Curie Intra-European Fellowship for career development. C.H. was supported by a Marie Curie Career Integration Grant. D.C.D. is supported by DFG CRC 779 and 854. Research on proteomic labelling at Caltech is supported by the Programmable Molecular Technology Initiative of the Gordon and Betty Moore Foundation.
Author information
Authors and Affiliations
Contributions
B.A.-C., C.T.S. and C.H.: conception and design of experiments, acquisition, analysis and interpretation of data. C.G., S.t.D. and A.R.D.: conception and design of experiments, acquisition of data. I.B., B.N.-A. and E.C.: acquisition of data. A.M.: provided reagents. D.D.: provided reagents and advice on experiments. D.A.T.: conception and design of experiments. J.D.L.: acquisition of data, analysis and interpretation of data. E.M.S.: conception and design of experiments, analysis and interpretation of data, drafting, writing and revising the article. All authors contributed to the writing and revision of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 (PDF 80295 kb)
Supplementary Table 1
CaMK2a (standard cage) excitatory hippocampal proteome. (XLSX 1098 kb)
Supplementary Table 2
Gene ontology: CaMK2a (standard cage) excitatoryhippocampal proteome. (XLSX 40 kb)
Supplementary Table 3
Cell-type specific protein and gene markers (XLSX 265 kb)
Supplementary Table 4
GAD2 proteome. (XLSX 764 kb)
Supplementary Table 5
Gene ontology: GAD2 Purkinje neuron proteome. (XLSX 360 kb)
Supplementary Table 6
Differentially expressed genes in CaMK2a (standard cage) vs GAD2 proteomes. (XLSX 289 kb)
Supplementary Table 7
CaMK2a enriched environment excitatory hippocampalproteome. (XLSX 1068 kb)
Supplementary Table 8
Differentially expressed genes in CaMK2a-standard cage vs CaMK2a- enriched environment proteomes. (XLSX 128 kb)
Supplementary Table 9
Gene ontology of differentially expressed genes inCaMK2a-standard cage vs CaMK2a-enriched environment proteomes. (XLSX 38 kb)
Supplementary Table 10
Primers used for mouse genotyping. (DOCX 59 kb)
Supplementary Table 11
Custom parameters used for MS analysis in MaxQuantand Perseus. (XLSX 20 kb)
Supplementary Table 12
Full hippocampus, cerebellum and glia proteomes. (XLSX 2312 kb)
Rights and permissions
About this article
Cite this article
Alvarez-Castelao, B., Schanzenbächer, C., Hanus, C. et al. Cell-type-specific metabolic labeling of nascent proteomes in vivo. Nat Biotechnol 35, 1196–1201 (2017). https://doi.org/10.1038/nbt.4016
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.4016
This article is cited by
-
Spatial multi-omics: novel tools to study the complexity of cardiovascular diseases
Genome Medicine (2024)
-
Native-state proteomics of Parvalbumin interneurons identifies unique molecular signatures and vulnerabilities to early Alzheimer’s pathology
Nature Communications (2024)
-
A mutant methionyl-tRNA synthetase-based toolkit to assess induced-mesenchymal stromal cell secretome in mixed-culture disease models
Stem Cell Research & Therapy (2023)
-
FEAST: A flow cytometry-based toolkit for interrogating microglial engulfment of synaptic and myelin proteins
Nature Communications (2023)
-
Ex vivo immunocapture and functional characterization of cell-type-specific mitochondria using MitoTag mice
Nature Protocols (2023)