Abstract
Biological systems can generate microstructured materials that combine organic and inorganic components and possess diverse physical and chemical properties. However, these natural processes in materials fabrication are not readily programmable. Here, we use a synthetic-biology approach to assemble patterned materials. We demonstrate programmable fabrication of three-dimensional (3D) materials by printing engineered self-patterning bacteria on permeable membranes that serve as a structural scaffold. Application of gold nanoparticles to the colonies creates hybrid organic-inorganic dome structures. The dynamics of the dome structures' response to pressure is determined by their geometry (colony size, dome height, and pattern), which is easily modified by varying the properties of the membrane (e.g., pore size and hydrophobicity). We generate resettable pressure sensors that process signals in response to varying pressure intensity and duration.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Currey, J.D. Mechanical-properties of mother of pearl in tension. Proc. R. Soc. Lond. B. 196, 443–463 (1977).
Luz, G.M. & Mano, J.F. Mineralized structures in nature: Examples and inspirations for the design of new composite materials and biomaterials. Compos. Sci. Technol. 70, 1777–1788 (2010).
Jackson, A.P., Vincent, J.F.V. & Turner, R.M. The mechanical design of nacre. Proc. R. Soc. Lond. B. 234, 415–440 (1988).
Chen, A.Y., Zhong, C. & Lu, T.K. Engineering living functional materials. ACS Synth. Biol. 4, 8–11 (2015).
Purnick, P.E. & Weiss, R. The second wave of synthetic biology: from modules to systems. Nat. Rev. Mol. Cell Biol. 10, 410–422 (2009).
Khalil, A.S. & Collins, J.J. Synthetic biology: applications come of age. Nat. Rev. Genet. 11, 367–379 (2010).
Ringler, P. & Schulz, G.E. Self-assembly of proteins into designed networks. Science 302, 106–109 (2003).
Zhong, C. et al. Strong underwater adhesives made by self-assembling multi-protein nanofibres. Nat. Nanotechnol. 9, 858–866 (2014).
Ryadnov, M.G. & Woolfson, D.N. Engineering the morphology of a self-assembling protein fibre. Nat. Mater. 2, 329–332 (2003).
Lee, Y.J. et al. Fabricating genetically engineered high-power lithium-ion batteries using multiple virus genes. Science 324, 1051–1055 (2009).
Aggeli, A. et al. pH as a trigger of peptide beta-sheet self-assembly and reversible switching between nematic and isotropic phases. J. Am. Chem. Soc. 125, 9619–9628 (2003).
Fichman, G. & Gazit, E. Self-assembly of short peptides to form hydrogels: design of building blocks, physical properties and technological applications. Acta Biomater. 10, 1671–1682 (2014).
Yan, H., Park, S.H., Finkelstein, G., Reif, J.H. & LaBean, T.H. DNA-templated self-assembly of protein arrays and highly conductive nanowires. Science 301, 1882–1884 (2003).
Seeman, N.C. & Belcher, A.M. Emulating biology: building nanostructures from the bottom up. Proc. Natl. Acad. Sci. USA 99 (Suppl. 2), 6451–6455 (2002).
Seeman, N.C. Nanomaterials based on DNA. Annu. Rev. Biochem. 79, 65–87 (2010).
Elbaz, J., Yin, P. & Voigt, C.A. Genetic encoding of DNA nanostructures and their self-assembly in living bacteria. Nat. Commun. 7, 11179 (2016).
Sleytr, U.B. & Beveridge, T.J. Bacterial S-layers. Trends Microbiol. 7, 253–260 (1999).
Feng, Y.Y. et al. Hybrid (organic/inorganic) electrodes from bacterially precipitated CdS for PEC/storage applications. J. Phys. Chem. C 121, 3734–3743 (2017).
Shenton, W., Pum, D., Sleytr, U.B. & Mann, S. Synthesis of cadmium sulphide superlattices using self-assembled bacterial S-layers. Nature 389, 585–587 (1997).
Marusak, K.E. et al. Cadmium sulphide quantum dots with tunable electronic properties by bacterial precipitation. RSC Advances 6, 76158–76166 (2016).
Chen, A.Y. et al. Synthesis and patterning of tunable multiscale materials with engineered cells. Nat. Mater. 13, 515–523 (2014).
Barnhart, M.M. & Chapman, M.R. Curli biogenesis and function. Annu. Rev. Microbiol. 60, 131–147 (2006).
Payne, S. et al. Temporal control of self-organized pattern formation without morphogen gradients in bacteria. Mol. Syst. Biol. 9, 697 (2013).
Tan, C., Marguet, P. & You, L. Emergent bistability by a growth-modulating positive feedback circuit. Nat. Chem. Biol. 5, 842–848 (2009).
Stano, N.M. & Patel, S.S. T7 lysozyme represses T7 RNA polymerase transcription by destabilizing the open complex during initiation. J. Biol. Chem. 279, 16136–16143 (2004).
Cao, Y. et al. Collective space-sensing coordinates pattern scaling in engineered bacteria. Cell 165, 620–630 (2016).
Zhang, R., Xu, Y., Wen, B., Sheng, N. & Fang, H. Enhanced permeation of a hydrophobic fluid through particles with hydrophobic and hydrophilic patterned surfaces. Sci. Rep. 4, 5738 (2014).
Nagapudi, K. et al. Viscoelastic and mechanical behavior of recombinant protein elastomers. Biomaterials 26, 4695–4706 (2005).
Tuson, H.H. et al. Measuring the stiffness of bacterial cells from growth rates in hydrogels of tunable elasticity. Mol. Microbiol. 84, 874–891 (2012).
Basu, S., Gerchman, Y., Collins, C.H., Arnold, F.H. & Weiss, R. A synthetic multicellular system for programmed pattern formation. Nature 434, 1130–1134 (2005).
Liu, C. et al. Sequential establishment of stripe patterns in an expanding cell population. Science 334, 238–241 (2011).
Tabor, J.J. et al. A synthetic genetic edge detection program. Cell 137, 1272–1281 (2009).
Schaerli, Y. et al. A unified design space of synthetic stripe-forming networks. Nat. Commun. 5, 4905 (2014).
Song, H., Payne, S., Gray, M. & You, L. Spatiotemporal modulation of biodiversity in a synthetic chemical-mediated ecosystem. Nat. Chem. Biol. 5, 929–935 (2009).
Moon, T.S., Lou, C., Tamsir, A., Stanton, B.C. & Voigt, C.A. Genetic programs constructed from layered logic gates in single cells. Nature 491, 249–253 (2012).
Fernandez-Rodriguez, J., Yang, L., Gorochowski, T.E., Gordon, D.B. & Voigt, C.A. Memory and combinatorial logic based on DNA inversions: dynamics and evolutionary stability. ACS Synth. Biol. 4, 1361–1372 (2015).
Andrianantoandro, E., Basu, S., Karig, D.K. & Weiss, R. Synthetic biology: new engineering rules for an emerging discipline. Mol. Syst. Biol. 2, 0028 (2006).
Siuti, P., Yazbek, J. & Lu, T.K. Synthetic circuits integrating logic and memory in living cells. Nat. Biotechnol. 31, 448–452 (2013).
Tamsir, A., Tabor, J.J. & Voigt, C.A. Robust multicellular computing using genetically encoded NOR gates and chemical 'wires'. Nature 469, 212–215 (2011).
Rosenfeld, N., Young, J.W., Alon, U., Swain, P.S. & Elowitz, M.B. Gene regulation at the single-cell level. Science 307, 1962–1965 (2005).
Bennett, M.R. & Hasty, J. Microfluidic devices for measuring gene network dynamics in single cells. Nat. Rev. Genet. 10, 628–638 (2009).
Din, M.O. et al. Synchronized cycles of bacterial lysis for in vivo delivery. Nature 536, 81–85 (2016).
Balagaddé, F.K. et al. A synthetic Escherichia coli predator-prey ecosystem. Mol. Syst. Biol. 4, 187 (2008).
Chen, Y., Kim, J.K., Hirning, A.J., Josić, K. & Bennett, M.R. SYNTHETIC BIOLOGY. Emergent genetic oscillations in a synthetic microbial consortium. Science 349, 986–989 (2015).
Lander, A.D. Pattern, growth, and control. Cell 144, 955–969 (2011).
Nguyen, P.Q., Botyanszki, Z., Tay, P.K. & Joshi, N.S. Programmable biofilm-based materials from engineered curli nanofibres. Nat. Commun. 5, 4945 (2014).
Li, Y. et al. Surface plasmon coupling enhanced dielectric environment sensitivity in a quasi-three-dimensional metallic nanohole array. Opt. Express 18, 3546–3555 (2010).
Grandidier, J., Callahan, D.M., Munday, J.N. & Atwater, H.A. Light absorption enhancement in thin-film solar cells using whispering gallery modes in dielectric nanospheres. Adv. Mater. 23, 1272–1276 (2011).
Han, G.Q. et al. Controllable synthesis of three dimensional electrodeposited Co-P nanosphere arrays as efficient electrocatalysts for overall water splitting. RSC Adv. 6, 52761–52771 (2016).
Sun, F., Zhang, W.B., Mahdavi, A., Arnold, F.H. & Tirrell, D.A. Synthesis of bioactive protein hydrogels by genetically encoded SpyTag-SpyCatcher chemistry. Proc. Natl. Acad. Sci. USA 111, 11269–11274 (2014).
Chen, L. et al. Two-dimensionality of yeast colony expansion accompanied by pattern formation. PLoS Comput. Biol. 10, e1003979 (2014).
Vallet-Regí, M., Colilla, M. & González, B. Medical applications of organic-inorganic hybrid materials within the field of silica-based bioceramics. Chem. Soc. Rev. 40, 596–607 (2011).
Hirst, A.R., Escuder, B., Miravet, J.F. & Smith, D.K. High-tech applications of self-assembling supramolecular nanostructured gel-phase materials: from regenerative medicine to electronic devices. Angew. Chem. Int. Edn Engl. 47, 8002–8018 (2008).
Goldberg, M., Langer, R. & Jia, X. Nanostructured materials for applications in drug delivery and tissue engineering. J. Biomater. Sci. Polym. Ed. 18, 241–268 (2007).
Langer, R. & Tirrell, D.A. Designing materials for biology and medicine. Nature 428, 487–492 (2004).
Niu, Z., Liu, L., Zhang, L. & Chen, X. Porous graphene materials for water remediation. Small 10, 3434–3441 (2014).
Li, H., Liu, L.F. & Yang, F.L. Covalent assembly of 3D graphene/polypyrrole foams for oil spill cleanup. J. Mater. Chem. A Mater. Energy Sustain. 1, 3446–3453 (2013).
Newman, S.A. & Frisch, H.L. Dynamics of skeletal pattern formation in developing chick limb. Science 205, 662–668 (1979).
Jernvall, J. & Thesleff, I. Reiterative signaling and patterning during mammalian tooth morphogenesis. Mech. Dev. 92, 19–29 (2000).
Kavanagh, K.D., Evans, A.R. & Jernvall, J. Predicting evolutionary patterns of mammalian teeth from development. Nature 449, 427–432 (2007).
Davies, D.G. et al. The involvement of cell-to-cell signals in the development of a bacterial biofilm. Science 280, 295–298 (1998).
Asally, M. et al. Localized cell death focuses mechanical forces during 3D patterning in a biofilm. Proc. Natl. Acad. Sci. USA 109, 18891–18896 (2012).
Sambrook, J. & Russell, D.W. Molecular Cloning: a Laboratory Manual 3 edn. (Cold Spring Harbor Laboratory Press, 2001).
Cohen, D.J., Morfino, R.C. & Maharbiz, M.M. A modified consumer inkjet for spatiotemporal control of gene expression. PLoS One 4, e7086 (2009).
Braunovic, M., Konchits, V.V. & Myshkin, N.K. Electrical Contacts: Fundamentals, Applications and Technology (CRC Press, 2007).
Li, L.Q., Song, W.P., Zhang, G.Y. & Jia, D. An electrical contact resistance model including roughness effect for a rough MEMS switch. J. Micromech. Microeng. 22, 115023 (2012).
Vogler, M. & Sheppard, S. Electrical contact resistance under high loads and elevated-temperatures. Weld. J. 72, S231–S238 (1993).
Acknowledgements
We thank R. Tsoi, C. Zhang, Z. Dai for discussions and comments; Y. Gao for assistance with confocal microscopy; Duke Light Microscopy Core Facility (LMCF) for access to confocal microscopes and imaging software; M. Plue for assistance with the TEM and SEM; Duke Shared Materials Instrumentation Facility (SMIF) for access to TEM and SEM. This study was partially supported by the Office of Naval Research (N00014-12-1-0631), National Science Foundation (L.Y.: MCB-1412459; M.D.R.: DMS-1614838), Army Research Office (L.Y., #W911NF-14-1-0490), National Institutes of Health (L.Y.: 1R01-GM098642; K99CA207872-01), Swiss National Science Foundation (M.D.R.: P300P2_154583), a David and Lucile Packard Fellowship (L.Y.).
Author information
Authors and Affiliations
Contributions
L.Y. and Y.C. conceived the project. Y.C. generated and analyzed all the experimental data. Y.C. developed MATLAB codes for image analysis. Y.C. and Y.F. designed and carried out the electrochemical pressure-sensing experiments. M.D.R. and G.H. developed the numerical simulator for 3D pattern formation. Y.C. conducted parameter fittings in all simulations and generated the final simulation results. C.C. developed the finite element simulations for strain analysis. K.Z. assisted with immunolabeling and TEM imaging. K.M. assisted with TEM imaging. Y.C., S.Z., and L.Y. wrote the manuscript, with input from Y.F., M.D.R., K.Z., G.H., and K.M.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Construction of pattern-curli circuit
(a) TEM image of MG1655 ΔcsgA cells after 24 hrs incubation at 30 ˚C. The scale bar is 200nm. (b) TEM image of MG1655 QUOTE csgA cells carrying the curli-pattern circuit incubated after 24 hrs incubation at 30 ˚C. The scale bar is 500nm. (c) Crystal violet (CV) staining assay of MG1655 ΔcsgA cells carrying the curli-pattern circuit and MG1655 ΔcsgA cells carrying inducible curli circuit. CV staining assay is a standard method to stain amyloid1. In this case, the presence of CV stain (blue) indicates the production of curli fibril. Both samples were washed in 1× PBS for three times, then immersed in 500 μL 0.1% CV solution (Sigma) for 15 mins at room temperature. After that, the samples were washed with 1× PBS four times. Left: there was no detectable CV stain in the absence of AHL-mediated CsgA induction. Middle: CV stain was detected when CsgA was induced by 50 nM AHL. Right: CV stain was detected when bacteria carrying the curli-pattern circuit were induced by 1000 μM IPTG. (d) OD600 of CV stain dissolved in acetic acid. The samples in Supplementary Fig. 1C were immersed in 500 μL of 30% acetic acid for 15 mins at room temperature. The ODs of dissolved solutions were measured at 600 nm absorbance using a Perkin-Elmer VICTOR3 plate reader. MG1655 ∆csgA curli-pattern circuit was used in this measurement. Circuit was induced by different amount of IPTG and AHL. The error bars represent the +/- s.d. for n = 5 (AHL concentration varying from 0-10nM) and n=3 (AHL concentration varying from 20-100nM). (e) TEM images of curli secreted from MG1655 ∆csgA carrying the curli-pattern circuit assembling with 5 nm NiNTA-AuNP. Before assembly, cells were incubated at 30 ˚C after 24 hrs. Then 5 nm NiNTA-AuNP were assembled to curli using the protocol described as NiNTA-AuNP labelling. The scale bar is 100nm. Right: TEM image at larger magnification with scale bar of 50 nm. (f) TEM images of curli secreted from MG1655 ∆csgA carrying the curli-pattern circuit binding with gold nanogold using immunolabeling. The 6×-His tag first binds with 1st antibody (mouse anti-6× His tag antibody conjugated with biotin); then 1st antibody binds to 2nd antibody (goat anti-mouse IgG conjugated with 10 nm gold). By this means, 10 nm gold nanogold particles were assembled to the curli. The scale bar is 500nm. Right: TEM image at larger magnification with scale bar of 200nm. (g) TEM images of curli immunolabeling with 2nd antibody without 1st antibody. Antibody agents and constructs were maintained the same with Supplementary Fig. 1F. The scale bar is 500nm. (h) TEM images of curli secreted from MG1655 ∆csgA carrying the curli-pattern circuit binding with quantum dots (Qdots) using immunolabeling. The 6×-His tag first binds with 1st antibody (mouse anti-6× His tag antibody conjugated with biotin); then 1st antibody binds to 2nd antibody (Streptavidin-655Qdots). The scale bar is 200nm. Right: TEM image at larger magnification with scale bar of 100nm.
1 Elghetany, M. T. & Saleem, A. Methods for staining amyloid in tissues: a review. Stain Technol 63, 201-212 (1988).
Supplementary Figure 2 Bacterial growth and pattern formation on permeable membranes.
(a) Printing bacteria on a membrane. An initial overnight LB culture of MG1655 ∆csgA carrying the full gene circuit was grown for 16 hrs at 37 °C. 0.3% molten agar in 2×YT (PH = 6.5) medium was prepared. While allowing it to cool, we diluted the overnight culture to an OD of 0.2, and then diluted the resulting culture another 50-fold into 10 mL fresh LB culture before loading it into the inkjet printer cartridge. After the agar cooled down to 50 °C, IPTG and appropriate antibiotics were added. 170 μL of the agar was added to each culture well. After the agar solidified, a permeable membrane was placed on top of the agar. We then used the inject printer to print the bacteria solution onto the membrane surface. After incubating the whole device under 30 ºC for 32 hrs, the permeable membrane carrying the colony was carefully removed by using a tweezer. (b) Heights and radii of colonies growing on PC membranes with different pore sizes after 32 hrs incubation at 30 °C. The heights and radii were extracted by a custom MATLAB image analysis code from the confocal microscopy2. The contact angles of these membranes were 64.0º, 59.0º, 58.5º, 57.7º, and 55.3º (corresponding to increasing pore sizes). The error bars represent the +/- s.d. for n = 2 (pore size was 0.03 μm) and n=3 (pore size varied from 0.05-0.4 μm). (c) Simulated heights and radii of colonies when the radius expansion rate υ and the fitting constant for nutrient loss by transport α1 were varied according to Eqs (2) and (3), respectively. (d) Heights and radii of colonies growing on membranes with different contact angles after 32 hrs incubation at 30 ºC. The heights and radii were extracted the same way described in Supplementary Fig. 2B. The pore size for all the membranes was 0.45 μm. Contact angles were 1.7º, 38.0º, 63.1º, and 134.3º. The error bars represent the +/- s.d. for n = 2 (contact angle was 38.0º) and n=3 (contact angles were 1.7º, 63.1º, and 134.3º). (e) Simulated heights and radii of colonies when the radius expansion rate υ was varied according to Eq (2).
2 Cao, Y. et al. Collective Space-Sensing Coordinates Pattern Scaling in Engineered Bacteria. Cell 165, 620-630, doi:https://doi.org/10.1016/j.cell.2016.03.006 (2016).
Supplementary Figure 3 Generation of structured materials using bacteria carrying the curli-pattern circuit.
(a) Assembly of nanoparticles to a colony of bacteria carrying curli-pattern circuit. We first fixed the colony, and then used the immunolabeling to bind nanoparticles to the colony. (b) Gold particles distribute similarly as mCherry in colonies. To examine if curli was co-expressed with T7 Lysozyme sufficiently, we compared the mCherry signal with the emission signal from the patterned nanoparticles. Left column: cells carrying curli-pattern circuit were grown on a PC membrane with a 0.4 μm pore size. Gold nanoparticles conjugated with an Alexa Fluor 488 fluorescent dye (a bright, green-fluorescent dye with excitation ideally suited to the 488 nm laser line, Thermo Fisher Scientific) were assembled onto the colony through immunolabeling. The colony was excited at 555 nm (left panel), or 488 nm (middle panel) during confocal microscopy. The mCherry signal (red data points) matched with the emission signal from Fluor 488 dye (green data points) very well: both form dome structures, indicating that gold particles distribute similarly as mCherry within the colony (right panel). The x-axis and y-axis are the same for all three panels. (c) Cells carrying curli-pattern circuit were grown on PC membrane with a 0.03 μm pore size. No gold nanoparticles were assembled onto the colony. When the colony was excited at 488 nm, no emission signal was detected (middle panel). This result demonstrated that the signal observed in the middle panel in Supplementary Fig. 3B was emitted from the Fluor 488 dye, not from the colony itself. Right panel: bacteria carrying the inducible curli circuit were grown on a PC membrane with a 0.03 μm pore size. Gold nanoparticles conjugated with the Alexa Fluor 488 fluorescent dye were assembled on colony through immunolabeling. These bacteria did not express CFP or mCherry reporter. When excited at 488 nm, the emission signal from the Fluor 488 dye formed a solid spherical cap instead of a dome (Fig. 3D). The x-axis and y-axis are the same for all three panels. (d) TEM of bacteria scooped from the outer-layer of the colony (dome part) carrying curli-pattern circuit after immunolabeling. The colony was grown on top of a PC membrane with a 0.4 μm pore size. The TEM image shows 10nm gold nanoparticles bound to curli fibrils. This result shows that the bacteria from the curli dome have efficiently assembled the nanoparticles.
Supplementary Figure 4 Assembly of the pressure sensor.
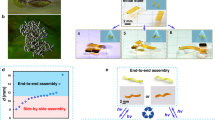
(a) Design concept. Black lines represent conductive wires. In the absence of pressing, the two colonies are separate and there is no electric current passing through. When pressed together lightly, a small current can go through the two colonies via their contact. The current increases with increasing pressure that enhances the contact. (b) Implementation of the sensor. Two PC membranes carrying colonies were mounted to the surfaces of two glass covers with nitrocellulose (Electron Microscopy Science, 72180). Copper wires were used to link the edge of the colony. The two glass covers (each carrying a colony) were placed to face each other with a 0.5 mm spacer (Grace Bio-Labs Press-To-Seal silicone isolator) in between. The other ends of both copper wires were linked to the positive and negative terminals of an electrochemistry workstation. A syringe needle controlled by a programmable syringe pump was used as a “presser” (18ga × 1.5", Blunt Tip) to press the colony sensor. The diameter of the syringe needle was 1.5 mm. It was placed 1 mm right above the top of the glass cover. The top projection of the cylinder aligned with the location of the colony. (c) Configuration of the overall testing device for the pressure sensor. The electrochemical workstation provides a constant voltage, and measures and records the electric currents resulting from the bacterial pressure sensors.
Supplementary Figure 5 Finite element simulation of two domes during pressing.
(a) Simulation configuration. Yellow part indicates the mixture layer of organic matters and gold nanoparticles. Gray part indicates organic matters. The height of all domes (H) is 200 μm. Two dome radii (R= 300 μm or 420 μm) are tested. The thickness of the dome is 1/5 of the dome height. All domes are three-dimensional (right panel). (b) Comparison of the average strain of the two sets of domes at different compressing distance. The strain (y-axis) refers to the average strain along the middle interface of the conducting mixture layer. In particular, each element node in the simulation will have its strain value, then all the strain values were integrally averaged along the line of middle interface of the conducing layer. The average strain along the middle interface of the conducting mixture layer in the smaller colony domes are is ~23.8% larger than that in the larger colony domes. (c) Contour plots of strain distributions in two sets of domes. The compressing distance for both sets of domes is 50 μm. Due to the symmetric configuration, only the domes on the bottom are shown in the figure. The color coding of the heatmap is the same for both sets of domes. The strain in smaller dome are overall higher than that in the larger dome. (d) Comparison of the average contact pressure of the two sets of domes at different compressing distance. The average pressure (y-axis) refers to the average contact pressure at the contact surface area of the domes. The smaller dome demonstrates a larger (~14.5% more) average contact pressure at the contact surface area of the domes. (e) Comparison of the average contact pressure of the two sets of domes at certain compressing distance with different Young’s modulus ratio. The compressing distance for both sets of domes is 45 μm. The conclusion that the smaller dome demonstrates a larger average contact pressure at the contact surface area of the domes maintains for a wide range of variations in the modulus ratio (E2/E1) of the materials. (f) Comparison of the average contact pressure of the two sets of domes at certain compressing distance with different thickness of the mixture layer. The compressing distance for both sets of domes is 45 μm. The conclusion that the smaller dome demonstrates a larger average contact pressure at the contact surface area of the domes maintains for a wide range of variations in the mixed layer’s thickness.
Supplementary Figure 6 Life span and storage condition for bacterial pressure sensor.
(a) Responses of bacterial pressure sensors to repeated pressing with different frequencies Top panel: The bacterial pressure sensors generated 8 strong current signals when the time gap between two pressing pulses is 5s. Bottom panel: The bacterial pressure sensors generated 57 strong current signals when the time gap between two pressing pulses is 20s. The pore size of the membrane is 0.2 μm. The bacterial pressure sensors in both panels were fabricated under the same experimental conditions and using the same protocols. (b) Experimental configuration for testing bacterial pressure sensor’s life span with different storage conditions (e.g. humidity). i. After both devices were pressed for the first time (as in Figure 3A), they were stored in petri dishes containing (left panel) or not containing (right panel) a wet towel for one day at 4 °C. The wet towel served to maintain the humidity in the first petri dish. Then both devices were pressed the same way like Figure 3A. The electric current from both devices was recorded and compared. Then both devices were stored for another day. The iteration continued until both devices failed to respond to pressing. ii. The electric current signal measured right after the bacterial sensors being assembled. Bacterial colonies grew on membranes with pore size of 0.03 μm. iii. The electric current signal measured of the same device in Figure R5B after 1day storage with wet towel at 4 °C.
Supplementary Figure 7 Using a load cell to measure the force loaded on the bacterial sensor.
(a) Experimental configuration. The load cell (Honeywell load cell, model 31) was mounted to an aluminum base (10 cm ×15 cm ×15 mm). The bacterial sensor device was placed right on top of the testing platform. A syringe needle controlled by a programmable syringe pump was used as a “presser” (18ga × 1.5", Blunt Tip) to press the testing platform. It was placed 1 mm right above the top of the testing platform. The top projection of the cylinder aligned with the central axis of the load cell and the colony. (b) Controlled displacement of the presser. The distance indicates the displacement of the presser from its starting position. The presser starts to contact the testing platform when the displacement is >1 mm. (c) Force readout from the load cell. Regardless of the dome sizes (pore size of the membrane is 0.1 μm and 0.2 μm), the forces experienced by the bacterial devices were similar.
Supplementary Figure 8 Multi-colony location sensors.
(a) Design of two-colony, three-colony, and four-colony location sensors. Similar to Supplementary Fig. 4, in each device, colonies and conductive wires were mounted onto two membranes. Mounted to the bottom membrane, the yellow wires connected colonies to the negative terminal of the voltage. Mounted to the top membrane, the red wires connected colonies with to positive terminal of the voltage. To differentiate which colony was pressed, we labelled them a, b, c, and d based on the locations. Note: red and yellow wires are the same type of conductive copper wires. (b) Current responses to pressing at different locations. The top panel indicates the pressure input – the pressing distance as a function of time at different locations. When applicable, the first pulse was applied to location ‘a’, the second pulse to ‘b’, the third pulse to ‘c’, and the forth pulse to ‘d’. The bottom panel indicates the current responses from pressing at different locations. In the same device and at the same strength of pressing, the amplitude of the output current is determined by the location of the pressing. For example, in 2-colony device, the peak current from location ‘b’ is higher than location ‘a’. The pore size of all the membranes used in this set of experiments were 0.1 μm.
Supplementary Figure 9 Construction of 2×2 bacterial touch-pad.
(a) Design of the bacterially assembled sensor array. The schematic illustrates a “touch pad”, that could sense and transduce local pressure variations. The dome shape represents the micro-structured material made from the colony; the orange lines represent conductive wires; and the two blue planes represent supporting surfaces. The current from the bio-fabricated electronic broad will input to a Darlington transistor array (ULN2003ANE4, Texas Instrument). The transistor will amplify the original signal and control the LED dot matrix. (b) Individual LED light up. Individual LED will light up accordingly based on the specific bacterial sensor being pressed. (c) Multiple LEDs light up. Multiple LEDs will light up accordingly based on the combination of bacterial sensors being pressed.
Supplementary information
Supplementary Figures
Supplementary Figures 1–9 (PDF 1458 kb)
Supplementary Tables
Supplementary Table1–2 (PDF 572 kb)
Rights and permissions
About this article
Cite this article
Cao, Y., Feng, Y., Ryser, M. et al. Programmable assembly of pressure sensors using pattern-forming bacteria. Nat Biotechnol 35, 1087–1093 (2017). https://doi.org/10.1038/nbt.3978
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3978
This article is cited by
-
Gold nanoparticles exhibit anti-osteoarthritic effects via modulating interaction of the “microbiota-gut-joint” axis
Journal of Nanobiotechnology (2024)
-
Accelerating the design of pili-enabled living materials using an integrative technological workflow
Nature Chemical Biology (2024)
-
Applications, challenges, and needs for employing synthetic biology beyond the lab
Nature Communications (2021)
-
Living fabrication of functional semi-interpenetrating polymeric materials
Nature Communications (2021)
-
Enhancing the tropism of bacteria via genetically programmed biosensors
Nature Biomedical Engineering (2021)