Abstract
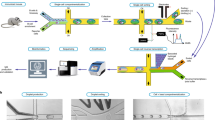
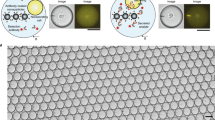
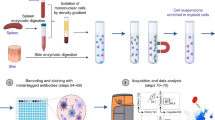
Studies of the dynamics of the antibody-mediated immune response have been hampered by the absence of quantitative, high-throughput systems to analyze individual antibody-secreting cells1,2,3,4,5. Here we describe a simple microfluidic system, DropMap, in which single cells are compartmentalized in tens of thousands of 40-pL droplets and analyzed in two-dimensional droplet arrays using a fluorescence relocation-based immunoassay. Using DropMap, we characterized antibody-secreting cells in mice immunized with tetanus toxoid (TT) over a 7-week protocol, simultaneously analyzing the secretion rate and affinity of IgG from over 0.5 million individual cells enriched from spleen and bone marrow. Immunization resulted in dramatic increases in the range of both single-cell secretion rates and affinities, which spanned at maximum 3 and 4 logs, respectively. We observed differences over time in dynamics of secretion rate and affinity within and between anatomical compartments. This system will not only enable immune monitoring and optimization of immunization and vaccination protocols but also potentiate antibody screening6,7.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Nossal, G.J.V. & Mäkelä, O. Elaboration of antibodies by single cells. Annu. Rev. Microbiol. 16, 53–74 (1962).
Helmreich, E., Kern, M. & Eisen, H.N. The secretion of antibody by isolated lymph node cells. J. Biol. Chem. 236, 464–473 (1961).
Hibi, T. & Dosch, H.M. Limiting dilution analysis of the B cell compartment in human bone marrow. Eur. J. Immunol. 16, 139–145 (1986).
Bromage, E., Stephens, R. & Hassoun, L. The third dimension of ELISPOTs: quantifying antibody secretion from individual plasma cells. J. Immunol. Methods 346, 75–79 (2009).
Salmon, S.E. & Smith, B.A. Immunoglobulin synthesis and total body tumor cell number in IgG multiple myeloma. J. Clin. Invest. 49, 1114–1121 (1970).
El Debs, B., Utharala, R., Balyasnikova, I.V., Griffiths, A.D. & Merten, C.A. Functional single-cell hybridoma screening using droplet-based microfluidics. Proc. Natl. Acad. Sci. USA 109, 11570–11575 (2012).
Mazutis, L. et al. Single-cell analysis and sorting using droplet-based microfluidics. Nat. Protoc. 8, 870–891 (2013).
Radbruch, A. et al. Competence and competition: the challenge of becoming a long-lived plasma cell. Nat. Rev. Immunol. 6, 741–750 (2006).
Eisen, H.N. Affinity enhancement of antibodies: how low-affinity antibodies produced early in immune responses are followed by high-affinity antibodies later and in memory B-cell responses. Cancer Immunol. Res. 2, 381–392 (2014).
Wine, Y. et al. Molecular deconvolution of the monoclonal antibodies that comprise the polyclonal serum response. Proc. Natl. Acad. Sci. USA 110, 2993–2998 (2013).
Lavinder, J.J. et al. Identification and characterization of the constituent human serum antibodies elicited by vaccination. Proc. Natl. Acad. Sci. USA 111, 2259–2264 (2014).
Nossal, G.J.V. & Lederberg, J. Antibody production by single cells. Nature 181, 1419–1420 (1958).
Georgiou, G. et al. The promise and challenge of high-throughput sequencing of the antibody repertoire. Nat. Biotechnol. 32, 158–168 (2014).
Tas, J.M. et al. Visualizing antibody affinity maturation in germinal centers. Science 351, 1048–1054 (2016).
Czerkinsky, C.C., Nilsson, L.A., Nygren, H., Ouchterlony, O. & Tarkowski, A. A solid-phase enzyme-linked immunospot (ELISPOT) assay for enumeration of specific antibody-secreting cells. J. Immunol. Methods 65, 109–121 (1983).
Saletti, G., Çuburu, N., Yang, J.S., Dey, A. & Czerkinsky, C. Enzyme-linked immunospot assays for direct ex vivo measurement of vaccine-induced human humoral immune responses in blood. Nat. Protoc. 8, 1073–1087 (2013).
Clargo, A.M. et al. The rapid generation of recombinant functional monoclonal antibodies from individual, antigen-specific bone marrow-derived plasma cells isolated using a novel fluorescence-based method. MAbs 6, 143–159 (2014).
Love, J.C., Ronan, J.L., Grotenbreg, G.M., van der Veen, A.G. & Ploegh, H.L. A microengraving method for rapid selection of single cells producing antigen-specific antibodies. Nat. Biotechnol. 24, 703–707 (2006).
Jin, A. et al. Rapid isolation of antigen-specific antibody-secreting cells using a chip-based immunospot array. Nat. Protoc. 6, 668–676 (2011).
Köster, S. et al. Drop-based microfluidic devices for encapsulation of single cells. Lab Chip 8, 1110–1115 (2008).
Boitard, L. et al. Monitoring single-cell bioenergetics via the coarsening of emulsion droplets. Proc. Natl. Acad. Sci. USA 109, 7181–7186 (2012).
Anna, S.L., Bontoux, N. & Stone, H.A. Formation of dispersions using “flow focusing” in microchannels. Appl. Phys. Lett. 82, 364–366 (2003).
Foote, J. & Eisen, H.N. Kinetic and affinity limits on antibodies produced during immune responses. Proc. Natl. Acad. Sci. USA 92, 1254–1256 (1995).
Poulsen, T.R., Meijer, P.J., Jensen, A., Nielsen, L.S. & Andersen, P.S. Kinetic, affinity, and diversity limits of human polyclonal antibody responses against tetanus toxoid. J. Immunol. 179, 3841–3850 (2007).
Pilbrough, W., Munro, T.P. & Gray, P. Intraclonal protein expression heterogeneity in recombinant CHO cells. PLoS One 4, e8432 (2009).
Sigal, A. et al. Variability and memory of protein levels in human cells. Nature 444, 643–646 (2006).
Vieira, P. & Rajewsky, K. The half-lives of serum immunoglobulins in adult mice. Eur. J. Immunol. 18, 313–316 (1988).
Yokoyama, W.M. et al. Production of monoclonal antibodies. Curr. Protoc. Immunol. 102, 205 (2013).
Greenfield, E.A. Antibodies: A Laboratory Manual. 2nd edn. (CSH Press, 2014).
Bernasconi, N.L., Traggiai, E. & Lanzavecchia, A. Maintenance of serological memory by polyclonal activation of human memory B cells. Science 298, 2199–2202 (2002).
Duffy, D.C., McDonald, J.C., Schueller, O.J.A. & Whitesides, G.M. Rapid prototyping of microfluidic systems in poly(dimethylsiloxane). Anal. Chem. 70, 4974–4984 (1998).
Duda, R.O. & Hart, P.E. Use of the Hough transformation to detect lines and curves in pictures. Commun. ACM 15, 11–15 (1972).
Acknowledgements
We would like to thank the Institut Pierre-Gilles de Gennes (IPGG) for use of clean room facilities and the laser engraver (CII08, Axyslaser), and Pfizer for the generous gift of TT-Alexa488, and CHO cell lines secreting TT4, TT7, and TT10. TT11 and TT27 antibodies are Pfizer proprietary antibodies isolated from their collaboration with HiFiBiO. We thank Pfizer (M. Holsti, G. Cheung, and W. Somers) as well as HiFiBiO Team (A. Gérard, A. Woolfe, M. Reichen, A. Poitou, S. Essonno, R. Kumar, S. Ellouze, K. Grosselin, B. Shen, and C. Brenan) for identification, rapid cloning, and validation of TT11 and TT27 antibodies. This work received support from the French Investissements d'Avenir program under the grant agreements ANR-10-NANO-02, ANR-10-IDEX-0001-02 PSL, ANR-10-LABX-31 and ANR-10- EQPX-34, by the French Agence Nationale de la Recherche (ANR-14-CE16-0011 project DROPmAbs), from Région Ile-de-France (DIM NanoK) and by the Institut Carnot Pasteur Maladies Infectieuses. K.E. acknowledges financial support from the 'Fondation Pierre-Gilles de Gennes', and the 'Swiss National Science Foundation' and the 'Society in Science—The Branco Weiss Fellowship'. C.C. acknowledges financial support from CONCYTEC, Peru. We would like to further acknowledge R. Henson for making the dscatter function for Matlab publicly available, M. Spitzer, J. Wildenhain, J. Rappsilber and M. Tyers for BoxPlotR that has been used to create the box plots, and B. Iannascoli (Unit of Antibodies in Therapy and Pathology, Institut Pasteur) for help with antibody production and cell lines.
Author information
Authors and Affiliations
Contributions
K.E., A.J., A.D.G., J. Bibette, P.B. and J. Baudry designed the study; K.E., R.C.L.D., A.G., E.B.-L., C.N., A.D.G., J. Bibette and J. Baudry developed the technology; K.E., R.C.L.D., C.E.C., L.B.-R., V.M., G.M., P.E. and A.G. performed the experiments; K.E., A.D.G., P.B. and J. Baudry analyzed the data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
J. Bibette and A.D.G. are co-founders of Seven Pines Holding BV, and HiFiBiO SAS is a subsidiary of Seven Pines Holding BV. Patents have been filed on some aspects of this work and the inventors may receive payments related to exploitation of these under their employer's rewards-to-inventors scheme.
Supplementary information
Rights and permissions
About this article
Cite this article
Eyer, K., Doineau, R., Castrillon, C. et al. Single-cell deep phenotyping of IgG-secreting cells for high-resolution immune monitoring. Nat Biotechnol 35, 977–982 (2017). https://doi.org/10.1038/nbt.3964
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3964
This article is cited by
-
Small-molecule autocatalysis drives compartment growth, competition and reproduction
Nature Chemistry (2024)
-
Adaptive immune receptor repertoire analysis
Nature Reviews Methods Primers (2024)
-
Multiplexed fluorescence and scatter detection with single cell resolution using on-chip fiber optics for droplet microfluidic applications
Microsystems & Nanoengineering (2024)
-
Single-B cell analysis correlates high-lactate secretion with stress and increased apoptosis
Scientific Reports (2024)
-
Identification of druggable regulators of cell secretion via a kinome-wide screen and high-throughput immunomagnetic cell sorting
Nature Biomedical Engineering (2023)